"Time": models, code, and papers
A CT-based deep learning system for automatic assessment of aortic root morphology for TAVI planning
Feb 10, 2023
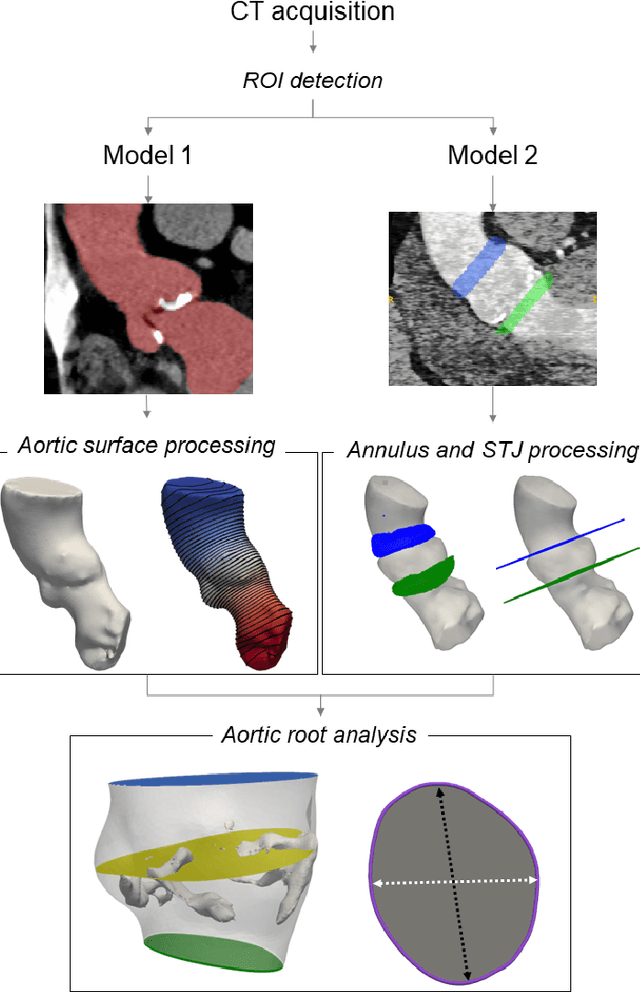
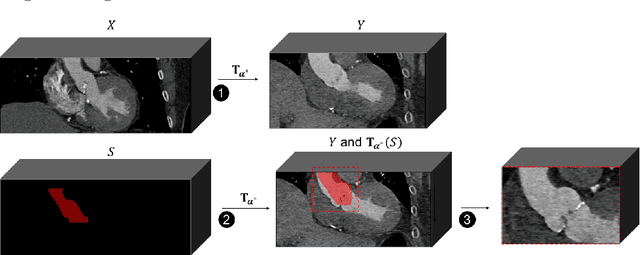
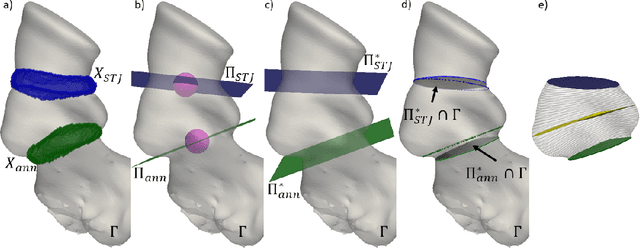
Accurate planning of transcatheter aortic implantation (TAVI) is important to minimize complications, and it requires anatomic evaluation of the aortic root (AR), commonly done through 3D computed tomography (CT) image analysis. Currently, there is no standard automated solution for this process. Two convolutional neural networks (CNNs) with 3D U-Net architectures (model 1 and model 2) were trained on 310 CT scans for AR analysis. Model 1 performed AR segmentation and model 2 identified the aortic annulus and sinotubular junction (STJ) contours. Results were validated against manual measurements of 178 TAVI candidates. After training, the two models were integrated into a fully automated pipeline for geometric analysis of the AR. The trained CNNs effectively segmented the AR, annulus and STJ, resulting in mean Dice scores of 0.93 for the AR, and mean surface distances of 1.16 mm and 1.30 mm for the annulus and STJ, respectively. Automatic measurements were in good agreement with manual annotations, yielding annulus diameters that differed by 0.52 [-2.96, 4.00] mm (bias and 95% limits of agreement for manual minus algorithm). Evaluating the area-derived diameter, bias and limits of agreement were 0.07 [-0.25, 0.39] mm. STJ and sinuses diameters computed by the automatic method yielded differences of 0.16 [-2.03, 2.34] and 0.1 [-2.93, 3.13] mm, respectively. The proposed tool is a fully automatic solution to quantify morphological biomarkers for pre-TAVI planning. The method was validated against manual annotation from clinical experts and showed to be quick and effective in assessing AR anatomy, with potential for time and cost savings.
Geometry of contact: contact planning for multi-legged robots via spin models duality
Feb 07, 2023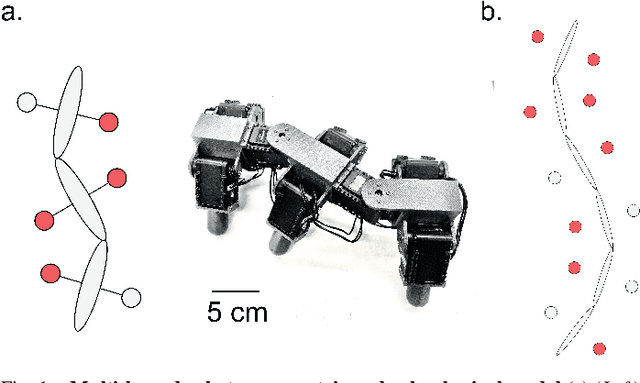
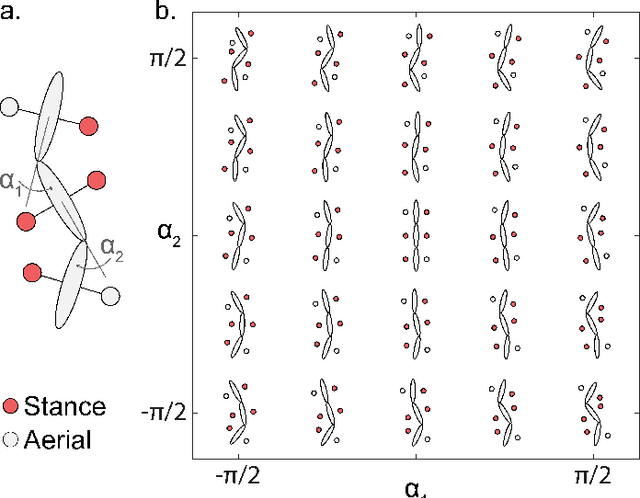
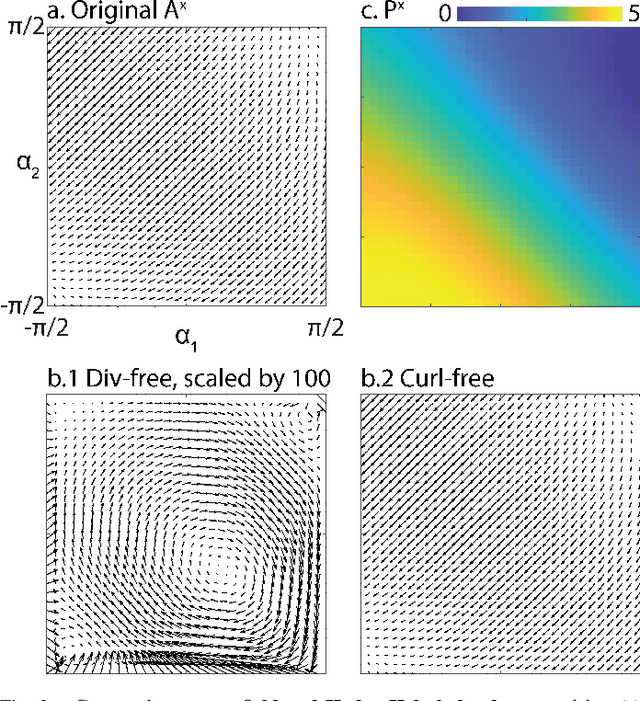
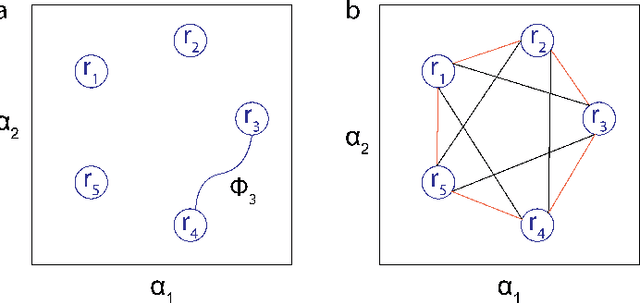
Contact planning is crucial in locomoting systems.Specifically, appropriate contact planning can enable versatile behaviors (e.g., sidewinding in limbless locomotors) and facilitate speed-dependent gait transitions (e.g., walk-trot-gallop in quadrupedal locomotors). The challenges of contact planning include determining not only the sequence by which contact is made and broken between the locomotor and the environments, but also the sequence of internal shape changes (e.g., body bending and limb shoulder joint oscillation). Most state-of-art contact planning algorithms focused on conventional robots (e.g.biped and quadruped) and conventional tasks (e.g. forward locomotion), and there is a lack of study on general contact planning in multi-legged robots. In this paper, we show that using geometric mechanics framework, we can obtain the global optimal contact sequence given the internal shape changes sequence. Therefore, we simplify the contact planning problem to a graph optimization problem to identify the internal shape changes. Taking advantages of the spatio-temporal symmetry in locomotion, we map the graph optimization problem to special cases of spin models, which allows us to obtain the global optima in polynomial time. We apply our approach to develop new forward and sidewinding behaviors in a hexapod and a 12-legged centipede. We verify our predictions using numerical and robophysical models, and obtain novel and effective locomotion behaviors.
Scalable Gaussian process regression enables accurate prediction of protein and small molecule properties with uncertainty quantitation
Feb 07, 2023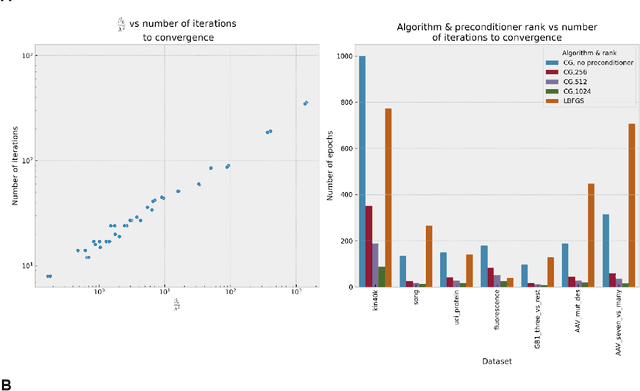
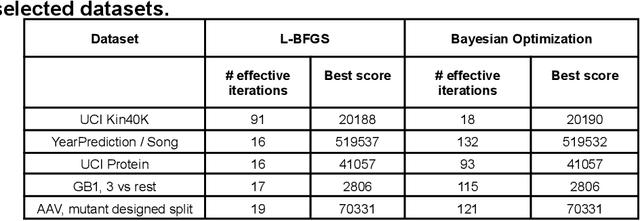
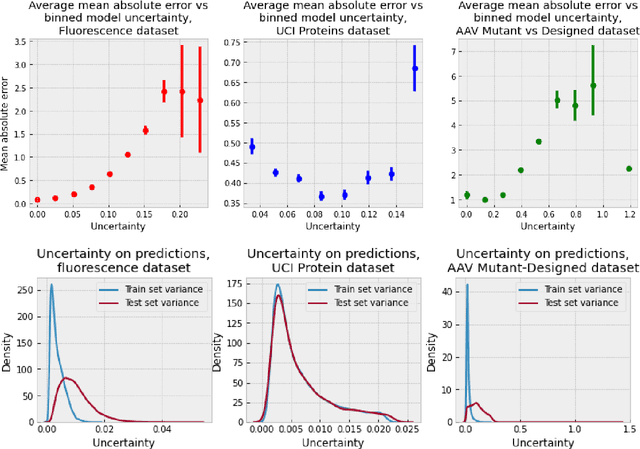
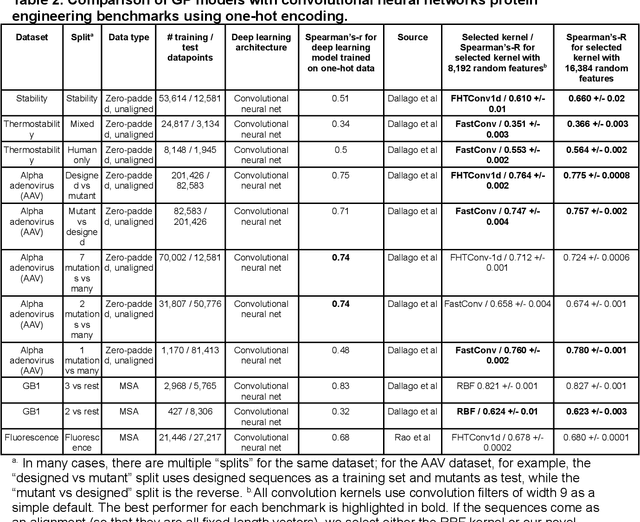
Gaussian process (GP) is a Bayesian model which provides several advantages for regression tasks in machine learning such as reliable quantitation of uncertainty and improved interpretability. Their adoption has been precluded by their excessive computational cost and by the difficulty in adapting them for analyzing sequences (e.g. amino acid and nucleotide sequences) and graphs (e.g. ones representing small molecules). In this study, we develop efficient and scalable approaches for fitting GP models as well as fast convolution kernels which scale linearly with graph or sequence size. We implement these improvements by building an open-source Python library called xGPR. We compare the performance of xGPR with the reported performance of various deep learning models on 20 benchmarks, including small molecule, protein sequence and tabular data. We show that xGRP achieves highly competitive performance with much shorter training time. Furthermore, we also develop new kernels for sequence and graph data and show that xGPR generally outperforms convolutional neural networks on predicting key properties of proteins and small molecules. Importantly, xGPR provides uncertainty information not available from typical deep learning models. Additionally, xGPR provides a representation of the input data that can be used for clustering and data visualization. These results demonstrate that xGPR provides a powerful and generic tool that can be broadly useful in protein engineering and drug discovery.
Layered State Discovery for Incremental Autonomous Exploration
Feb 07, 2023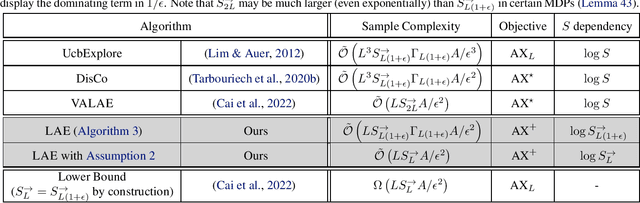
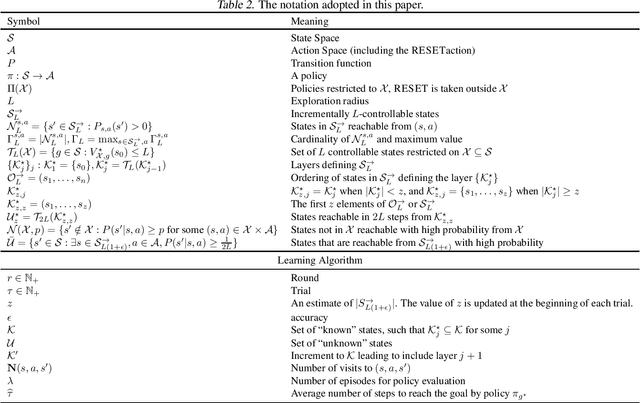

We study the autonomous exploration (AX) problem proposed by Lim & Auer (2012). In this setting, the objective is to discover a set of $\epsilon$-optimal policies reaching a set $\mathcal{S}_L^{\rightarrow}$ of incrementally $L$-controllable states. We introduce a novel layered decomposition of the set of incrementally $L$-controllable states that is based on the iterative application of a state-expansion operator. We leverage these results to design Layered Autonomous Exploration (LAE), a novel algorithm for AX that attains a sample complexity of $\tilde{\mathcal{O}}(LS^{\rightarrow}_{L(1+\epsilon)}\Gamma_{L(1+\epsilon)} A \ln^{12}(S^{\rightarrow}_{L(1+\epsilon)})/\epsilon^2)$, where $S^{\rightarrow}_{L(1+\epsilon)}$ is the number of states that are incrementally $L(1+\epsilon)$-controllable, $A$ is the number of actions, and $\Gamma_{L(1+\epsilon)}$ is the branching factor of the transitions over such states. LAE improves over the algorithm of Tarbouriech et al. (2020a) by a factor of $L^2$ and it is the first algorithm for AX that works in a countably-infinite state space. Moreover, we show that, under a certain identifiability assumption, LAE achieves minimax-optimal sample complexity of $\tilde{\mathcal{O}}(LS^{\rightarrow}_{L}A\ln^{12}(S^{\rightarrow}_{L})/\epsilon^2)$, outperforming existing algorithms and matching for the first time the lower bound proved by Cai et al. (2022) up to logarithmic factors.
Transfer learning for process design with reinforcement learning
Feb 07, 2023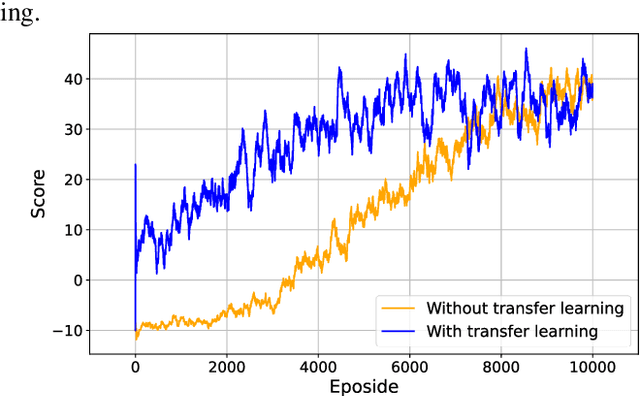
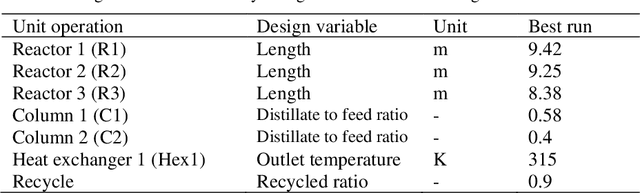

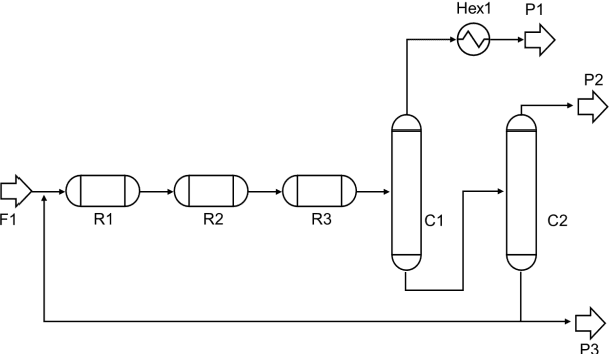
Process design is a creative task that is currently performed manually by engineers. Artificial intelligence provides new potential to facilitate process design. Specifically, reinforcement learning (RL) has shown some success in automating process design by integrating data-driven models that learn to build process flowsheets with process simulation in an iterative design process. However, one major challenge in the learning process is that the RL agent demands numerous process simulations in rigorous process simulators, thereby requiring long simulation times and expensive computational power. Therefore, typically short-cut simulation methods are employed to accelerate the learning process. Short-cut methods can, however, lead to inaccurate results. We thus propose to utilize transfer learning for process design with RL in combination with rigorous simulation methods. Transfer learning is an established approach from machine learning that stores knowledge gained while solving one problem and reuses this information on a different target domain. We integrate transfer learning in our RL framework for process design and apply it to an illustrative case study comprising equilibrium reactions, azeotropic separation, and recycles, our method can design economically feasible flowsheets with stable interaction with DWSIM. Our results show that transfer learning enables RL to economically design feasible flowsheets with DWSIM, resulting in a flowsheet with an 8% higher revenue. And the learning time can be reduced by a factor of 2.
CoViT: Real-time phylogenetics for the SARS-CoV-2 pandemic using Vision Transformers
Aug 09, 2022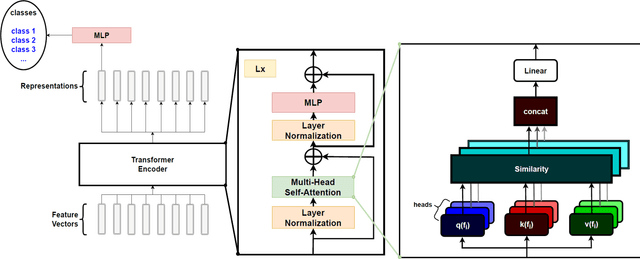
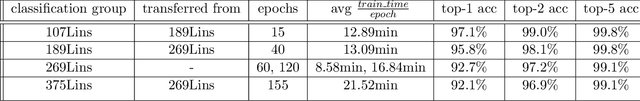
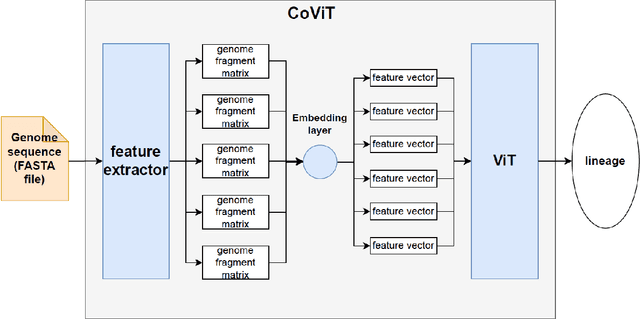

Real-time viral genome detection, taxonomic classification and phylogenetic analysis are critical for efficient tracking and control of viral pandemics such as Covid-19. However, the unprecedented and still growing amounts of viral genome data create a computational bottleneck, which effectively prevents the real-time pandemic tracking. We are attempting to alleviate this bottleneck by modifying and applying Vision Transformer, a recently developed neural network model for image recognition, to taxonomic classification and placement of viral genomes, such as SARS-CoV-2. Our solution, CoViT, places newly acquired samples onto the tree of SARS-CoV-2 lineages. One of the two potential placements returned by CoVit is the true one with the probability of 99.0%. The probability of the correct placement to be found among five potential placements generated by CoViT is 99.8%. The placement time is 1.45ms per individual genome running on NVIDIAs GeForce RTX 2080 Ti GPU. We make CoViT available to research community through GitHub: https://github.com/zuherJahshan/covit.
Neural Image Compression with a Diffusion-Based Decoder
Jan 23, 2023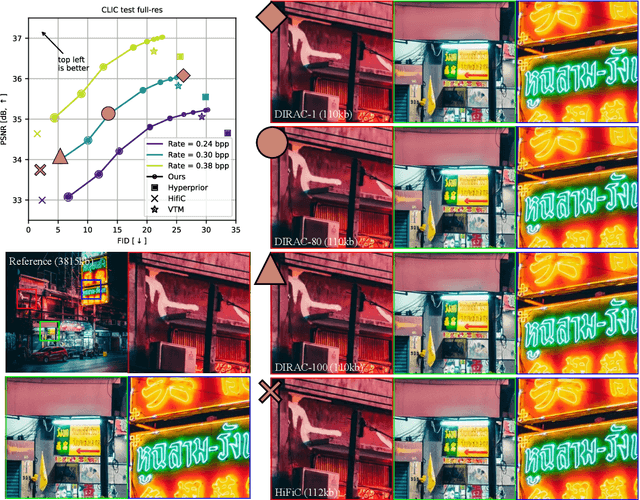
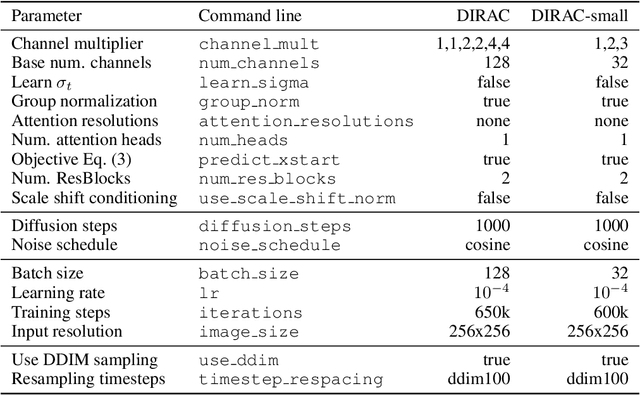
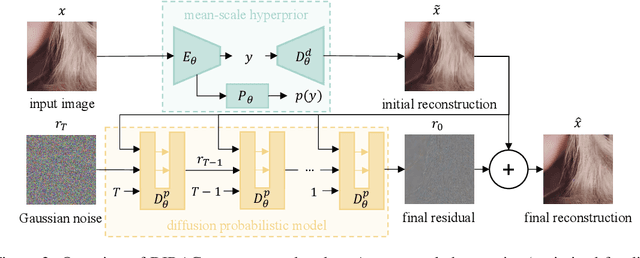
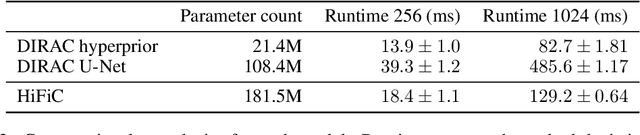
Diffusion probabilistic models have recently achieved remarkable success in generating high quality image and video data. In this work, we build on this class of generative models and introduce a method for lossy compression of high resolution images. The resulting codec, which we call DIffuson-based Residual Augmentation Codec (DIRAC),is the first neural codec to allow smooth traversal of the rate-distortion-perception tradeoff at test time, while obtaining competitive performance with GAN-based methods in perceptual quality. Furthermore, while sampling from diffusion probabilistic models is notoriously expensive, we show that in the compression setting the number of steps can be drastically reduced.
Combined Use of Federated Learning and Image Encryption for Privacy-Preserving Image Classification with Vision Transformer
Jan 23, 2023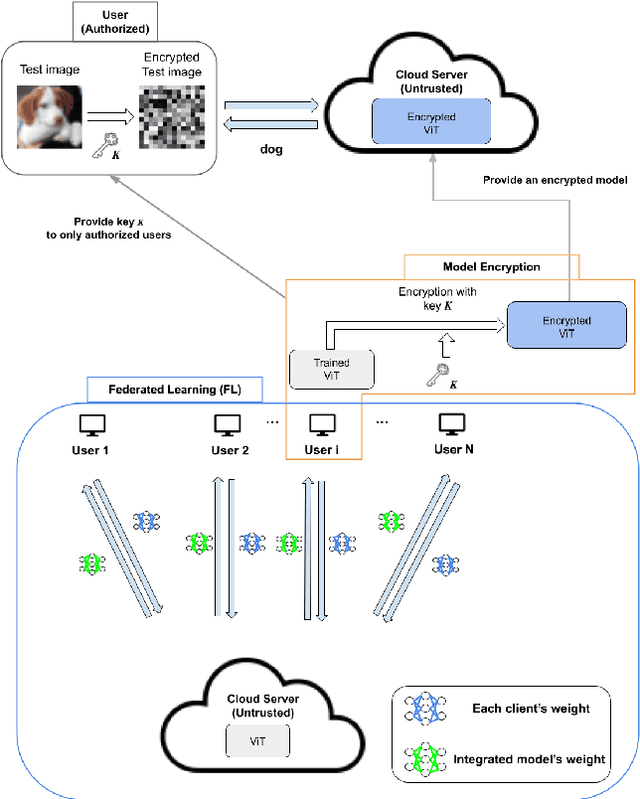
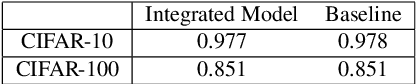
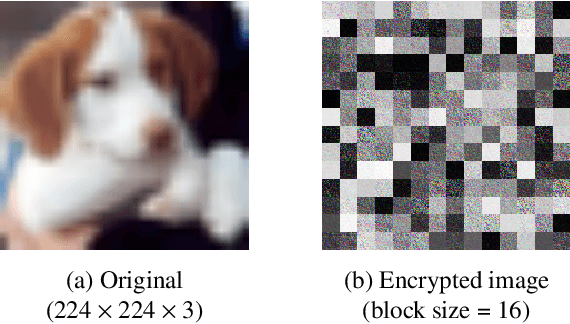


In recent years, privacy-preserving methods for deep learning have become an urgent problem. Accordingly, we propose the combined use of federated learning (FL) and encrypted images for privacy-preserving image classification under the use of the vision transformer (ViT). The proposed method allows us not only to train models over multiple participants without directly sharing their raw data but to also protect the privacy of test (query) images for the first time. In addition, it can also maintain the same accuracy as normally trained models. In an experiment, the proposed method was demonstrated to well work without any performance degradation on the CIFAR-10 and CIFAR-100 datasets.
Low-Rank Structured Clutter Covariance Matrix Estimation for Airborne STAP Radar
Jan 27, 2023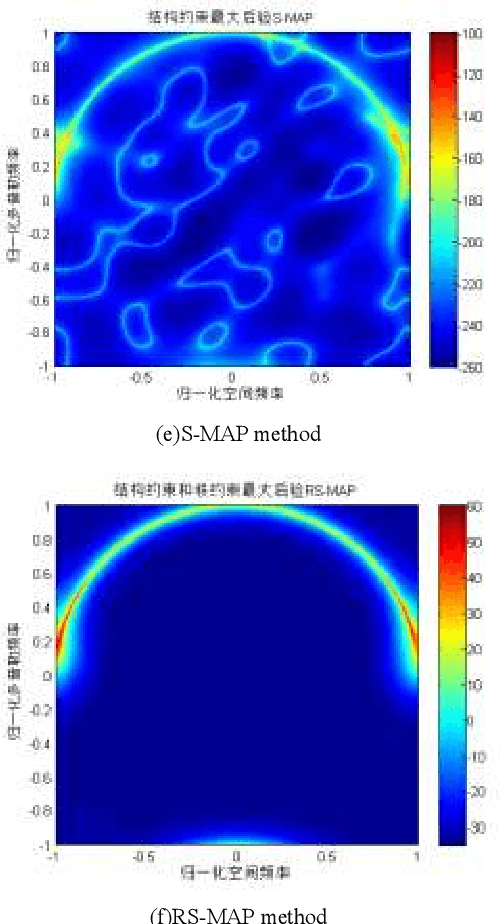
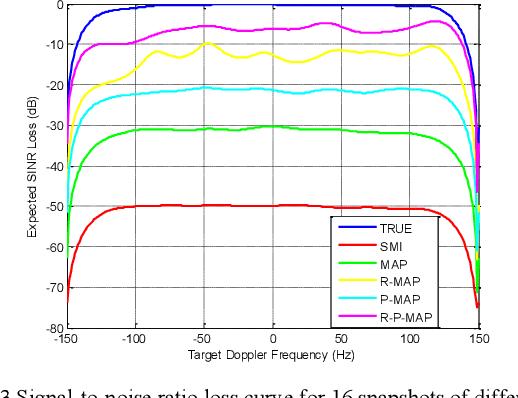
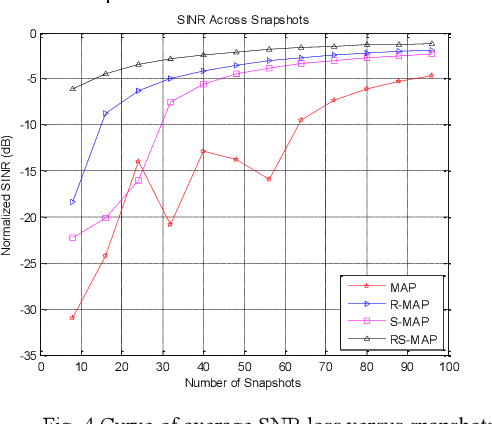
In space-time adaptive processing (STAP) of the airborne radar system, it is very important to realize sparse restoration of the clutter covariance matrix with a small number of samples. In this paper, a clutter suppression method for airborne forward-looking array radar based on joint statistics and structural priority is proposed, which can estimate the clutter covariance matrix in the case of small samples. Assuming that the clutter covariance matrix obeys the inverse Wishart prior distribution, the maximum posterior estimate is obtained by using the low-rank symmetry of the matrix itself. The simulation results based on the radar forward-looking array model show that compared with the traditional covariance matrix estimation method, the proposed method can effectively improve the clutter suppression performance of airborne radar while efficiently calculating.
Meta Temporal Point Processes
Jan 27, 2023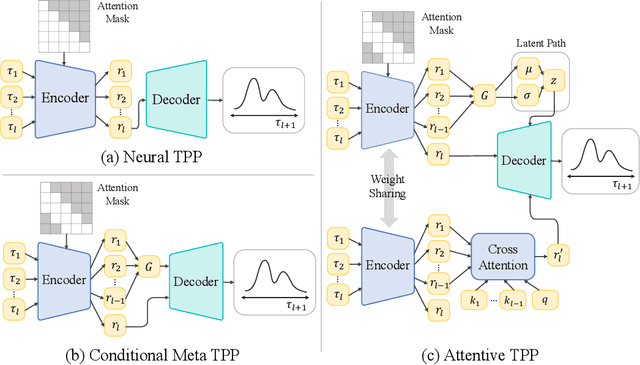
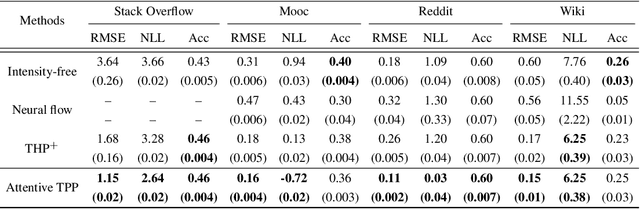
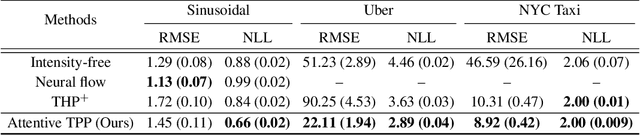
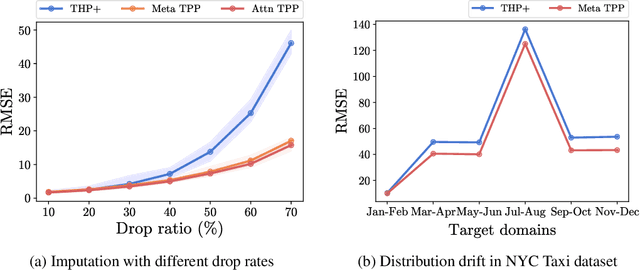
A temporal point process (TPP) is a stochastic process where its realization is a sequence of discrete events in time. Recent work in TPPs model the process using a neural network in a supervised learning framework, where a training set is a collection of all the sequences. In this work, we propose to train TPPs in a meta learning framework, where each sequence is treated as a different task, via a novel framing of TPPs as neural processes (NPs). We introduce context sets to model TPPs as an instantiation of NPs. Motivated by attentive NP, we also introduce local history matching to help learn more informative features. We demonstrate the potential of the proposed method on popular public benchmark datasets and tasks, and compare with state-of-the-art TPP methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge