"Time": models, code, and papers
CNTS: Cooperative Network for Time Series
Feb 20, 2023
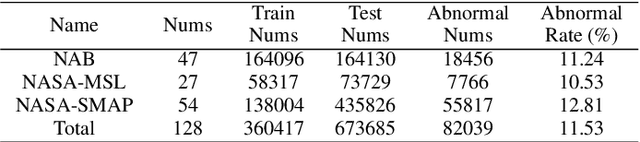
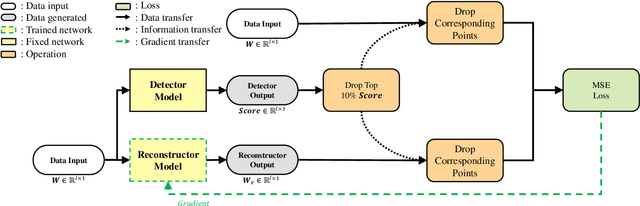
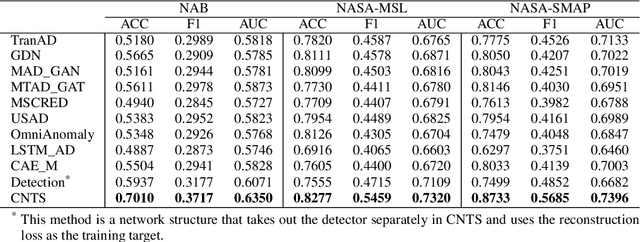
The use of deep learning techniques in detecting anomalies in time series data has been an active area of research with a long history of development and a variety of approaches. In particular, reconstruction-based unsupervised anomaly detection methods have gained popularity due to their intuitive assumptions and low computational requirements. However, these methods are often susceptible to outliers and do not effectively model anomalies, leading to suboptimal results. This paper presents a novel approach for unsupervised anomaly detection, called the Cooperative Network Time Series (CNTS) approach. The CNTS system consists of two components: a detector and a reconstructor. The detector is responsible for directly detecting anomalies, while the reconstructor provides reconstruction information to the detector and updates its learning based on anomalous information received from the detector. The central aspect of CNTS is a multi-objective optimization problem, which is solved through a cooperative solution strategy. Experiments on three real-world datasets demonstrate the state-of-the-art performance of CNTS and confirm the cooperative effectiveness of the detector and reconstructor. The source code for this study is publicly available on GitHub.
Contrastive Loss is All You Need to Recover Analogies as Parallel Lines
Jun 14, 2023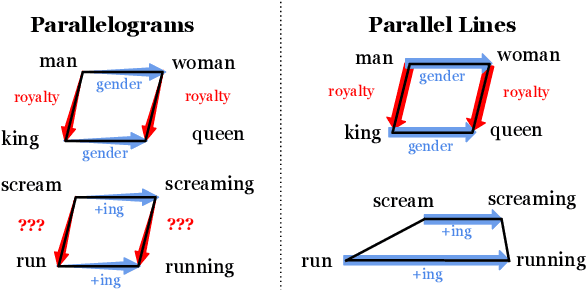
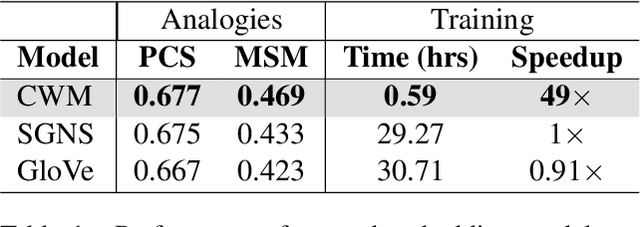

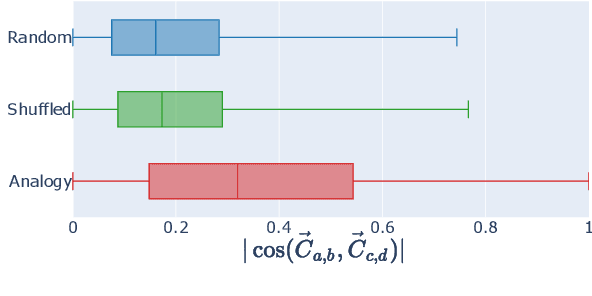
While static word embedding models are known to represent linguistic analogies as parallel lines in high-dimensional space, the underlying mechanism as to why they result in such geometric structures remains obscure. We find that an elementary contrastive-style method employed over distributional information performs competitively with popular word embedding models on analogy recovery tasks, while achieving dramatic speedups in training time. Further, we demonstrate that a contrastive loss is sufficient to create these parallel structures in word embeddings, and establish a precise relationship between the co-occurrence statistics and the geometric structure of the resulting word embeddings.
Energy-Efficient MIMO Integrated Sensing and Communications with On-off Non-transmission Power
Jun 27, 2023This paper investigates the energy efficiency of a multiple-input multiple-output (MIMO) integrated sensing and communications (ISAC) system, in which one multi-antenna base station (BS) transmits unified ISAC signals to a multi-antenna communication user (CU) and at the same time use the echo signals to estimate an extended target. We focus on one particular ISAC transmission block and take into account the practical on-off non-transmission power at the BS. Under this setup, we minimize the energy consumption at the BS while ensuring a minimum average data rate requirement for communication and a maximum Cram\'er-Rao bound (CRB) requirement for target estimation, by jointly optimizing the transmit covariance matrix and the ``on'' duration for active transmission. We obtain the optimal solution to the rate-and-CRB-constrained energy minimization problem in a semi-closed form. Interestingly, the obtained optimal solution is shown to unify the spectrum-efficient and energy-efficient communications and sensing designs. In particular, for the special MIMO sensing case with rate constraint inactive, the optimal solution follows the isotropic transmission with shortest ``on'' duration, in which the BS radiates the required sensing energy by using sufficiently high power over the shortest duration. For the general ISAC case, the optimal transmit covariance solution is of full rank and follows the eigenmode transmission based on the communication channel, while the optimal ``on'' duration is determined based on both the rate and CRB constraints. Numerical results show that the proposed ISAC design achieves significantly reduced energy consumption as compared to the benchmark schemes based on isotropic transmission, always-on transmission, and sensing or communications only designs, especially when the rate and CRB constraints become stringent.
Phase Space Analysis of Cardiac Spectra
Jun 27, 2023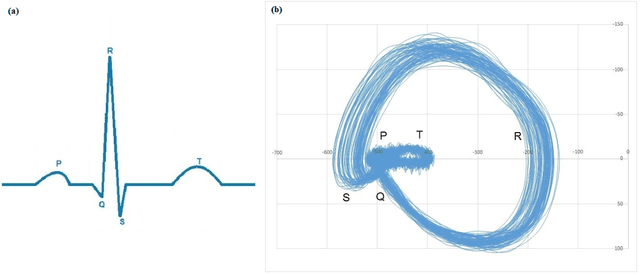
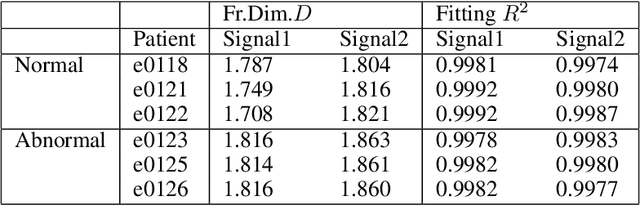
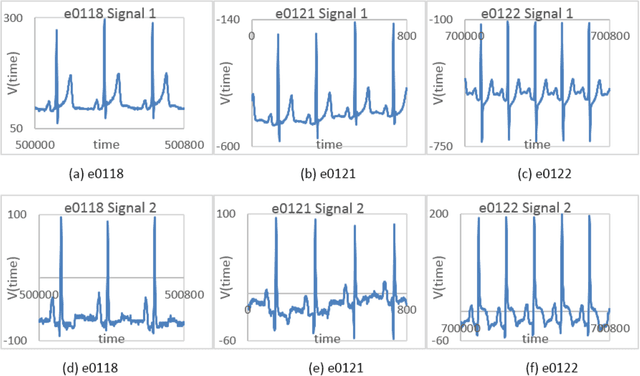
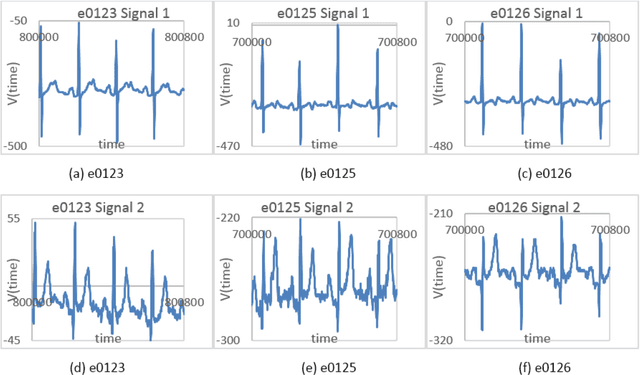
Cardiac diseases are one of the main reasons of mortality in modern, industrialized societies, and they cause high expenses in public health systems. Therefore, it is important to develop analytical methods to improve cardiac diagnostics. Electric activity of heart was first modeled by using a set of nonlinear differential equations. Latter, variations of cardiac spectra originated from deterministic dynamics are investigated. Analyzing the power spectra of a normal human heart presents His-Purkinje network, possessing a fractal like structure. Phase space trajectories are extracted from the time series graph of ECG. Lower values of fractal dimension, D indicate dynamics that are more coherent. If D has non-integer values greater than two when the system becomes chaotic or strange attractor. Recently, the development of a fast and robust method, which can be applied to multichannel physiologic signals, was reported. This manuscript investigates two different ECG systems produced from normal and abnormal human hearts to introduce an auxiliary phase space method in conjunction with ECG signals for diagnoses of heart diseases. Here, the data for each person includes two signals based on V_4 and modified lead III (MLIII) respectively. Fractal analysis method is employed on the trajectories constructed in phase space, from which the fractal dimension D is obtained using the box counting method. It is observed that, MLIII signals have larger D values than the first signals (V_4), predicting more randomness yet more information. The lowest value of D (1.708) indicates the perfect oscillation of the normal heart and the highest value of D (1.863) presents the randomness of the abnormal heart. Our significant finding is that the phase space picture presents the distribution of the peak heights from the ECG spectra, giving valuable information about heart activities in conjunction with ECG.
HyenaDNA: Long-Range Genomic Sequence Modeling at Single Nucleotide Resolution
Jun 27, 2023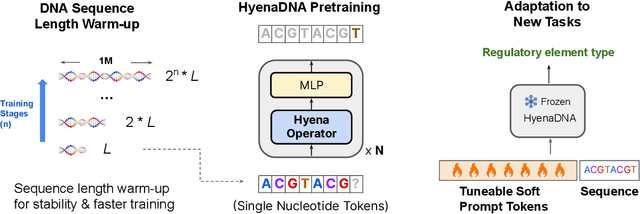
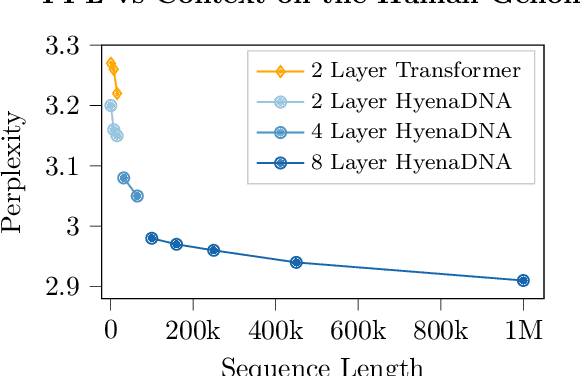
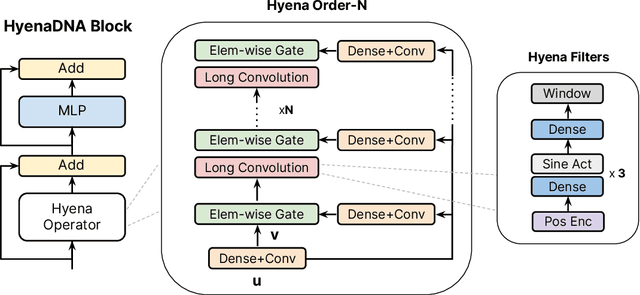
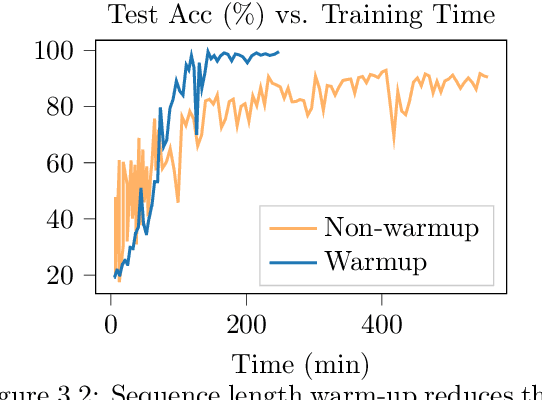
Genomic (DNA) sequences encode an enormous amount of information for gene regulation and protein synthesis. Similar to natural language models, researchers have proposed foundation models in genomics to learn generalizable features from unlabeled genome data that can then be fine-tuned for downstream tasks such as identifying regulatory elements. Due to the quadratic scaling of attention, previous Transformer-based genomic models have used 512 to 4k tokens as context (<0.001% of the human genome), significantly limiting the modeling of long-range interactions in DNA. In addition, these methods rely on tokenizers to aggregate meaningful DNA units, losing single nucleotide resolution where subtle genetic variations can completely alter protein function via single nucleotide polymorphisms (SNPs). Recently, Hyena, a large language model based on implicit convolutions was shown to match attention in quality while allowing longer context lengths and lower time complexity. Leveraging Hyenas new long-range capabilities, we present HyenaDNA, a genomic foundation model pretrained on the human reference genome with context lengths of up to 1 million tokens at the single nucleotide-level, an up to 500x increase over previous dense attention-based models. HyenaDNA scales sub-quadratically in sequence length (training up to 160x faster than Transformer), uses single nucleotide tokens, and has full global context at each layer. We explore what longer context enables - including the first use of in-context learning in genomics for simple adaptation to novel tasks without updating pretrained model weights. On fine-tuned benchmarks from the Nucleotide Transformer, HyenaDNA reaches state-of-the-art (SotA) on 12 of 17 datasets using a model with orders of magnitude less parameters and pretraining data. On the GenomicBenchmarks, HyenaDNA surpasses SotA on all 8 datasets on average by +9 accuracy points.
DeepVATS: Deep Visual Analytics for Time Series
Feb 08, 2023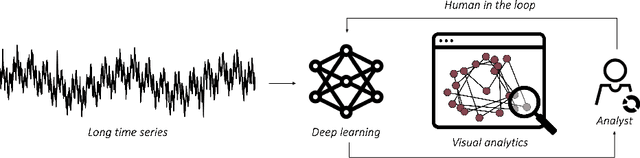
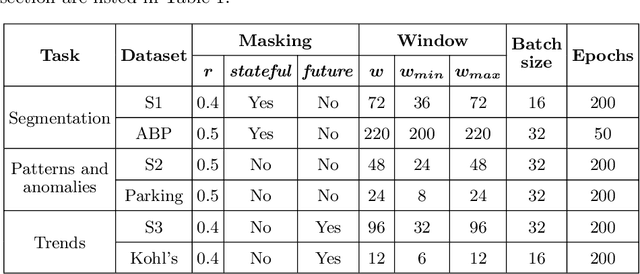
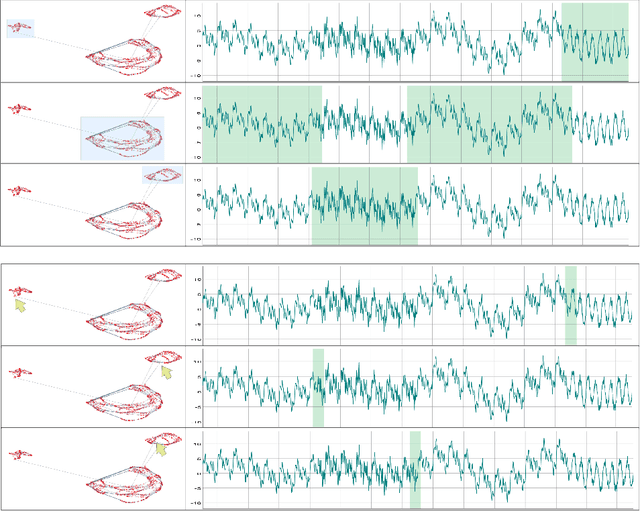
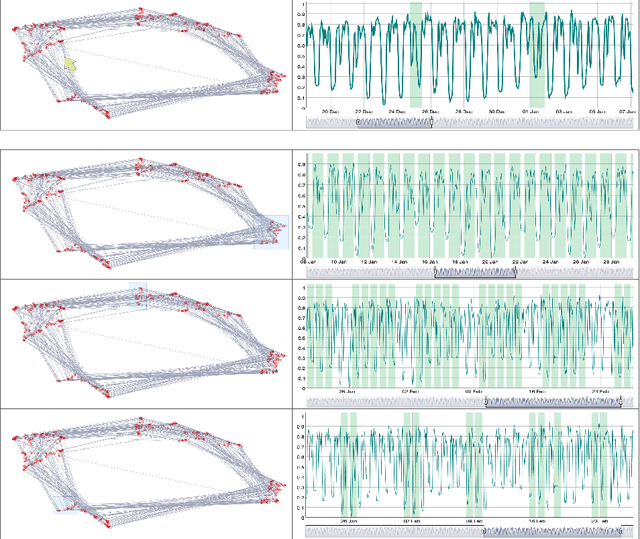
The field of Deep Visual Analytics (DVA) has recently arisen from the idea of developing Visual Interactive Systems supported by deep learning techniques, in order to provide them with large-scale data processing capabilities and to unify their implementation across different data modalities and domains of application. In this paper we present DeepVATS, an open-source tool that brings the field of DVA into time series data. DeepVATS trains, in a self-supervised way, a masked time series autoencoder that reconstructs patches of a time series, and projects the knowledge contained in the embeddings of that model in an interactive plot, from which time series patterns and anomalies emerge and can be easily spotted. The tool has been tested on both synthetic and real datasets, and its code is publicly available on https://github.com/vrodriguezf/deepvats
Onsets and Velocities: Affordable Real-Time Piano Transcription Using Convolutional Neural Networks
Mar 08, 2023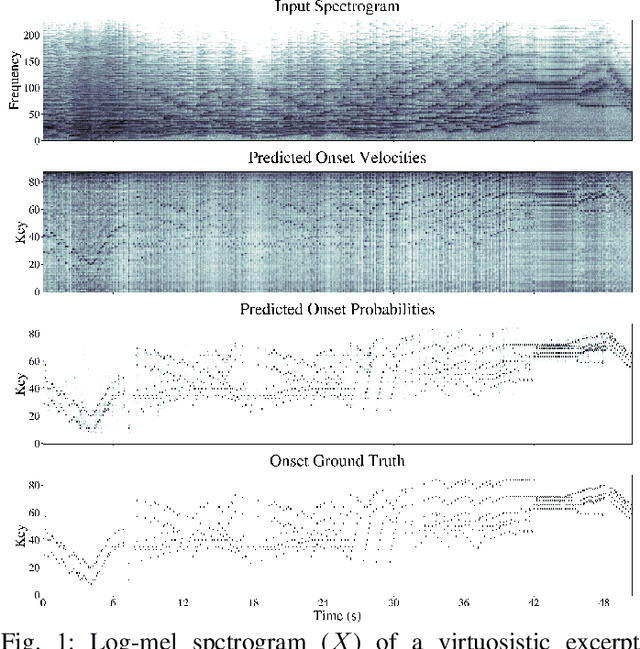
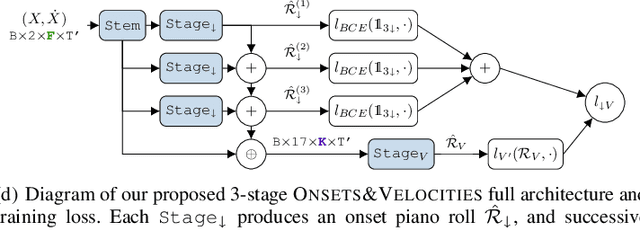
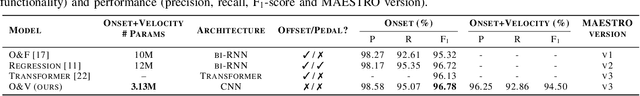
Polyphonic Piano Transcription has recently experienced substantial progress driven by the application of sophisticated Deep Learning setups and the introduction of new subtasks such as note onset, offset, velocity and pedal detection. In this work, we focus on onset and velocity detection, presenting a convolutional neural network with substantially reduced size (3.1M parameters) and a simple inference scheme that achieves state-of-the-art performance on the MAESTRO dataset for onset detection (F1=96.78%) and sets a good novel baseline for onset+velocity (F1=94.50%), while maintaining real-time capabilities on modest commodity hardware. Furthermore, our proposed ONSETS&VELOCITIES (O&V) model shows that a time resolution of 24ms is competitive, countering recent trends. We provide open-source software to reproduce our results and a real-time demo with a pretrained model.
Neuro-Modulated Hebbian Learning for Fully Test-Time Adaptation
Mar 02, 2023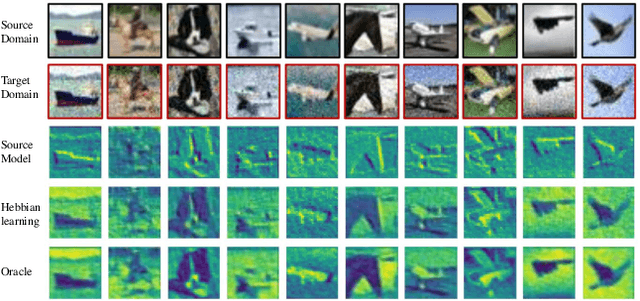
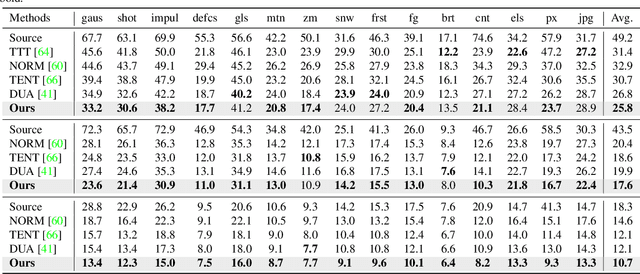
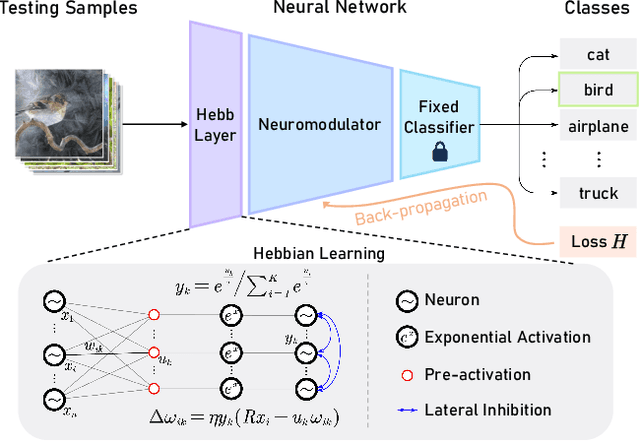
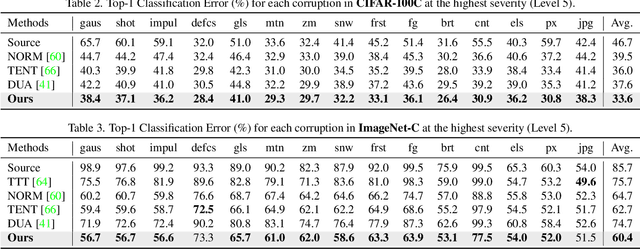
Fully test-time adaptation aims to adapt the network model based on sequential analysis of input samples during the inference stage to address the cross-domain performance degradation problem of deep neural networks. We take inspiration from the biological plausibility learning where the neuron responses are tuned based on a local synapse-change procedure and activated by competitive lateral inhibition rules. Based on these feed-forward learning rules, we design a soft Hebbian learning process which provides an unsupervised and effective mechanism for online adaptation. We observe that the performance of this feed-forward Hebbian learning for fully test-time adaptation can be significantly improved by incorporating a feedback neuro-modulation layer. It is able to fine-tune the neuron responses based on the external feedback generated by the error back-propagation from the top inference layers. This leads to our proposed neuro-modulated Hebbian learning (NHL) method for fully test-time adaptation. With the unsupervised feed-forward soft Hebbian learning being combined with a learned neuro-modulator to capture feedback from external responses, the source model can be effectively adapted during the testing process. Experimental results on benchmark datasets demonstrate that our proposed method can significantly improve the adaptation performance of network models and outperforms existing state-of-the-art methods.
Modeling Complex Object Changes in Satellite Image Time-Series: Approach based on CSP and Spatiotemporal Graph
May 24, 2023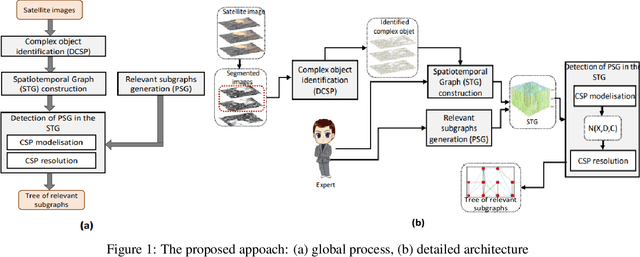
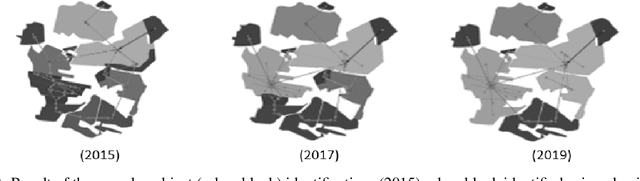
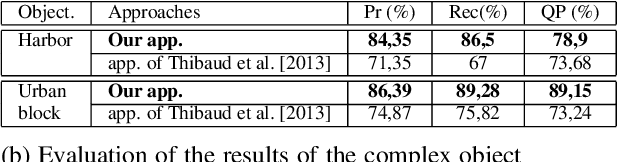
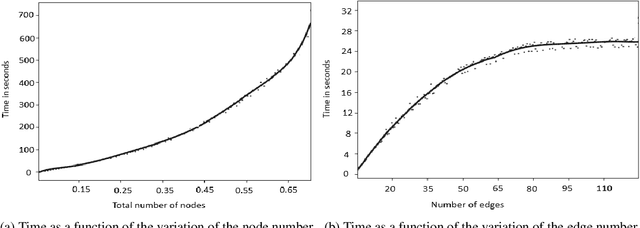
This paper proposes a method for automatically monitoring and analyzing the evolution of complex geographic objects. The objects are modeled as a spatiotemporal graph, which separates filiation relations, spatial relations, and spatiotemporal relations, and is analyzed by detecting frequent sub-graphs using constraint satisfaction problems (CSP). The process is divided into four steps: first, the identification of complex objects in each satellite image; second, the construction of a spatiotemporal graph to model the spatiotemporal changes of the complex objects; third, the creation of sub-graphs to be detected in the base spatiotemporal graph; and fourth, the analysis of the spatiotemporal graph by detecting the sub-graphs and solving a constraint network to determine relevant sub-graphs. The final step is further broken down into two sub-steps: (i) the modeling of the constraint network with defined variables and constraints, and (ii) the solving of the constraint network to find relevant sub-graphs in the spatiotemporal graph. Experiments were conducted using real-world satellite images representing several cities in Saudi Arabia, and the results demonstrate the effectiveness of the proposed approach.
G-CAME: Gaussian-Class Activation Mapping Explainer for Object Detectors
Jun 06, 2023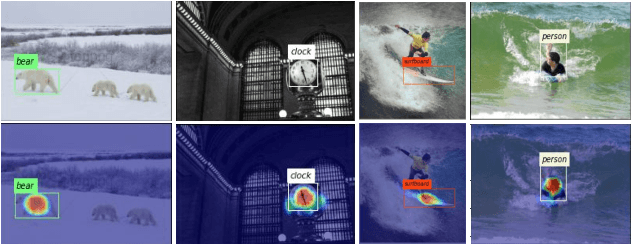
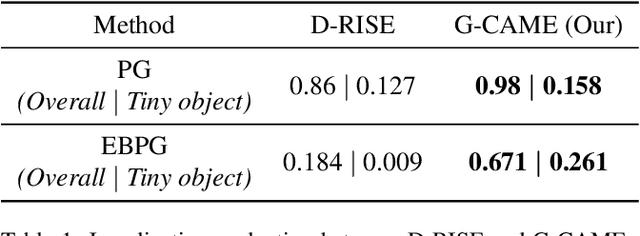
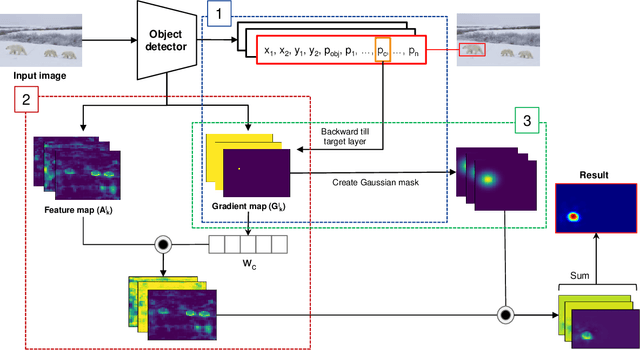
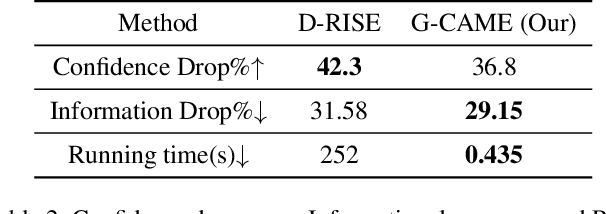
Nowadays, deep neural networks for object detection in images are very prevalent. However, due to the complexity of these networks, users find it hard to understand why these objects are detected by models. We proposed Gaussian Class Activation Mapping Explainer (G-CAME), which generates a saliency map as the explanation for object detection models. G-CAME can be considered a CAM-based method that uses the activation maps of selected layers combined with the Gaussian kernel to highlight the important regions in the image for the predicted box. Compared with other Region-based methods, G-CAME can transcend time constraints as it takes a very short time to explain an object. We also evaluated our method qualitatively and quantitatively with YOLOX on the MS-COCO 2017 dataset and guided to apply G-CAME into the two-stage Faster-RCNN model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge