"Information": models, code, and papers
Metric-Type Identification for Multi-Level Header Numerical Tables in Scientific Papers
Feb 01, 2021
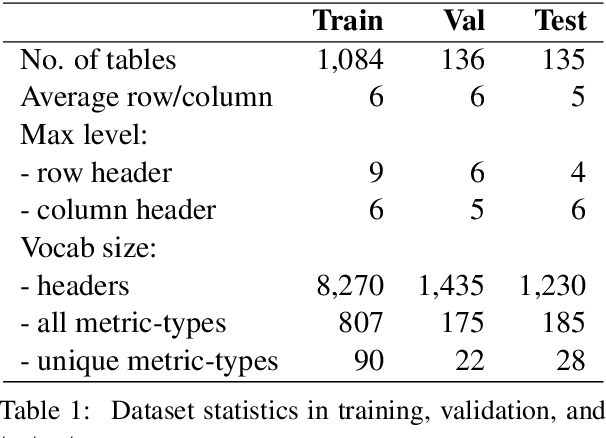
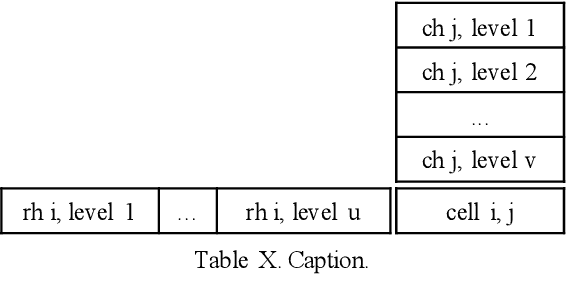
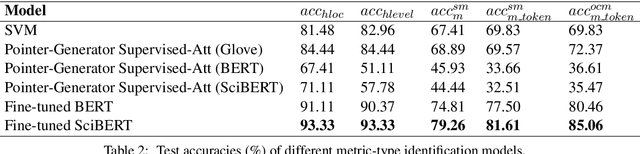
Numerical tables are widely used to present experimental results in scientific papers. For table understanding, a metric-type is essential to discriminate numbers in the tables. We introduce a new information extraction task, metric-type identification from multi-level header numerical tables, and provide a dataset extracted from scientific papers consisting of header tables, captions, and metric-types. We then propose two joint-learning neural classification and generation schemes featuring pointer-generator-based and BERT-based models. Our results show that the joint models can handle both in-header and out-of-header metric-type identification problems.
Location-aware Single Image Reflection Removal
Dec 13, 2020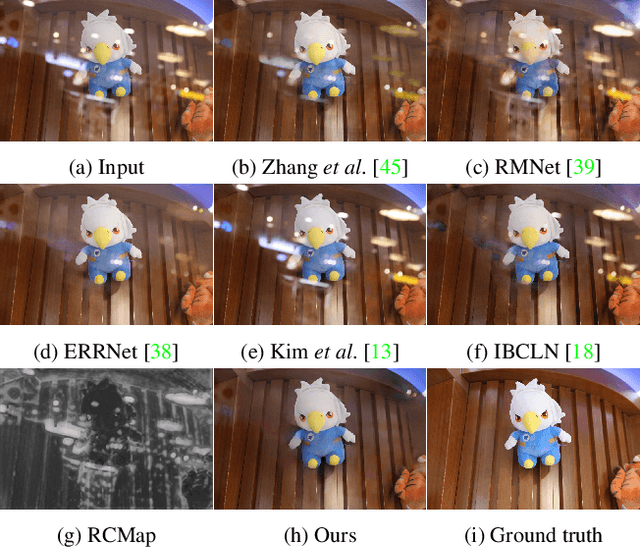
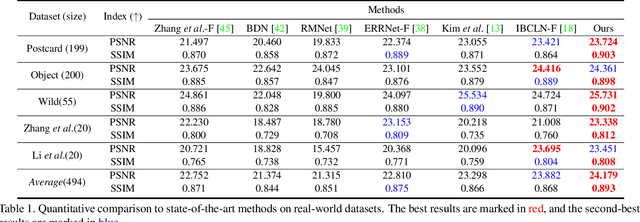
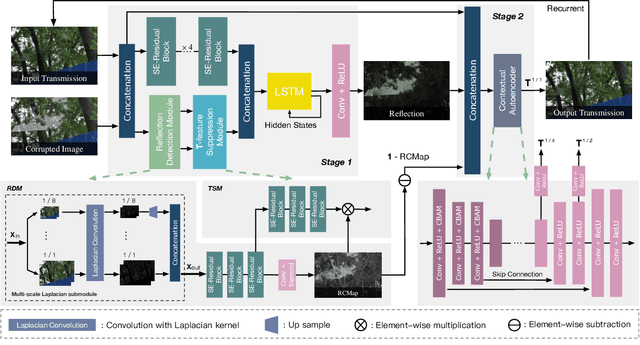
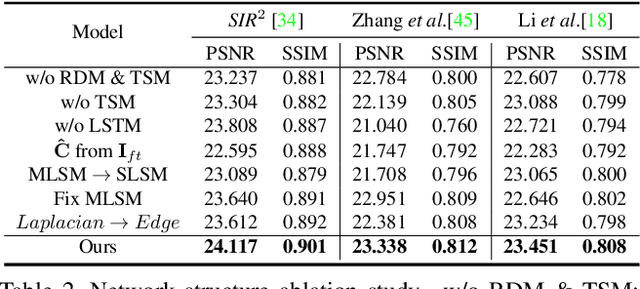
This paper proposes a novel location-aware deep learning-based single image reflection removal method. Our network has a reflection detection module to regress a probabilistic reflection confidence map, taking multi-scale Laplacian features as inputs. This probabilistic map tells whether a region is reflection-dominated or transmission-dominated. The novelty is that we use the reflection confidence map as the cues for the network to learn how to encode the reflection information adaptively and control the feature flow when predicting reflection and transmission layers. The integration of location information into the network significantly improves the quality of reflection removal results. Besides, a set of learnable Laplacian kernel parameters is introduced to facilitate the extraction of discriminative Laplacian features for reflection detection. We design our network as a recurrent network to progressively refine each iteration's reflection removal results. Extensive experiments verify the superior performance of the proposed method over state-of-the-art approaches.
Dilated Fully Convolutional Neural Network for Depth Estimation from a Single Image
Mar 12, 2021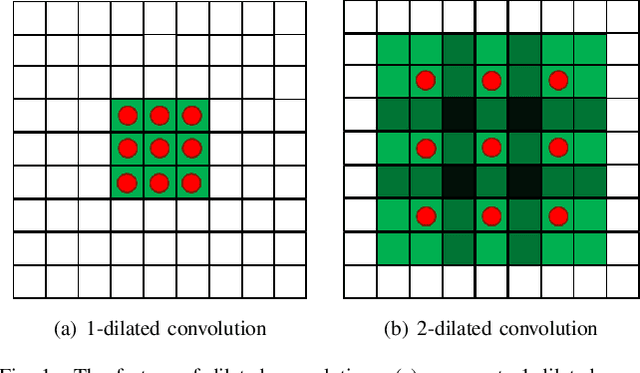
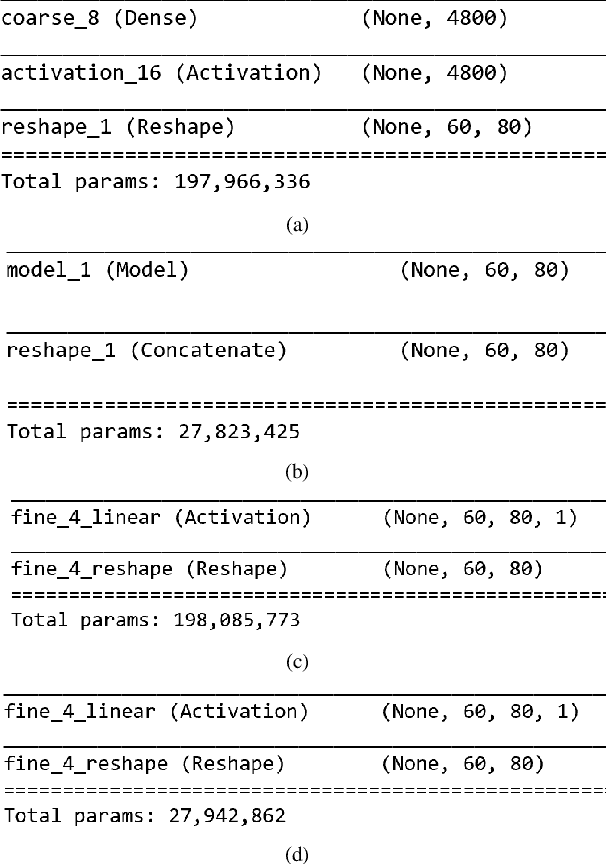
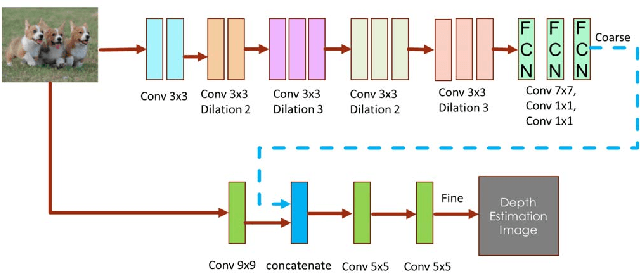


Depth prediction plays a key role in understanding a 3D scene. Several techniques have been developed throughout the years, among which Convolutional Neural Network has recently achieved state-of-the-art performance on estimating depth from a single image. However, traditional CNNs suffer from the lower resolution and information loss caused by the pooling layers. And oversized parameters generated from fully connected layers often lead to a exploded memory usage problem. In this paper, we present an advanced Dilated Fully Convolutional Neural Network to address the deficiencies. Taking advantages of the exponential expansion of the receptive field in dilated convolutions, our model can minimize the loss of resolution. It also reduces the amount of parameters significantly by replacing the fully connected layers with the fully convolutional layers. We show experimentally on NYU Depth V2 datasets that the depth prediction obtained from our model is considerably closer to ground truth than that from traditional CNNs techniques.
Graph Ensemble Learning over Multiple Dependency Trees for Aspect-level Sentiment Classification
Mar 12, 2021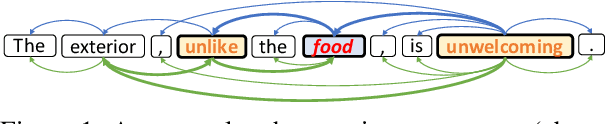
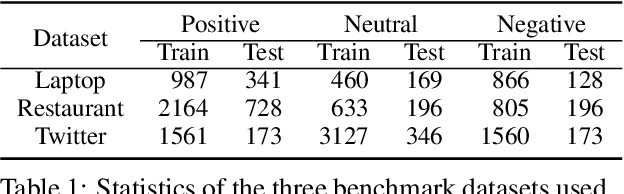
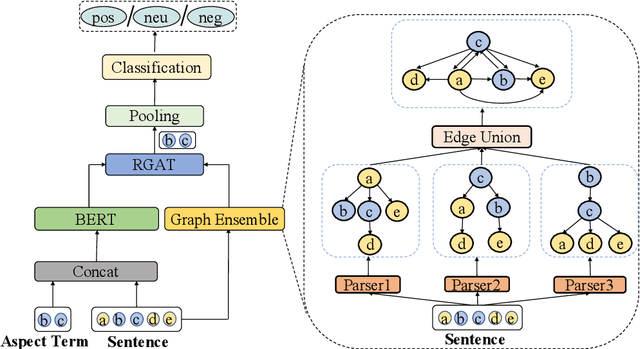
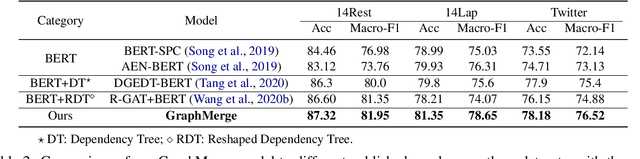
Recent work on aspect-level sentiment classification has demonstrated the efficacy of incorporating syntactic structures such as dependency trees with graph neural networks(GNN), but these approaches are usually vulnerable to parsing errors. To better leverage syntactic information in the face of unavoidable errors, we propose a simple yet effective graph ensemble technique, GraphMerge, to make use of the predictions from differ-ent parsers. Instead of assigning one set of model parameters to each dependency tree, we first combine the dependency relations from different parses before applying GNNs over the resulting graph. This allows GNN mod-els to be robust to parse errors at no additional computational cost, and helps avoid overparameterization and overfitting from GNN layer stacking by introducing more connectivity into the ensemble graph. Our experiments on the SemEval 2014 Task 4 and ACL 14 Twitter datasets show that our GraphMerge model not only outperforms models with single dependency tree, but also beats other ensemble mod-els without adding model parameters.
Topological Data Analysis of copy number alterations in cancer
Nov 22, 2020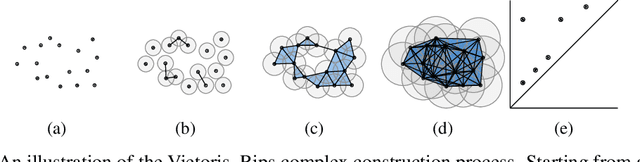
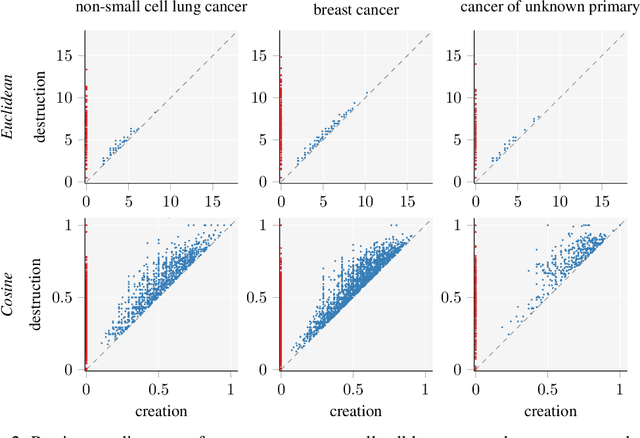
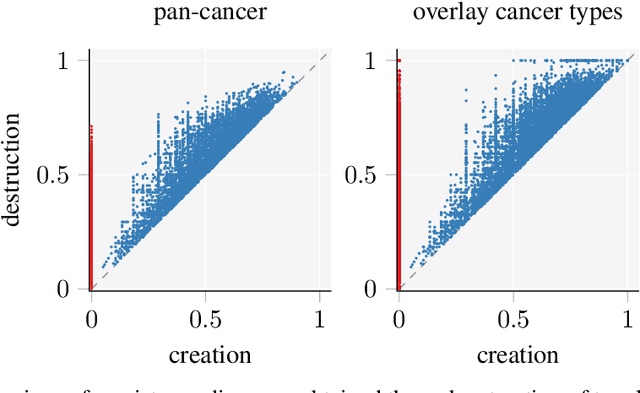
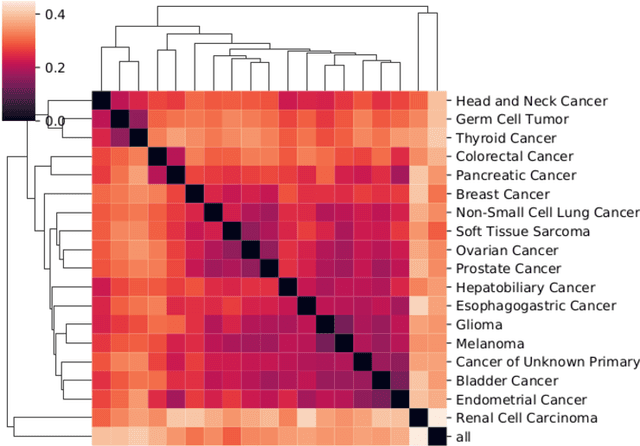
Identifying subgroups and properties of cancer biopsy samples is a crucial step towards obtaining precise diagnoses and being able to perform personalized treatment of cancer patients. Recent data collections provide a comprehensive characterization of cancer cell data, including genetic data on copy number alterations (CNAs). We explore the potential to capture information contained in cancer genomic information using a novel topology-based approach that encodes each cancer sample as a persistence diagram of topological features, i.e., high-dimensional voids represented in the data. We find that this technique has the potential to extract meaningful low-dimensional representations in cancer somatic genetic data and demonstrate the viability of some applications on finding substructures in cancer data as well as comparing similarity of cancer types.
Explainable AI For COVID-19 CT Classifiers: An Initial Comparison Study
Apr 25, 2021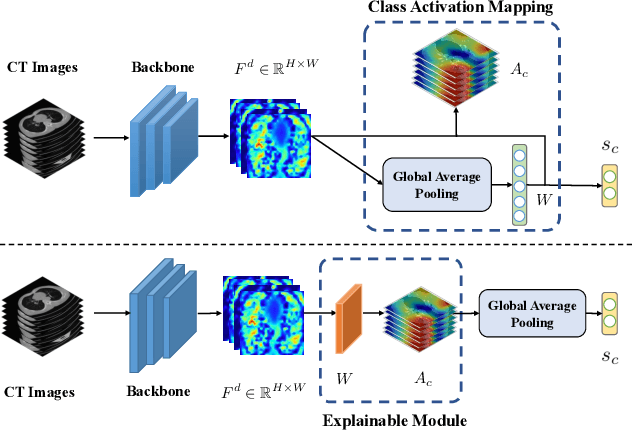
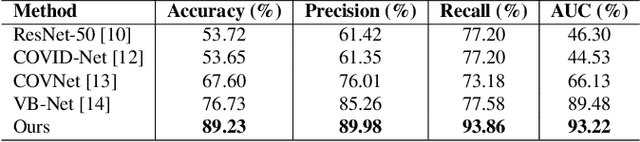
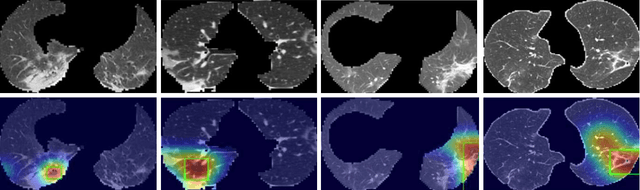
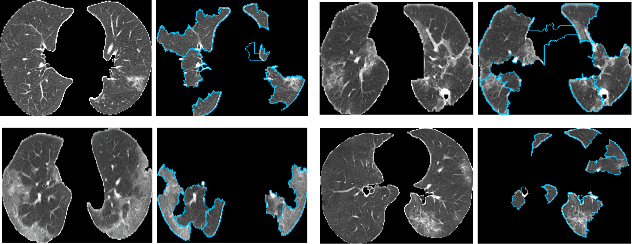
Artificial Intelligence (AI) has made leapfrogs in development across all the industrial sectors especially when deep learning has been introduced. Deep learning helps to learn the behaviour of an entity through methods of recognising and interpreting patterns. Despite its limitless potential, the mystery is how deep learning algorithms make a decision in the first place. Explainable AI (XAI) is the key to unlocking AI and the black-box for deep learning. XAI is an AI model that is programmed to explain its goals, logic, and decision making so that the end users can understand. The end users can be domain experts, regulatory agencies, managers and executive board members, data scientists, users that use AI, with or without awareness, or someone who is affected by the decisions of an AI model. Chest CT has emerged as a valuable tool for the clinical diagnostic and treatment management of the lung diseases associated with COVID-19. AI can support rapid evaluation of CT scans to differentiate COVID-19 findings from other lung diseases. However, how these AI tools or deep learning algorithms reach such a decision and which are the most influential features derived from these neural networks with typically deep layers are not clear. The aim of this study is to propose and develop XAI strategies for COVID-19 classification models with an investigation of comparison. The results demonstrate promising quantification and qualitative visualisations that can further enhance the clinician's understanding and decision making with more granular information from the results given by the learned XAI models.
Saliency-Guided Deep Learning Network for Automatic Tumor Bed Volume Delineation in Post-operative Breast Irradiation
May 06, 2021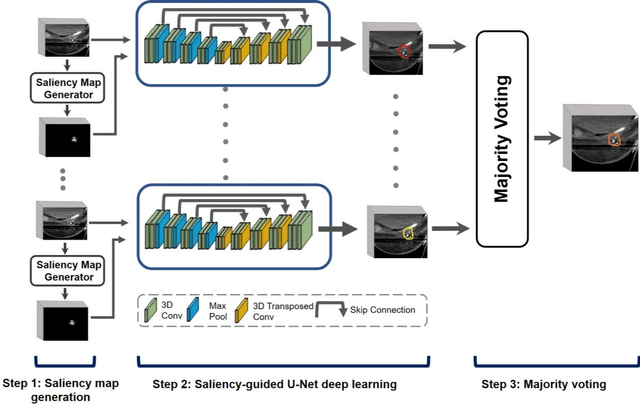
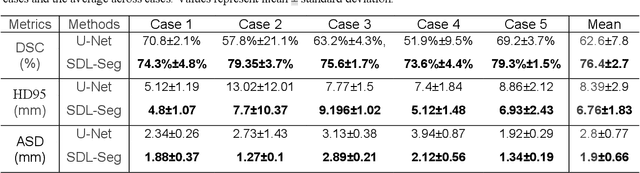
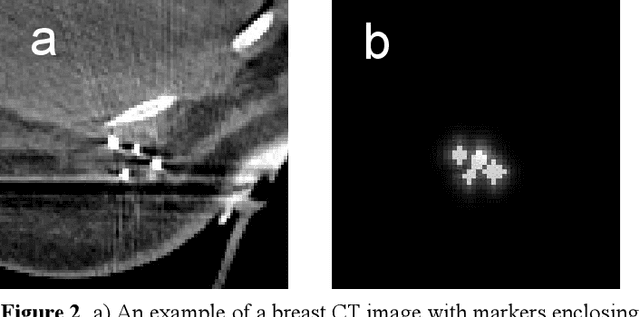
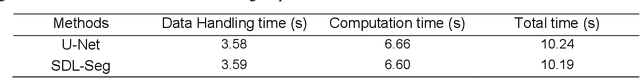
Efficient, reliable and reproducible target volume delineation is a key step in the effective planning of breast radiotherapy. However, post-operative breast target delineation is challenging as the contrast between the tumor bed volume (TBV) and normal breast tissue is relatively low in CT images. In this study, we propose to mimic the marker-guidance procedure in manual target delineation. We developed a saliency-based deep learning segmentation (SDL-Seg) algorithm for accurate TBV segmentation in post-operative breast irradiation. The SDL-Seg algorithm incorporates saliency information in the form of markers' location cues into a U-Net model. The design forces the model to encode the location-related features, which underscores regions with high saliency levels and suppresses low saliency regions. The saliency maps were generated by identifying markers on CT images. Markers' locations were then converted to probability maps using a distance-transformation coupled with a Gaussian filter. Subsequently, the CT images and the corresponding saliency maps formed a multi-channel input for the SDL-Seg network. Our in-house dataset was comprised of 145 prone CT images from 29 post-operative breast cancer patients, who received 5-fraction partial breast irradiation (PBI) regimen on GammaPod. The performance of the proposed method was compared against basic U-Net. Our model achieved mean (standard deviation) of 76.4 %, 6.76 mm, and 1.9 mm for DSC, HD95, and ASD respectively on the test set with computation time of below 11 seconds per one CT volume. SDL-Seg showed superior performance relative to basic U-Net for all the evaluation metrics while preserving low computation cost. The findings demonstrate that SDL-Seg is a promising approach for improving the efficiency and accuracy of the on-line treatment planning procedure of PBI, such as GammaPod based PBI.
Discovering key topics from short, real-world medical inquiries via natural language processing and unsupervised learning
Dec 08, 2020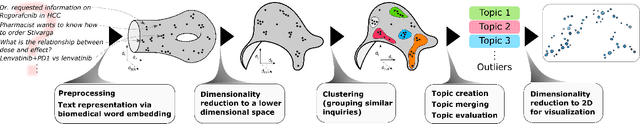
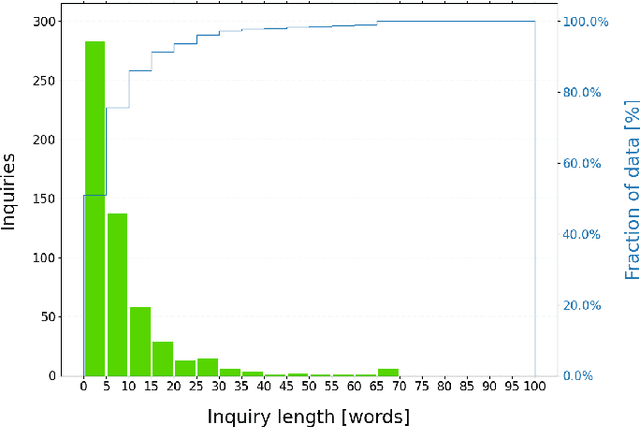
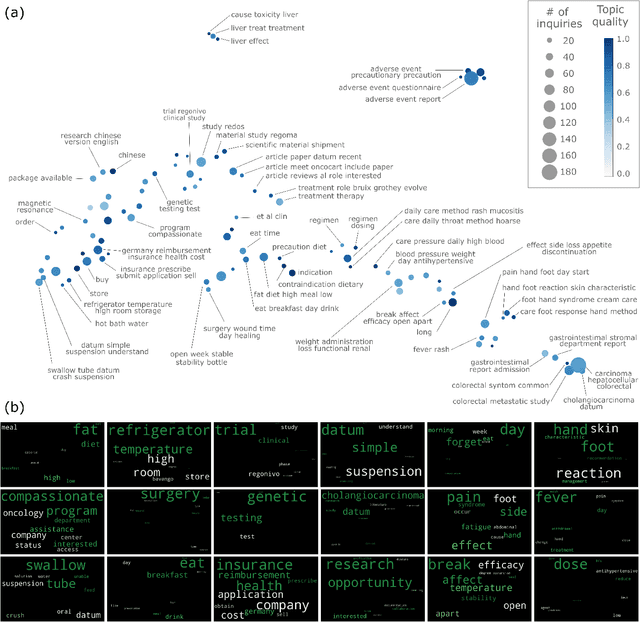
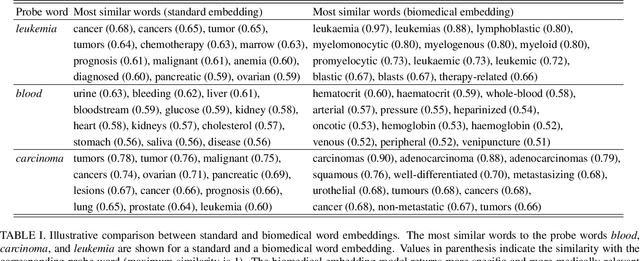
Millions of unsolicited medical inquiries are received by pharmaceutical companies every year. It has been hypothesized that these inquiries represent a treasure trove of information, potentially giving insight into matters regarding medicinal products and the associated medical treatments. However, due to the large volume and specialized nature of the inquiries, it is difficult to perform timely, recurrent, and comprehensive analyses. Here, we propose a machine learning approach based on natural language processing and unsupervised learning to automatically discover key topics in real-world medical inquiries from customers. This approach does not require ontologies nor annotations. The discovered topics are meaningful and medically relevant, as judged by medical information specialists, thus demonstrating that unsolicited medical inquiries are a source of valuable customer insights. Our work paves the way for the machine-learning-driven analysis of medical inquiries in the pharmaceutical industry, which ultimately aims at improving patient care.
Structure-Aware Audio-to-Score Alignment using Progressively Dilated Convolutional Neural Networks
Feb 14, 2021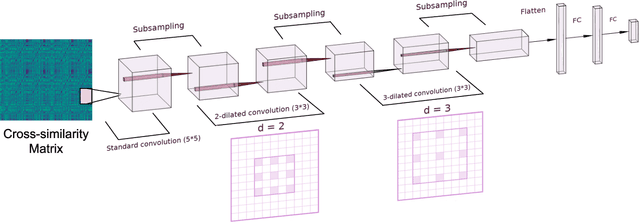
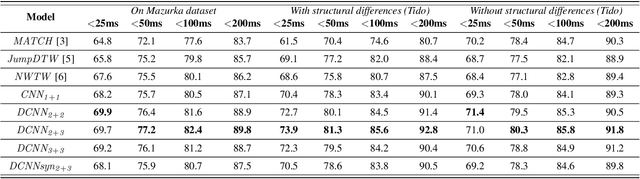
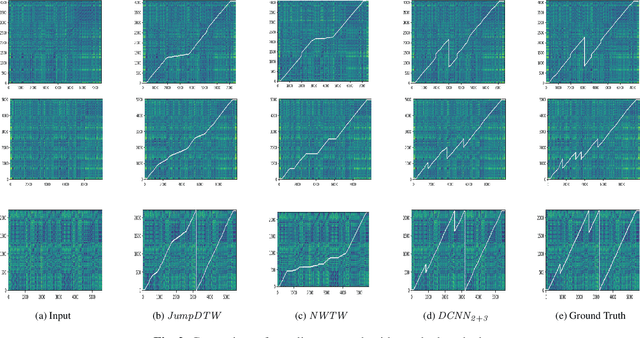
The identification of structural differences between a music performance and the score is a challenging yet integral step of audio-to-score alignment, an important subtask of music information retrieval. We present a novel method to detect such differences between the score and performance for a given piece of music using progressively dilated convolutional neural networks. Our method incorporates varying dilation rates at different layers to capture both short-term and long-term context, and can be employed successfully in the presence of limited annotated data. We conduct experiments on audio recordings of real performances that differ structurally from the score, and our results demonstrate that our models outperform standard methods for structure-aware audio-to-score alignment.
Future-Guided Incremental Transformer for Simultaneous Translation
Dec 23, 2020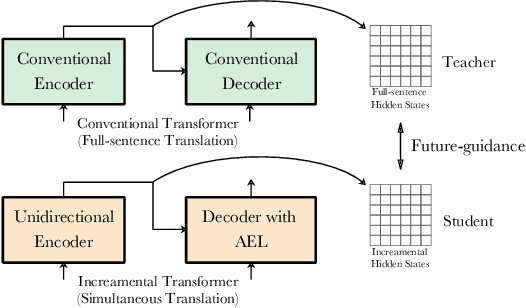
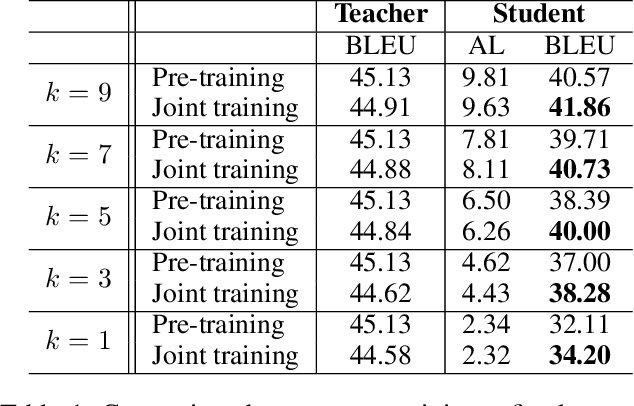
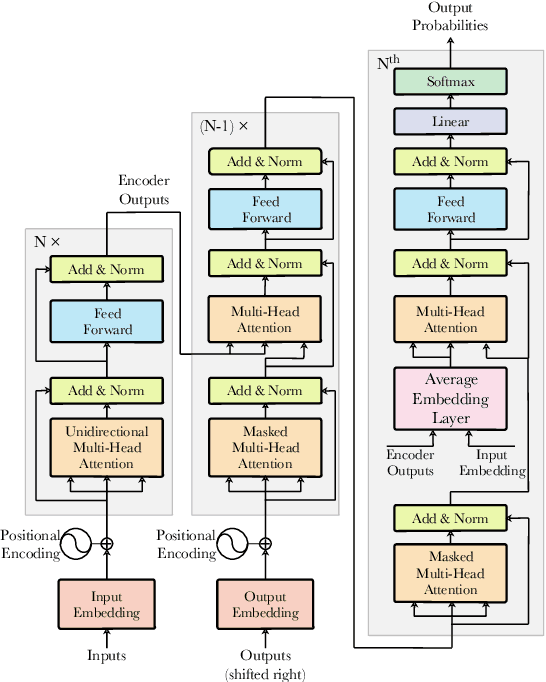
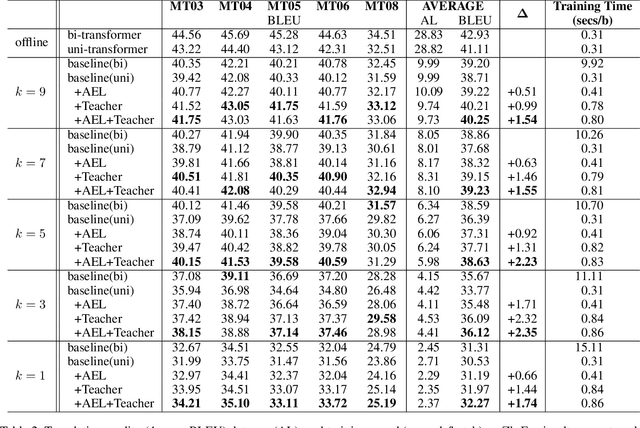
Simultaneous translation (ST) starts translations synchronously while reading source sentences, and is used in many online scenarios. The previous wait-k policy is concise and achieved good results in ST. However, wait-k policy faces two weaknesses: low training speed caused by the recalculation of hidden states and lack of future source information to guide training. For the low training speed, we propose an incremental Transformer with an average embedding layer (AEL) to accelerate the speed of calculation of the hidden states during training. For future-guided training, we propose a conventional Transformer as the teacher of the incremental Transformer, and try to invisibly embed some future information in the model through knowledge distillation. We conducted experiments on Chinese-English and German-English simultaneous translation tasks and compared with the wait-k policy to evaluate the proposed method. Our method can effectively increase the training speed by about 28 times on average at different k and implicitly embed some predictive abilities in the model, achieving better translation quality than wait-k baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge