"Information": models, code, and papers
Measuring Model Fairness under Noisy Covariates: A Theoretical Perspective
May 20, 2021
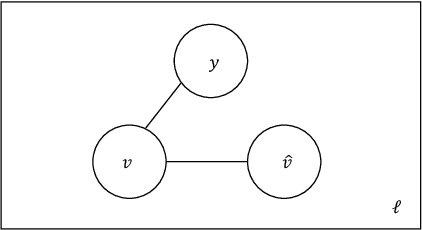
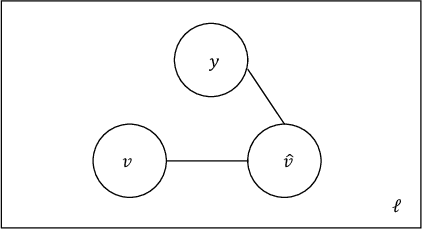
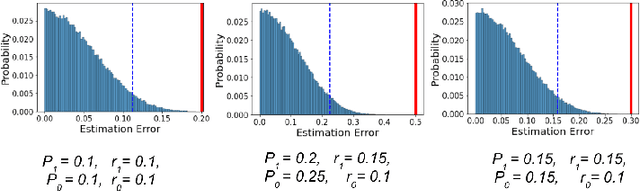
In this work we study the problem of measuring the fairness of a machine learning model under noisy information. Focusing on group fairness metrics, we investigate the particular but common situation when the evaluation requires controlling for the confounding effect of covariate variables. In a practical setting, we might not be able to jointly observe the covariate and group information, and a standard workaround is to then use proxies for one or more of these variables. Prior works have demonstrated the challenges with using a proxy for sensitive attributes, and strong independence assumptions are needed to provide guarantees on the accuracy of the noisy estimates. In contrast, in this work we study using a proxy for the covariate variable and present a theoretical analysis that aims to characterize weaker conditions under which accurate fairness evaluation is possible. Furthermore, our theory identifies potential sources of errors and decouples them into two interpretable parts $\gamma$ and $\epsilon$. The first part $\gamma$ depends solely on the performance of the proxy such as precision and recall, whereas the second part $\epsilon$ captures correlations between all the variables of interest. We show that in many scenarios the error in the estimates is dominated by $\gamma$ via a linear dependence, whereas the dependence on the correlations $\epsilon$ only constitutes a lower order term. As a result we expand the understanding of scenarios where measuring model fairness via proxies can be an effective approach. Finally, we compare, via simulations, the theoretical upper-bounds to the distribution of simulated estimation errors and show that assuming some structure on the data, even weak, is key to significantly improve both theoretical guarantees and empirical results.
Local Disentanglement in Variational Auto-Encoders Using Jacobian $L_1$ Regularization
Jun 05, 2021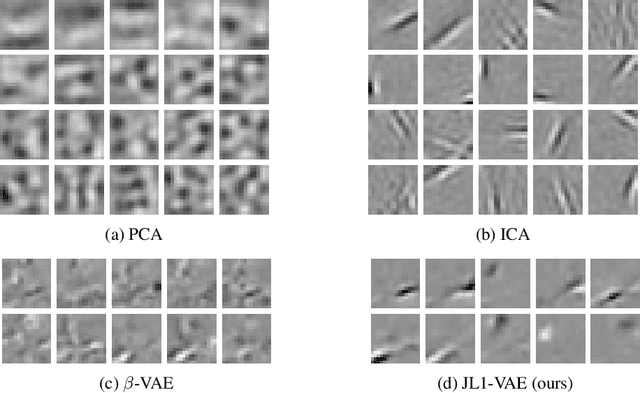
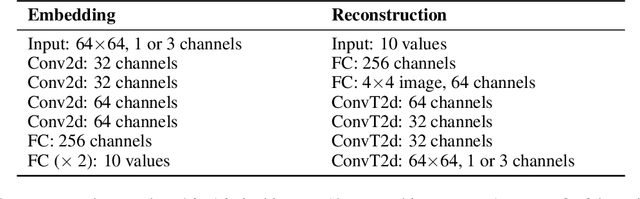
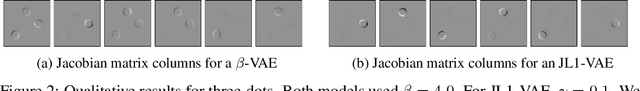
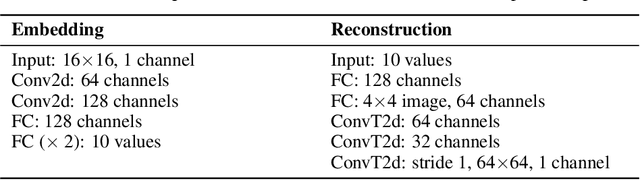



There have been many recent advances in representation learning; however, unsupervised representation learning can still struggle with model identification issues. Variational Auto-Encoders (VAEs) and their extensions such as $\beta$-VAEs have been shown to locally align latent variables with PCA directions, which can help to improve model disentanglement under some conditions. Borrowing inspiration from Independent Component Analysis (ICA) and sparse coding, we propose applying an $L_1$ loss to the VAE's generative Jacobian during training to encourage local latent variable alignment with independent factors of variation in the data. We demonstrate our results on a variety of datasets, giving qualitative and quantitative results using information theoretic and modularity measures that show our added $L_1$ cost encourages local axis alignment of the latent representation with individual factors of variation.
Inferring Black Hole Properties from Astronomical Multivariate Time Series with Bayesian Attentive Neural Processes
Jun 18, 2021
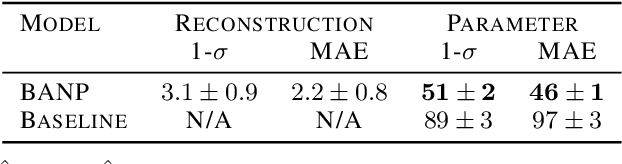
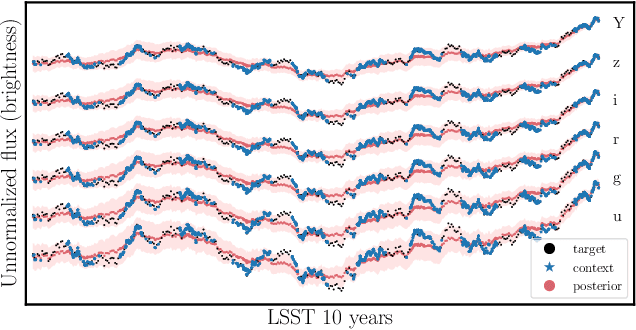
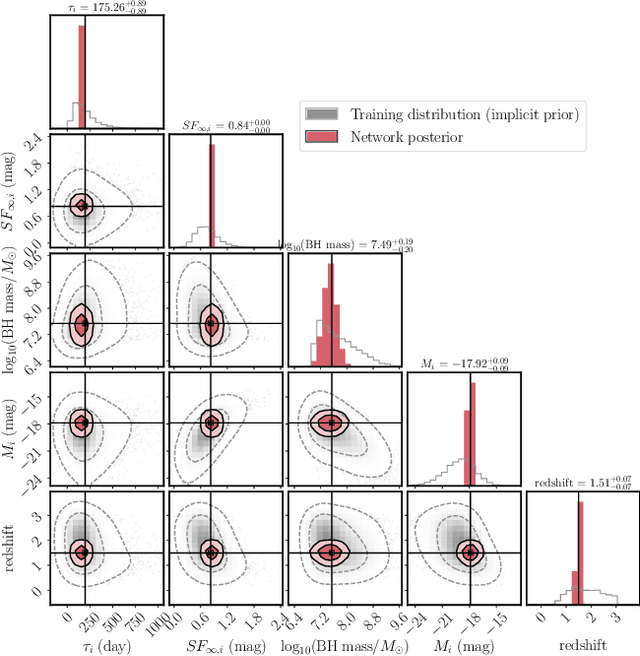
Among the most extreme objects in the Universe, active galactic nuclei (AGN) are luminous centers of galaxies where a black hole feeds on surrounding matter. The variability patterns of the light emitted by an AGN contain information about the physical properties of the underlying black hole. Upcoming telescopes will observe over 100 million AGN in multiple broadband wavelengths, yielding a large sample of multivariate time series with long gaps and irregular sampling. We present a method that reconstructs the AGN time series and simultaneously infers the posterior probability density distribution (PDF) over the physical quantities of the black hole, including its mass and luminosity. We apply this method to a simulated dataset of 11,000 AGN and report precision and accuracy of 0.4 dex and 0.3 dex in the inferred black hole mass. This work is the first to address probabilistic time series reconstruction and parameter inference for AGN in an end-to-end fashion.
Icebreaker: Element-wise Active Information Acquisition with Bayesian Deep Latent Gaussian Model
Aug 14, 2019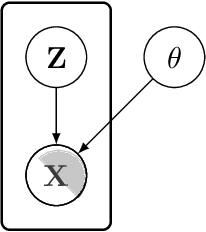
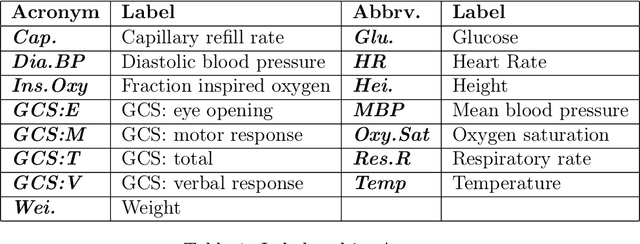
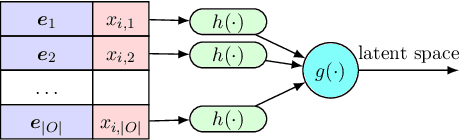
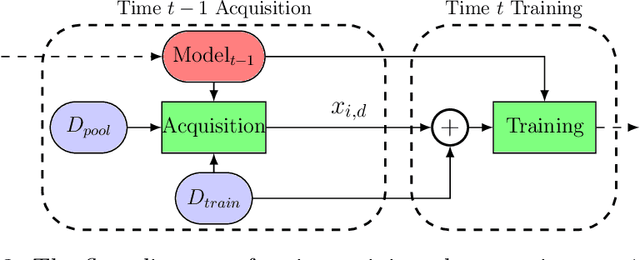
In this paper we introduce the ice-start problem, i.e., the challenge of deploying machine learning models when only little or no training data is initially available, and acquiring each feature element of data is associated with costs. This setting is representative for the real-world machine learning applications. For instance, in the health-care domain, when training an AI system for predicting patient metrics from lab tests, obtaining every single measurement comes with a high cost. Active learning, where only the label is associated with a cost does not apply to such problem, because performing all possible lab tests to acquire a new training datum would be costly, as well as unnecessary due to redundancy. We propose Icebreaker, a principled framework to approach the ice-start problem. Icebreaker uses a full Bayesian Deep Latent Gaussian Model (BELGAM) with a novel inference method. Our proposed method combines recent advances in amortized inference and stochastic gradient MCMC to enable fast and accurate posterior inference. By utilizing BELGAM's ability to fully quantify model uncertainty, we also propose two information acquisition functions for imputation and active prediction problems. We demonstrate that BELGAM performs significantly better than the previous VAE (Variational autoencoder) based models, when the data set size is small, using both machine learning benchmarks and real-world recommender systems and health-care applications. Moreover, based on BELGAM, Icebreaker further improves the performance and demonstrate the ability to use minimum amount of the training data to obtain the highest test time performance.
End-to-end Temporal Action Detection with Transformer
Jun 18, 2021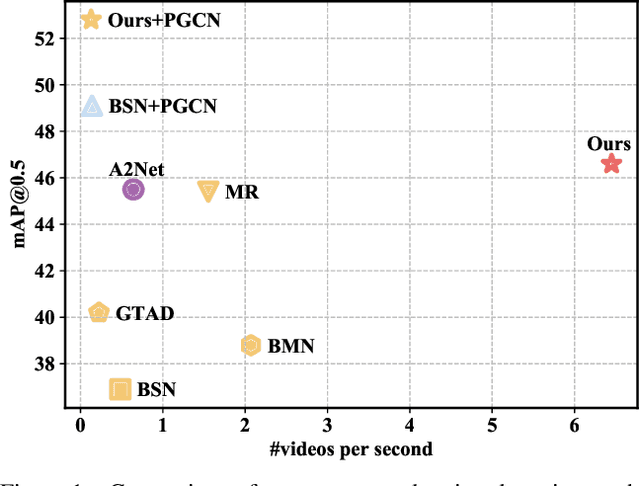
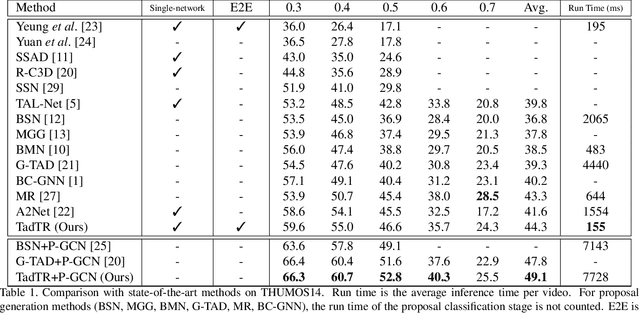
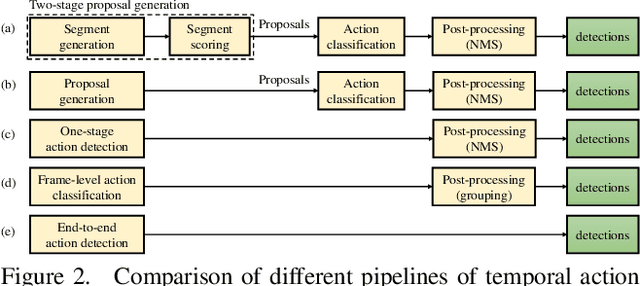
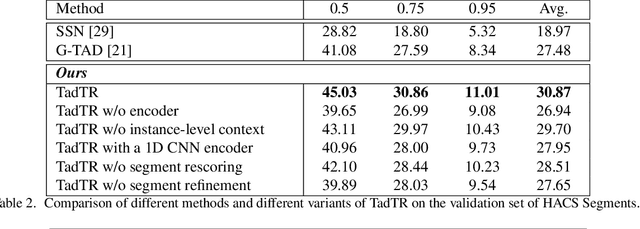
Temporal action detection (TAD) aims to determine the semantic label and the boundaries of every action instance in an untrimmed video. It is a fundamental task in video understanding and significant progress has been made in TAD. Previous methods involve multiple stages or networks and hand-designed rules or operations, which fall short in efficiency and flexibility. Here, we construct an end-to-end framework for TAD upon Transformer, termed \textit{TadTR}, which simultaneously predicts all action instances as a set of labels and temporal locations in parallel. TadTR is able to adaptively extract temporal context information needed for making action predictions, by selectively attending to a number of snippets in a video. It greatly simplifies the pipeline of TAD and runs much faster than previous detectors. Our method achieves state-of-the-art performance on HACS Segments and THUMOS14 and competitive performance on ActivityNet-1.3. Our code will be made available at \url{https://github.com/xlliu7/TadTR}.
Interpretable Drug Synergy Prediction with Graph Neural Networks for Human-AI Collaboration in Healthcare
May 14, 2021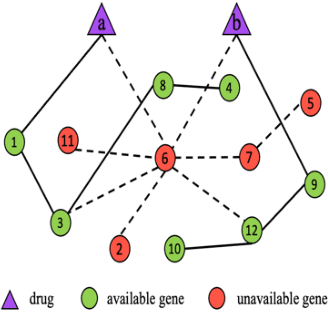
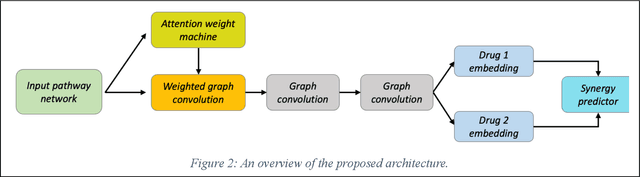
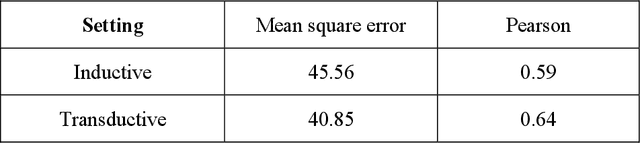
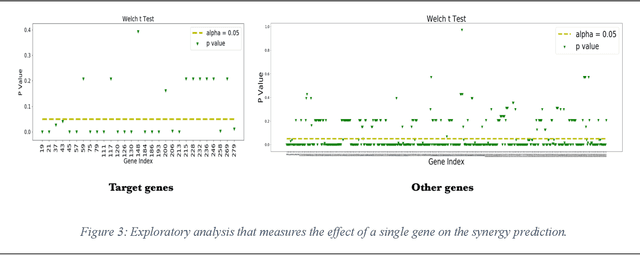
We investigate molecular mechanisms of resistant or sensitive response of cancer drug combination therapies in an inductive and interpretable manner. Though deep learning algorithms are widely used in the drug synergy prediction problem, it is still an open problem to formulate the prediction model with biological meaning to investigate the mysterious mechanisms of synergy (MoS) for the human-AI collaboration in healthcare systems. To address the challenges, we propose a deep graph neural network, IDSP (Interpretable Deep Signaling Pathways), to incorporate the gene-gene as well as gene-drug regulatory relationships in synergic drug combination predictions. IDSP automatically learns weights of edges based on the gene and drug node relations, i.e., signaling interactions, by a multi-layer perceptron (MLP) and aggregates information in an inductive manner. The proposed architecture generates interpretable drug synergy prediction by detecting important signaling interactions, and can be implemented when the underlying molecular mechanism encounters unseen genes or signaling pathways. We test IDWSP on signaling networks formulated by genes from 46 core cancer signaling pathways and drug combinations from NCI ALMANAC drug combination screening data. The experimental results demonstrated that 1) IDSP can learn from the underlying molecular mechanism to make prediction without additional drug chemical information while achieving highly comparable performance with current state-of-art methods; 2) IDSP show superior generality and flexibility to implement the synergy prediction task on both transductive tasks and inductive tasks. 3) IDSP can generate interpretable results by detecting different salient signaling patterns (i.e. MoS) for different cell lines.
Surrogate Scoring Rules and a Dominant Truth Serum for Information Elicitation
May 04, 2018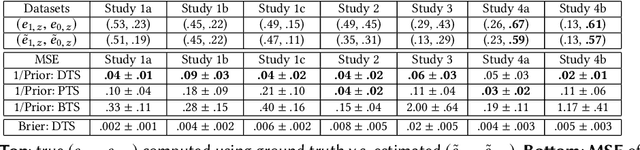
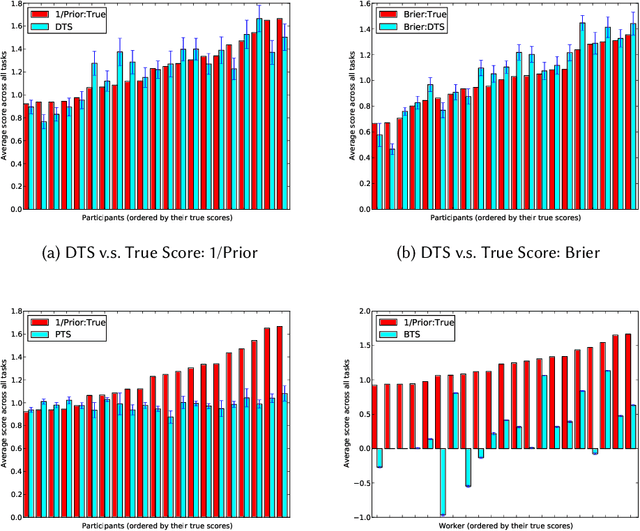
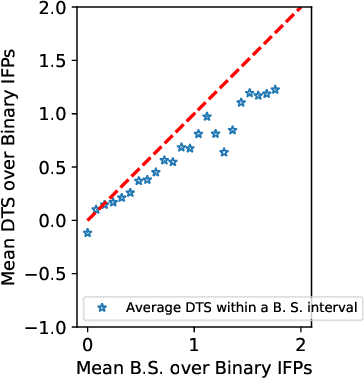
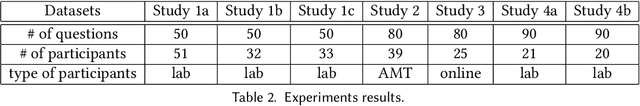
We study information elicitation without verification (IEWV) and ask the following question: Can we achieve truthfulness in dominant strategy in IEWV? This paper considers two elicitation settings. The first setting is when the mechanism designer has access to a random variable that is a noisy or proxy version of the ground truth, with known biases. The second setting is the standard peer prediction setting where agents' reports are the only source of information that the mechanism designer has. We introduce surrogate scoring rules (SSR) for the first setting, which use the noisy ground truth to evaluate quality of elicited information, and show that SSR achieve truthful elicitation in dominant strategy. Built upon SSR, we develop a multi-task mechanism, dominant truth serum (DTS), to achieve truthful elicitation in dominant strategy when the mechanism designer only has access to agents' reports (the second setting). The method relies on an estimation procedure to accurately estimate the average bias in the reports of other agents. With the accurate estimation, a random peer agent's report serves as a noisy ground truth and SSR can then be applied to achieve truthfulness in dominant strategy. A salient feature of SSR and DTS is that they both quantify the quality or value of information despite lack of ground truth, just as proper scoring rules do for the with verification setting. Our work complements both the strictly proper scoring rule literature by solving the case where the mechanism designer only has access to a noisy or proxy version of the ground truth, and the peer prediction literature by achieving truthful elicitation in dominant strategy.
Machine Learning Characterization of Cancer Patients-Derived Extracellular Vesicles using Vibrational Spectroscopies
Jul 21, 2021
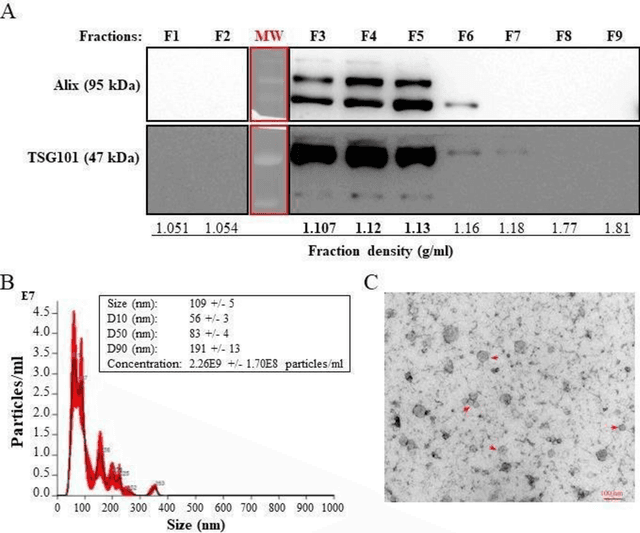
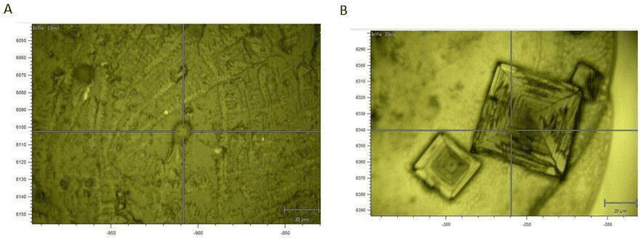
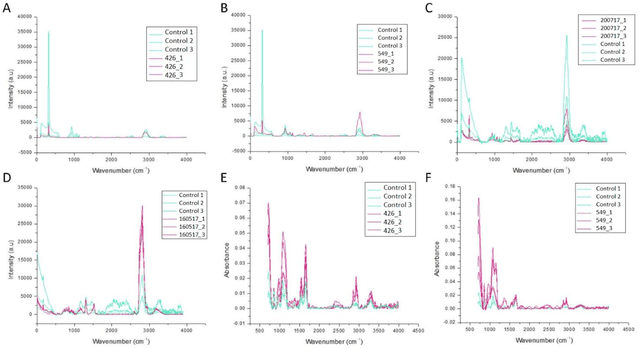
The early detection of cancer is a challenging problem in medicine. The blood sera of cancer patients are enriched with heterogeneous secretory lipid bound extracellular vesicles (EVs), which present a complex repertoire of information and biomarkers, representing their cell of origin, that are being currently studied in the field of liquid biopsy and cancer screening. Vibrational spectroscopies provide non-invasive approaches for the assessment of structural and biophysical properties in complex biological samples. In this study, multiple Raman spectroscopy measurements were performed on the EVs extracted from the blood sera of 9 patients consisting of four different cancer subtypes (colorectal cancer, hepatocellular carcinoma, breast cancer and pancreatic cancer) and five healthy patients (controls). FTIR(Fourier Transform Infrared) spectroscopy measurements were performed as a complementary approach to Raman analysis, on two of the four cancer subtypes. The AdaBoost Random Forest Classifier, Decision Trees, and Support Vector Machines (SVM) distinguished the baseline corrected Raman spectra of cancer EVs from those of healthy controls (18 spectra) with a classification accuracy of greater than 90% when reduced to a spectral frequency range of 1800 to 1940 inverse cm, and subjected to a 0.5 training/testing split. FTIR classification accuracy on 14 spectra showed an 80% classification accuracy. Our findings demonstrate that basic machine learning algorithms are powerful tools to distinguish the complex vibrational spectra of cancer patient EVs from those of healthy patients. These experimental methods hold promise as valid and efficient liquid biopsy for machine intelligence-assisted early cancer screening.
Deep Learning Through the Lens of Example Difficulty
Jun 18, 2021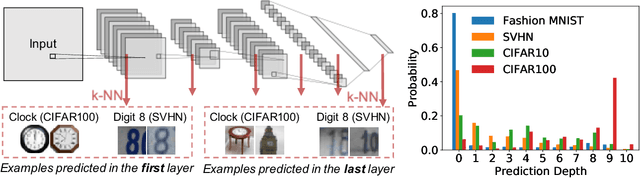
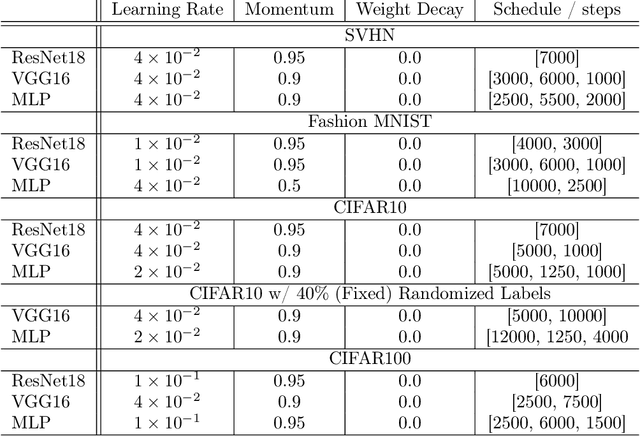
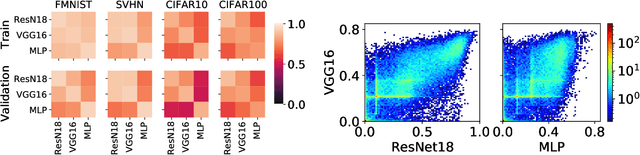

Existing work on understanding deep learning often employs measures that compress all data-dependent information into a few numbers. In this work, we adopt a perspective based on the role of individual examples. We introduce a measure of the computational difficulty of making a prediction for a given input: the (effective) prediction depth. Our extensive investigation reveals surprising yet simple relationships between the prediction depth of a given input and the model's uncertainty, confidence, accuracy and speed of learning for that data point. We further categorize difficult examples into three interpretable groups, demonstrate how these groups are processed differently inside deep models and showcase how this understanding allows us to improve prediction accuracy. Insights from our study lead to a coherent view of a number of separately reported phenomena in the literature: early layers generalize while later layers memorize; early layers converge faster and networks learn easy data and simple functions first.
Language Grounding with 3D Objects
Jul 26, 2021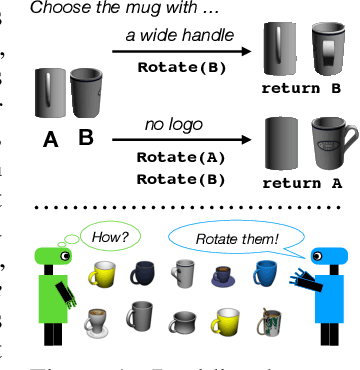
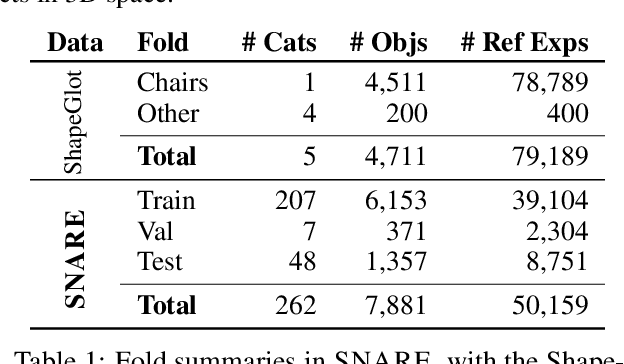
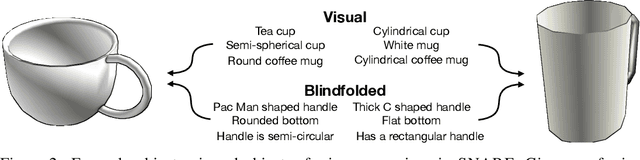
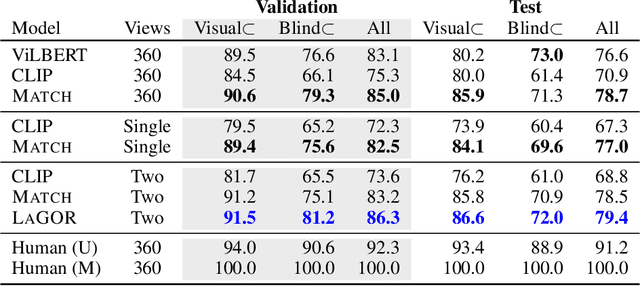
Seemingly simple natural language requests to a robot are generally underspecified, for example "Can you bring me the wireless mouse?" When viewing mice on the shelf, the number of buttons or presence of a wire may not be visible from certain angles or positions. Flat images of candidate mice may not provide the discriminative information needed for "wireless". The world, and objects in it, are not flat images but complex 3D shapes. If a human requests an object based on any of its basic properties, such as color, shape, or texture, robots should perform the necessary exploration to accomplish the task. In particular, while substantial effort and progress has been made on understanding explicitly visual attributes like color and category, comparatively little progress has been made on understanding language about shapes and contours. In this work, we introduce a novel reasoning task that targets both visual and non-visual language about 3D objects. Our new benchmark, ShapeNet Annotated with Referring Expressions (SNARE), requires a model to choose which of two objects is being referenced by a natural language description. We introduce several CLIP-based models for distinguishing objects and demonstrate that while recent advances in jointly modeling vision and language are useful for robotic language understanding, it is still the case that these models are weaker at understanding the 3D nature of objects -- properties which play a key role in manipulation. In particular, we find that adding view estimation to language grounding models improves accuracy on both SNARE and when identifying objects referred to in language on a robot platform.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge