"Information": models, code, and papers
A cautionary tale on fitting decision trees to data from additive models: generalization lower bounds
Oct 18, 2021
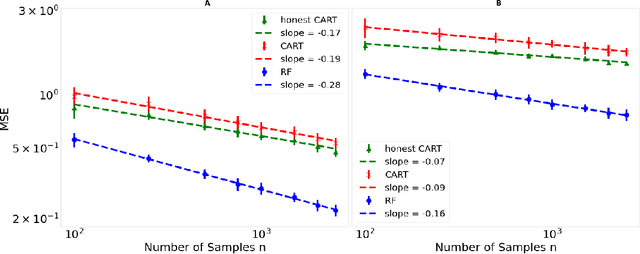
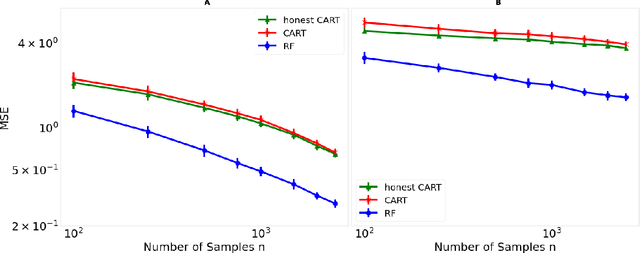
Decision trees are important both as interpretable models amenable to high-stakes decision-making, and as building blocks of ensemble methods such as random forests and gradient boosting. Their statistical properties, however, are not well understood. The most cited prior works have focused on deriving pointwise consistency guarantees for CART in a classical nonparametric regression setting. We take a different approach, and advocate studying the generalization performance of decision trees with respect to different generative regression models. This allows us to elicit their inductive bias, that is, the assumptions the algorithms make (or do not make) to generalize to new data, thereby guiding practitioners on when and how to apply these methods. In this paper, we focus on sparse additive generative models, which have both low statistical complexity and some nonparametric flexibility. We prove a sharp squared error generalization lower bound for a large class of decision tree algorithms fitted to sparse additive models with $C^1$ component functions. This bound is surprisingly much worse than the minimax rate for estimating such sparse additive models. The inefficiency is due not to greediness, but to the loss in power for detecting global structure when we average responses solely over each leaf, an observation that suggests opportunities to improve tree-based algorithms, for example, by hierarchical shrinkage. To prove these bounds, we develop new technical machinery, establishing a novel connection between decision tree estimation and rate-distortion theory, a sub-field of information theory.
EPIC-KITCHENS-100 Unsupervised Domain Adaptation Challenge for Action Recognition 2021: Team M3EM Technical Report
Jun 27, 2021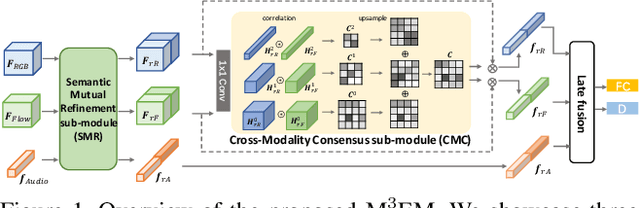

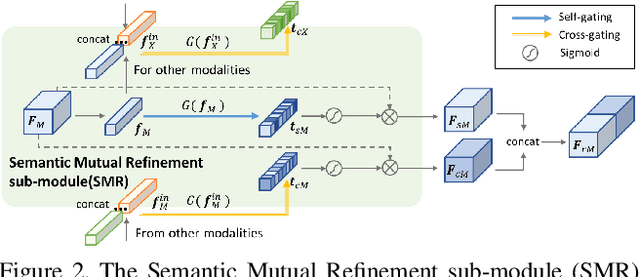
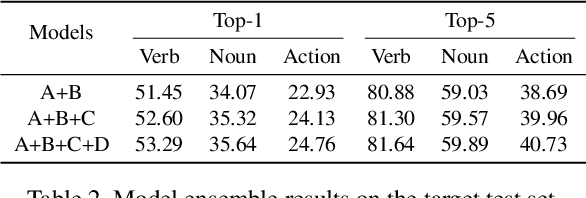
In this report, we describe the technical details of our submission to the 2021 EPIC-KITCHENS-100 Unsupervised Domain Adaptation Challenge for Action Recognition. Leveraging multiple modalities has been proved to benefit the Unsupervised Domain Adaptation (UDA) task. In this work, we present Multi-Modal Mutual Enhancement Module (M3EM), a deep module for jointly considering information from multiple modalities to find the most transferable representations across domains. We achieve this by implementing two sub-modules for enhancing each modality using the context of other modalities. The first sub-module exchanges information across modalities through the semantic space, while the second sub-module finds the most transferable spatial region based on the consensus of all modalities.
Neural Link Prediction with Walk Pooling
Oct 08, 2021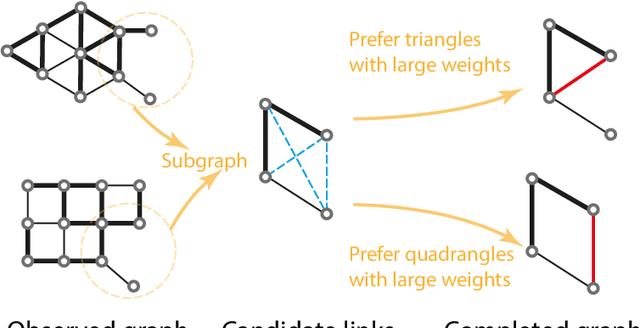
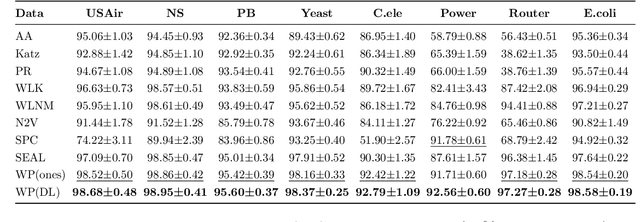
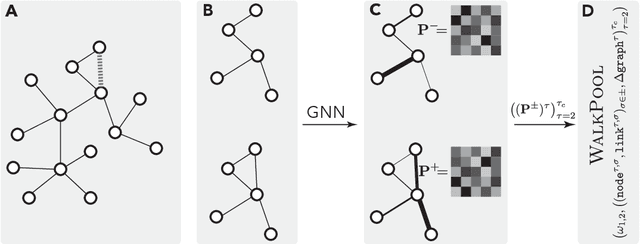
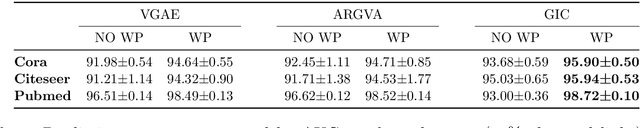
Graph neural networks achieve high accuracy in link prediction by jointly leveraging graph topology and node attributes. Topology, however, is represented indirectly; state-of-the-art methods based on subgraph classification label nodes with distance to the target link, so that, although topological information is present, it is tempered by pooling. This makes it challenging to leverage features like loops and motifs associated with network formation mechanisms. We propose a link prediction algorithm based on a new pooling scheme called WalkPool. WalkPool combines the expressivity of topological heuristics with the feature-learning ability of neural networks. It summarizes a putative link by random walk probabilities of adjacent paths. Instead of extracting transition probabilities from the original graph, it computes the transition matrix of a "predictive" latent graph by applying attention to learned features; this may be interpreted as feature-sensitive topology fingerprinting. WalkPool can leverage unsupervised node features or be combined with GNNs and trained end-to-end. It outperforms state-of-the-art methods on all common link prediction benchmarks, both homophilic and heterophilic, with and without node attributes. Applying WalkPool to a set of unsupervised GNNs significantly improves prediction accuracy, suggesting that it may be used as a general-purpose graph pooling scheme.
Trusted-Maximizers Entropy Search for Efficient Bayesian Optimization
Jul 30, 2021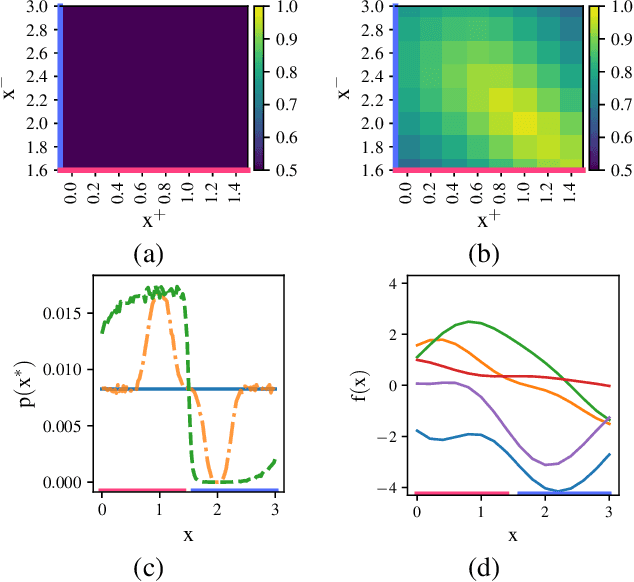
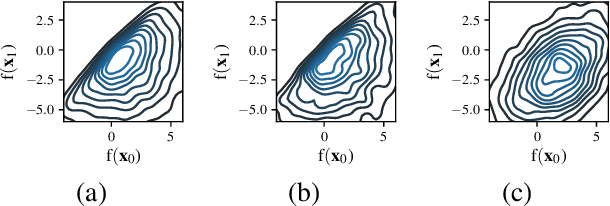
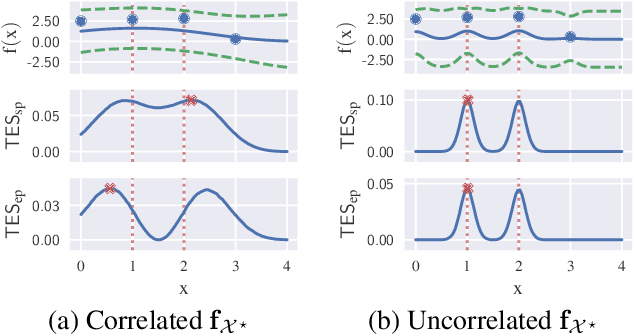
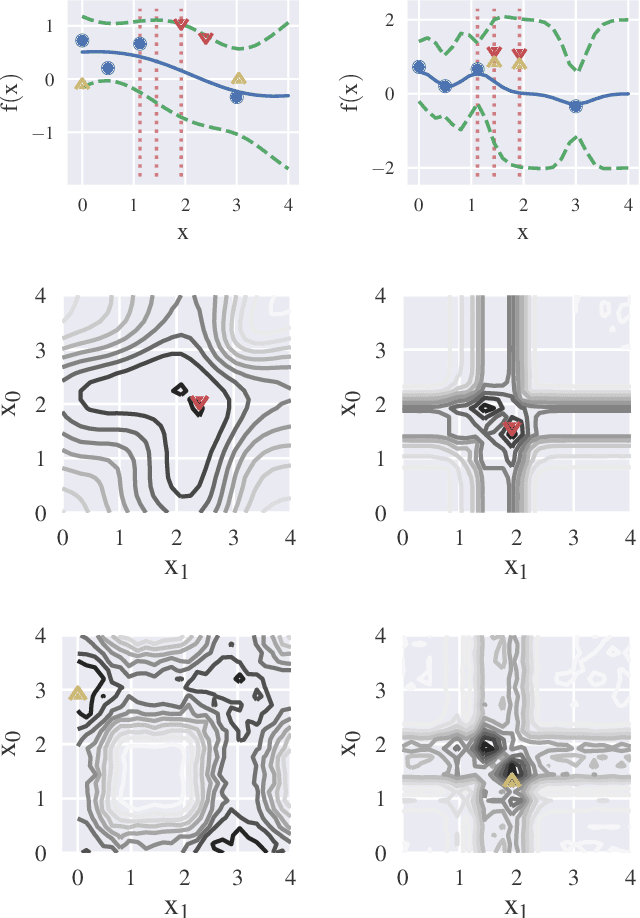
Information-based Bayesian optimization (BO) algorithms have achieved state-of-the-art performance in optimizing a black-box objective function. However, they usually require several approximations or simplifying assumptions (without clearly understanding their effects on the BO performance) and/or their generalization to batch BO is computationally unwieldy, especially with an increasing batch size. To alleviate these issues, this paper presents a novel trusted-maximizers entropy search (TES) acquisition function: It measures how much an input query contributes to the information gain on the maximizer over a finite set of trusted maximizers, i.e., inputs optimizing functions that are sampled from the Gaussian process posterior belief of the objective function. Evaluating TES requires either only a stochastic approximation with sampling or a deterministic approximation with expectation propagation, both of which are investigated and empirically evaluated using synthetic benchmark objective functions and real-world optimization problems, e.g., hyperparameter tuning of a convolutional neural network and synthesizing 'physically realizable' faces to fool a black-box face recognition system. Though TES can naturally be generalized to a batch variant with either approximation, the latter is amenable to be scaled to a much larger batch size in our experiments.
Improving GNSS Positioning using Neural Network-based Corrections
Oct 18, 2021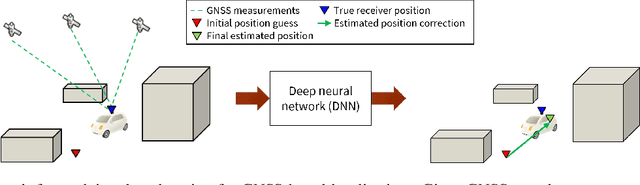
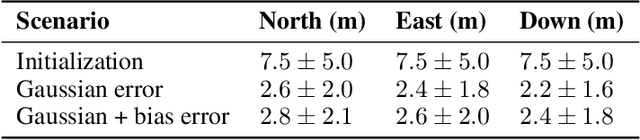

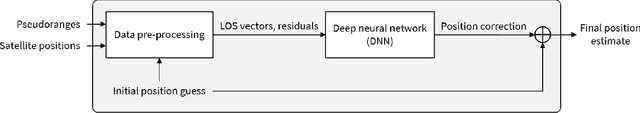

Deep Neural Networks (DNNs) are a promising tool for Global Navigation Satellite System (GNSS) positioning in the presence of multipath and non-line-of-sight errors, owing to their ability to model complex errors using data. However, developing a DNN for GNSS positioning presents various challenges, such as 1) poor numerical conditioning caused by large variations in measurements and position values across the globe, 2) varying number and order within the set of measurements due to changing satellite visibility, and 3) overfitting to available data. In this work, we address the aforementioned challenges and propose an approach for GNSS positioning by applying DNN-based corrections to an initial position guess. Our DNN learns to output the position correction using the set of pseudorange residuals and satellite line-of-sight vectors as inputs. The limited variation in these input and output values improves the numerical conditioning for our DNN. We design our DNN architecture to combine information from the available GNSS measurements, which vary both in number and order, by leveraging recent advancements in set-based deep learning methods. Furthermore, we present a data augmentation strategy for reducing overfitting in the DNN by randomizing the initial position guesses. We first perform simulations and show an improvement in the initial positioning error when our DNN-based corrections are applied. After this, we demonstrate that our approach outperforms a WLS baseline on real-world data. Our implementation is available at github.com/Stanford-NavLab/deep_gnss.
Local and Global Context-Based Pairwise Models for Sentence Ordering
Oct 08, 2021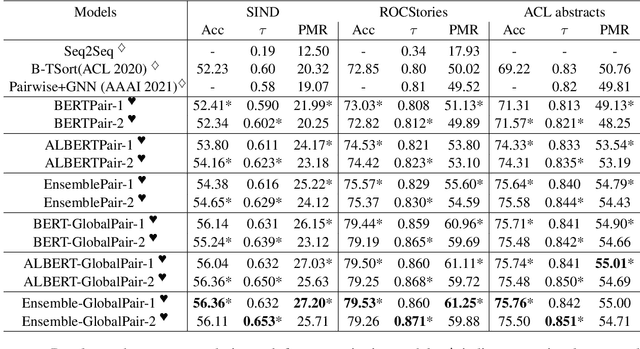
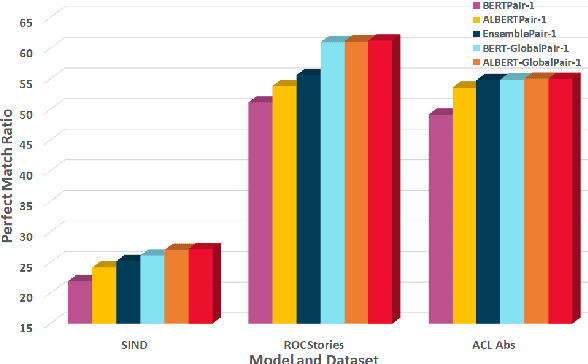
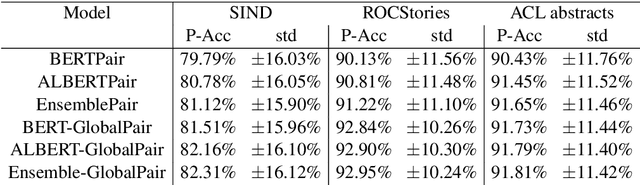
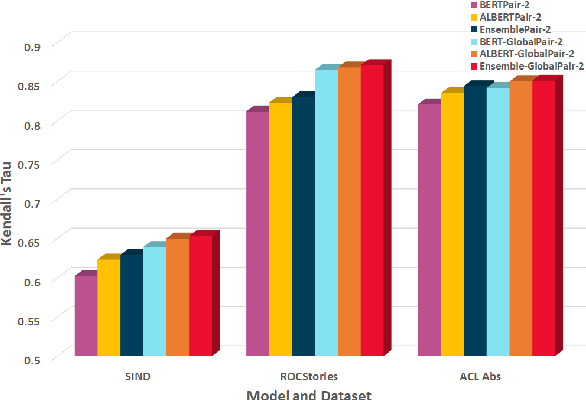
Sentence Ordering refers to the task of rearranging a set of sentences into the appropriate coherent order. For this task, most previous approaches have explored global context-based end-to-end methods using Sequence Generation techniques. In this paper, we put forward a set of robust local and global context-based pairwise ordering strategies, leveraging which our prediction strategies outperform all previous works in this domain. Our proposed encoding method utilizes the paragraph's rich global contextual information to predict the pairwise order using novel transformer architectures. Analysis of the two proposed decoding strategies helps better explain error propagation in pairwise models. This approach is the most accurate pure pairwise model and our encoding strategy also significantly improves the performance of other recent approaches that use pairwise models, including the previous state-of-the-art, demonstrating the research novelty and generalizability of this work. Additionally, we show how the pre-training task for ALBERT helps it to significantly outperform BERT, despite having considerably lesser parameters. The extensive experimental results, architectural analysis and ablation studies demonstrate the effectiveness and superiority of the proposed models compared to the previous state-of-the-art, besides providing a much better understanding of the functioning of pairwise models.
Structure-Aware Feature Generation for Zero-Shot Learning
Aug 16, 2021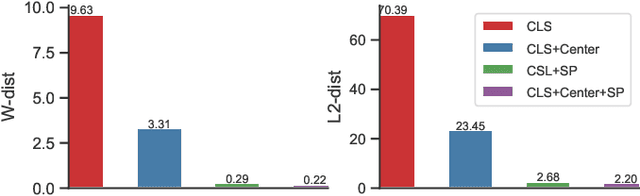
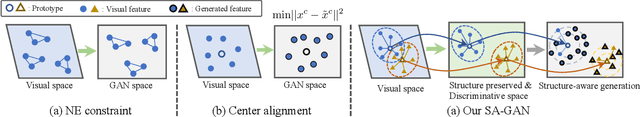
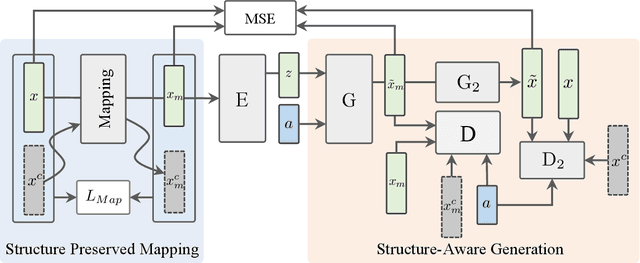
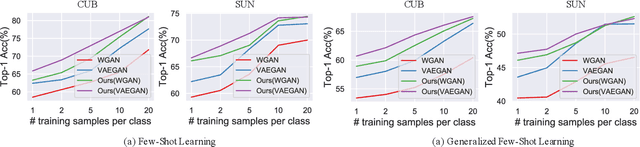
Zero-Shot Learning (ZSL) targets at recognizing unseen categories by leveraging auxiliary information, such as attribute embedding. Despite the encouraging results achieved, prior ZSL approaches focus on improving the discriminant power of seen-class features, yet have largely overlooked the geometric structure of the samples and the prototypes. The subsequent attribute-based generative adversarial network (GAN), as a result, also neglects the topological information in sample generation and further yields inferior performances in classifying the visual features of unseen classes. In this paper, we introduce a novel structure-aware feature generation scheme, termed as SA-GAN, to explicitly account for the topological structure in learning both the latent space and the generative networks. Specifically, we introduce a constraint loss to preserve the initial geometric structure when learning a discriminative latent space, and carry out our GAN training with additional supervising signals from a structure-aware discriminator and a reconstruction module. The former supervision distinguishes fake and real samples based on their affinity to class prototypes, while the latter aims to reconstruct the original feature space from the generated latent space. This topology-preserving mechanism enables our method to significantly enhance the generalization capability on unseen-classes and consequently improve the classification performance. Experiments on four benchmarks demonstrate that the proposed approach consistently outperforms the state of the art. Our code can be found in the supplementary material and will also be made publicly available.
RadGraph: Extracting Clinical Entities and Relations from Radiology Reports
Jun 28, 2021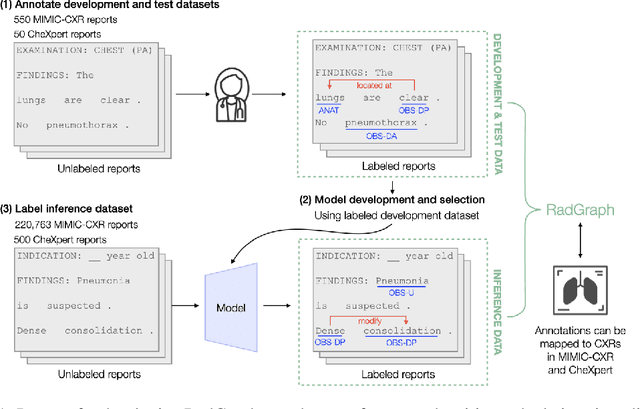
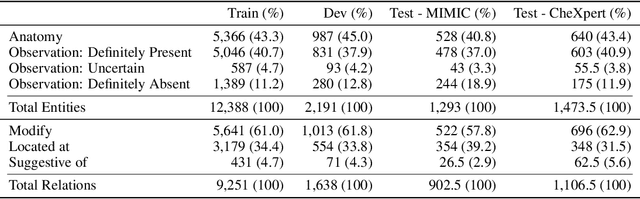
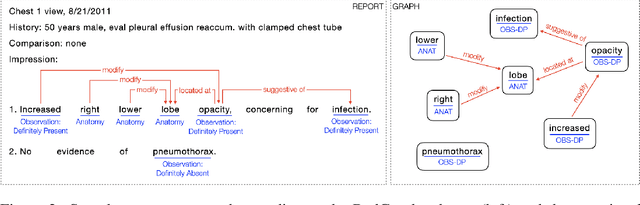
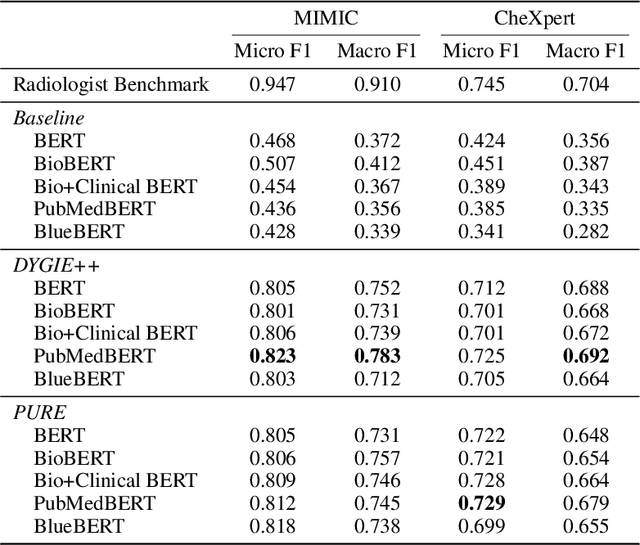
Extracting structured clinical information from free-text radiology reports can enable the use of radiology report information for a variety of critical healthcare applications. In our work, we present RadGraph, a dataset of entities and relations in full-text chest X-ray radiology reports based on a novel information extraction schema we designed to structure radiology reports. We release a development dataset, which contains board-certified radiologist annotations for 500 radiology reports from the MIMIC-CXR dataset (14,579 entities and 10,889 relations), and a test dataset, which contains two independent sets of board-certified radiologist annotations for 100 radiology reports split equally across the MIMIC-CXR and CheXpert datasets. Using these datasets, we train and test a deep learning model, RadGraph Benchmark, that achieves a micro F1 of 0.82 and 0.73 on relation extraction on the MIMIC-CXR and CheXpert test sets respectively. Additionally, we release an inference dataset, which contains annotations automatically generated by RadGraph Benchmark across 220,763 MIMIC-CXR reports (around 6 million entities and 4 million relations) and 500 CheXpert reports (13,783 entities and 9,908 relations) with mappings to associated chest radiographs. Our freely available dataset can facilitate a wide range of research in medical natural language processing, as well as computer vision and multi-modal learning when linked to chest radiographs.
CCMI : Classifier based Conditional Mutual Information Estimation
Jun 05, 2019
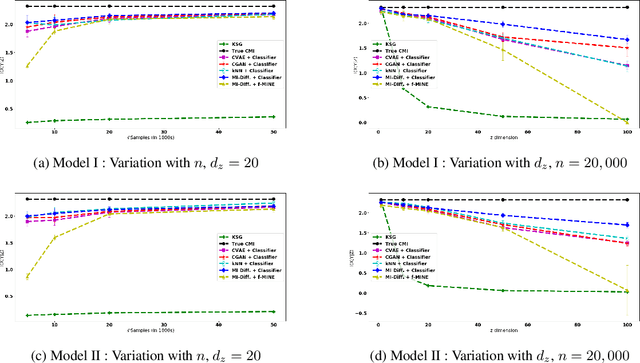
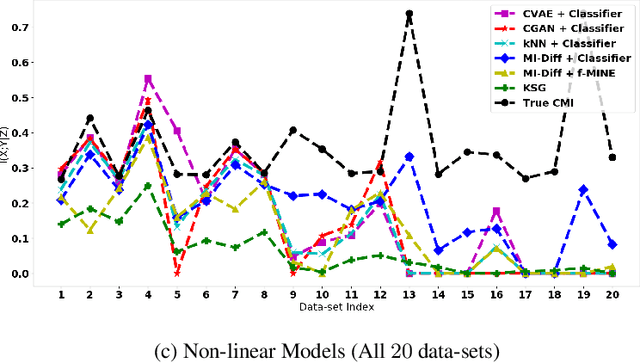
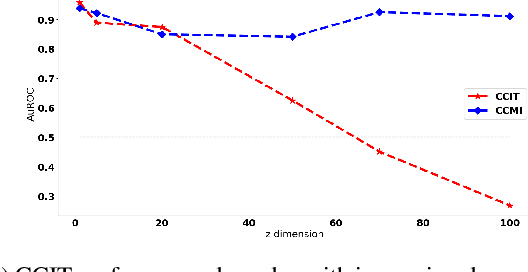
Conditional Mutual Information (CMI) is a measure of conditional dependence between random variables X and Y, given another random variable Z. It can be used to quantify conditional dependence among variables in many data-driven inference problems such as graphical models, causal learning, feature selection and time-series analysis. While k-nearest neighbor (kNN) based estimators as well as kernel-based methods have been widely used for CMI estimation, they suffer severely from the curse of dimensionality. In this paper, we leverage advances in classifiers and generative models to design methods for CMI estimation. Specifically, we introduce an estimator for KL-Divergence based on the likelihood ratio by training a classifier to distinguish the observed joint distribution from the product distribution. We then show how to construct several CMI estimators using this basic divergence estimator by drawing ideas from conditional generative models. We demonstrate that the estimates from our proposed approaches do not degrade in performance with increasing dimension and obtain significant improvement over the widely used KSG estimator. Finally, as an application of accurate CMI estimation, we use our best estimator for conditional independence testing and achieve superior performance than the state-of-the-art tester on both simulated and real data-sets.
Eigenbehaviour as an Indicator of Cognitive Abilities
Oct 18, 2021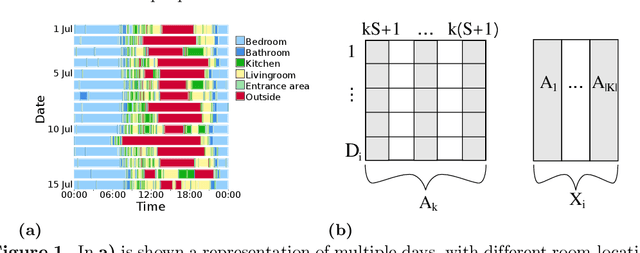

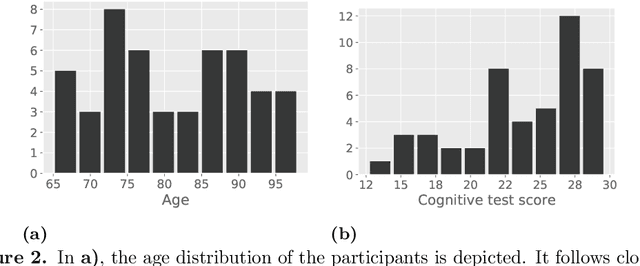
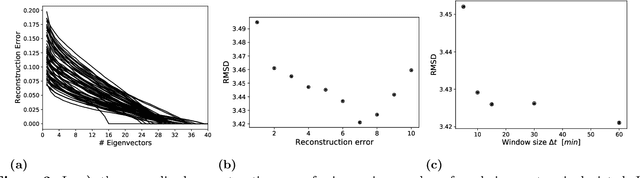
With growing usage of machine learning algorithms and big data in health applications, digital biomarkers have become an important key feature to ensure the success of those applications. In this paper, we focus on one important use-case, the long-term continuous monitoring of the cognitive ability of older adults. The cognitive ability is a factor both for long-term monitoring of people living alone as well as an outcome in clinical studies. In this work, we propose a new digital biomarker for cognitive abilities based on location eigenbehaviour obtained from contactless ambient sensors. Indoor location information obtained from passive infrared sensors is used to build a location matrix covering several weeks of measurement. Based on the eigenvectors of this matrix, the reconstruction error is calculated for various numbers of used eigenvectors. The reconstruction error is used to predict cognitive ability scores collected at baseline, using linear regression. Additionally, classification of normal versus pathological cognition level is performed using a support-vector-machine. Prediction performance is strong for high levels of cognitive ability, but grows weaker for low levels of cognitive ability. Classification into normal versus pathological cognitive ability level reaches high accuracy with a AUC = 0.94. Due to the unobtrusive method of measurement based on contactless ambient sensors, this digital biomarker of cognitive ability is easily obtainable. The usage of the reconstruction error is a strong digital biomarker for the binary classification and, to a lesser extent, for more detailed prediction of interindividual differences in cognition.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge