"Information": models, code, and papers
Enhancing Hyperbolic Graph Embeddings via Contrastive Learning
Jan 21, 2022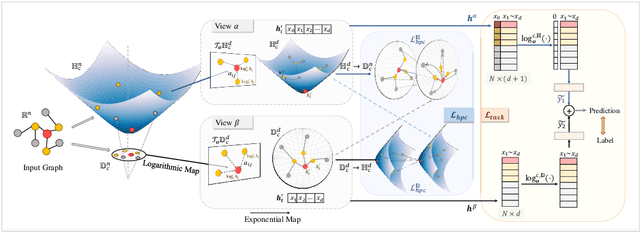
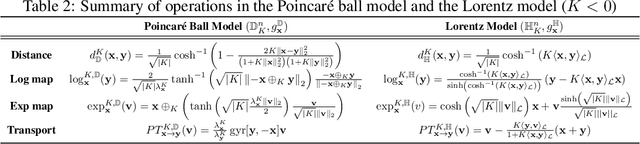
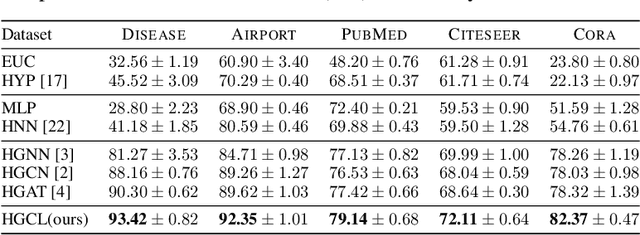
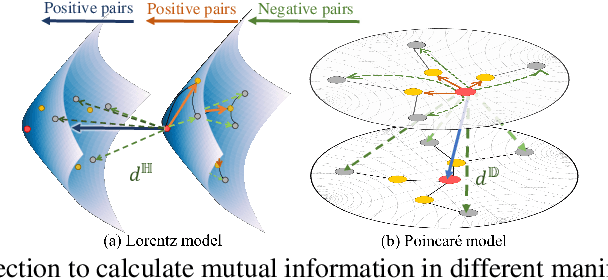

Recently, hyperbolic space has risen as a promising alternative for semi-supervised graph representation learning. Many efforts have been made to design hyperbolic versions of neural network operations. However, the inspiring geometric properties of this unique geometry have not been fully explored yet. The potency of graph models powered by the hyperbolic space is still largely underestimated. Besides, the rich information carried by abundant unlabelled samples is also not well utilized. Inspired by the recently active and emerging self-supervised learning, in this study, we attempt to enhance the representation power of hyperbolic graph models by drawing upon the advantages of contrastive learning. More specifically, we put forward a novel Hyperbolic Graph Contrastive Learning (HGCL) framework which learns node representations through multiple hyperbolic spaces to implicitly capture the hierarchical structure shared between different views. Then, we design a hyperbolic position consistency (HPC) constraint based on hyperbolic distance and the homophily assumption to make contrastive learning fit into hyperbolic space. Experimental results on multiple real-world datasets demonstrate the superiority of the proposed HGCL as it consistently outperforms competing methods by considerable margins for the node classification task.
Order Matters: Matching Multiple Knowledge Graphs
Nov 03, 2021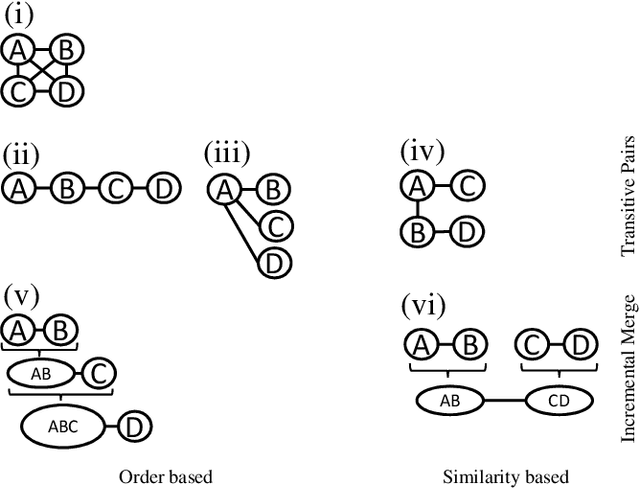
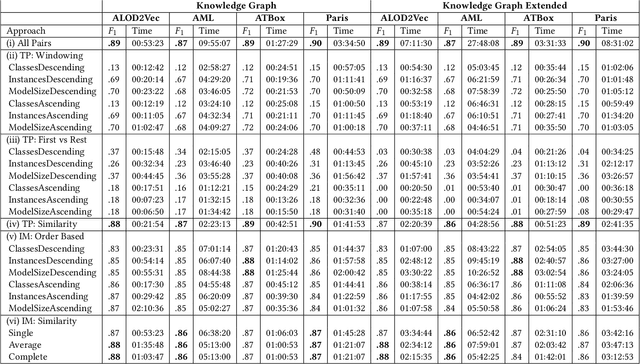
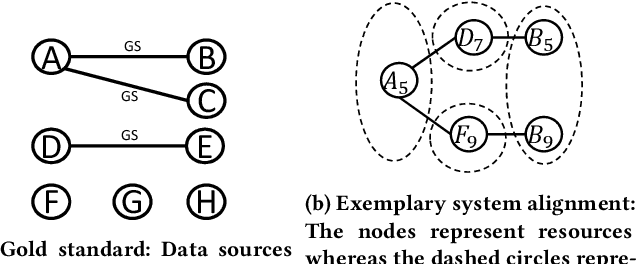
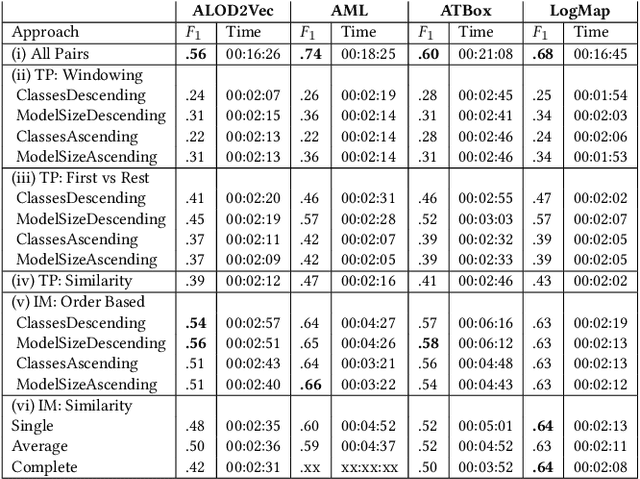
Knowledge graphs (KGs) provide information in machine interpretable form. In cases where multiple KGs are used in the same system, that information needs to be integrated. This is usually done by automated matching systems. Most of those systems consider only 1:1 (binary) matching tasks. Thus, matching a larger number of knowledge graphs with such systems would lead to quadratic efforts. In this paper, we empirically analyze different approaches to reduce the task of multi-source matching to a linear number of executions of binary matching systems. We show that the matching order of KGs and the multi-source strategy actually matter and that near-optimal results can be achieved with linear efforts.
SpectraNet: Learned Recognition of Artificial Satellites From High Contrast Spectroscopic Imagery
Jan 10, 2022
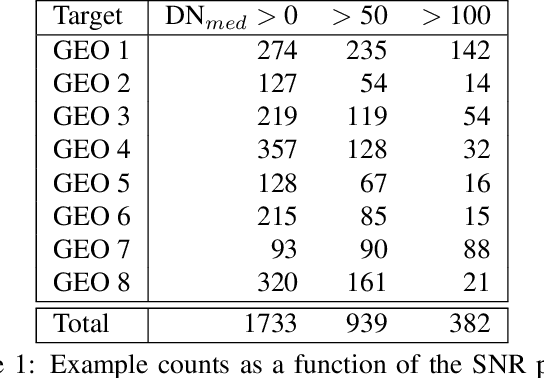
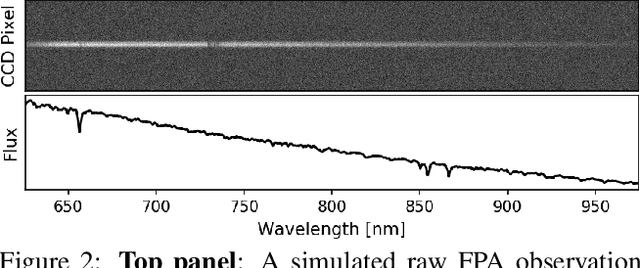
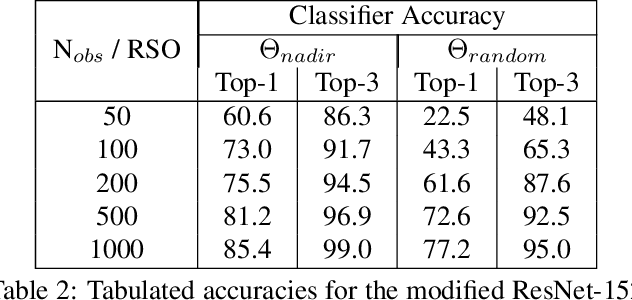
Effective space traffic management requires positive identification of artificial satellites. Current methods for extracting object identification from observed data require spatially resolved imagery which limits identification to objects in low earth orbits. Most artificial satellites, however, operate in geostationary orbits at distances which prohibit ground based observatories from resolving spatial information. This paper demonstrates an object identification solution leveraging modified residual convolutional neural networks to map distance-invariant spectroscopic data to object identity. We report classification accuracies exceeding 80% for a simulated 64-class satellite problem--even in the case of satellites undergoing constant, random re-orientation. An astronomical observing campaign driven by these results returned accuracies of 72% for a nine-class problem with an average of 100 examples per class, performing as expected from simulation. We demonstrate the application of variational Bayesian inference by dropout, stochastic weight averaging (SWA), and SWA-focused deep ensembling to measure classification uncertainties--critical components in space traffic management where routine decisions risk expensive space assets and carry geopolitical consequences.
* 8 pages, 8 figures, 5 tables. Published at WACV 2022
An AI-based Domain-Decomposition Non-Intrusive Reduced-Order Model for Extended Domains applied to Multiphase Flow in Pipes
Feb 13, 2022
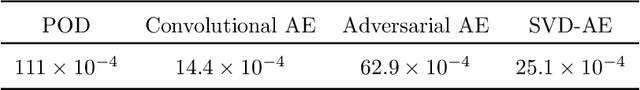
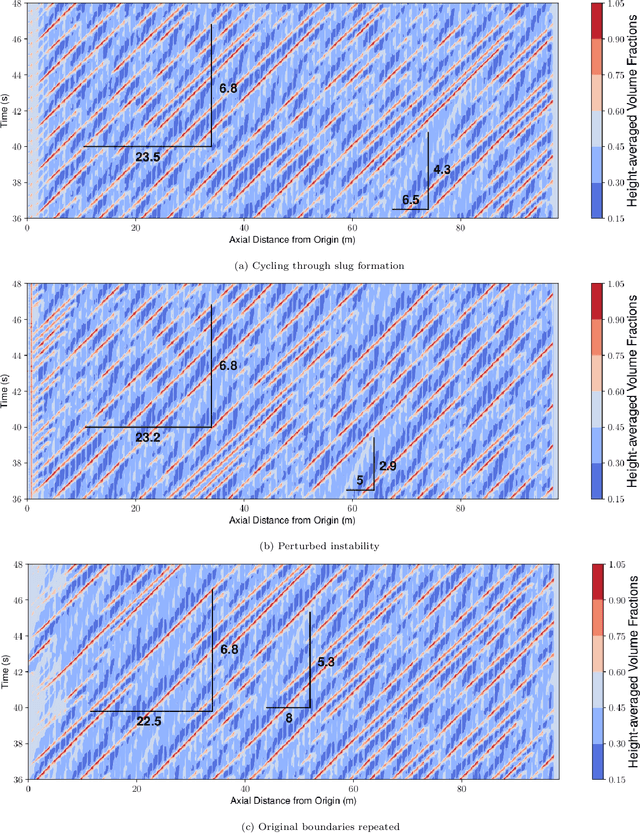
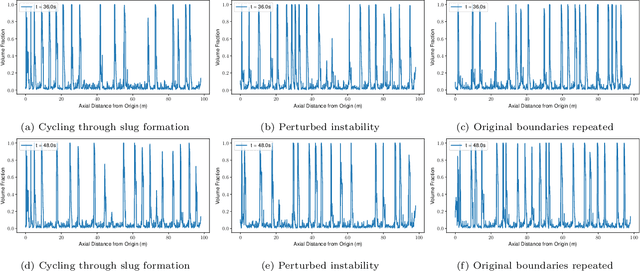
The modelling of multiphase flow in a pipe presents a significant challenge for high-resolution computational fluid dynamics (CFD) models due to the high aspect ratio (length over diameter) of the domain. In subsea applications, the pipe length can be several hundreds of kilometres versus a pipe diameter of just a few inches. In this paper, we present a new AI-based non-intrusive reduced-order model within a domain decomposition framework (AI-DDNIROM) which is capable of making predictions for domains significantly larger than the domain used in training. This is achieved by using domain decomposition; dimensionality reduction; training a neural network to make predictions for a single subdomain; and by using an iteration-by-subdomain technique to converge the solution over the whole domain. To find the low-dimensional space, we explore several types of autoencoder networks, known for their ability to compress information accurately and compactly. The performance of the autoencoders is assessed on two advection-dominated problems: flow past a cylinder and slug flow in a pipe. To make predictions in time, we exploit an adversarial network which aims to learn the distribution of the training data, in addition to learning the mapping between particular inputs and outputs. This type of network has shown the potential to produce realistic outputs. The whole framework is applied to multiphase slug flow in a horizontal pipe for which an AI-DDNIROM is trained on high-fidelity CFD simulations of a pipe of length 10 m with an aspect ratio of 13:1, and tested by simulating the flow for a pipe of length 98 m with an aspect ratio of almost 130:1. Statistics of the flows obtained from the CFD simulations are compared to those of the AI-DDNIROM predictions to demonstrate the success of our approach.
Differentially-Private Clustering of Easy Instances
Dec 29, 2021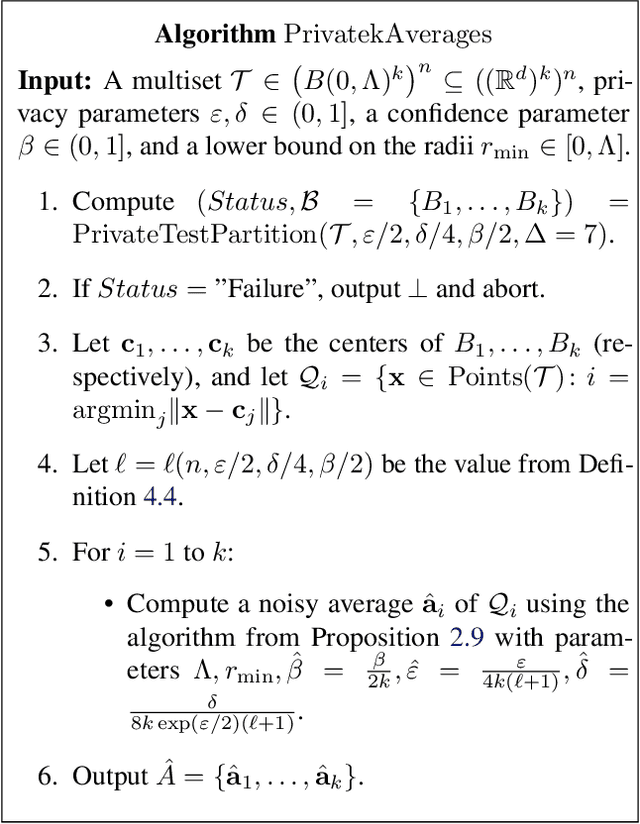
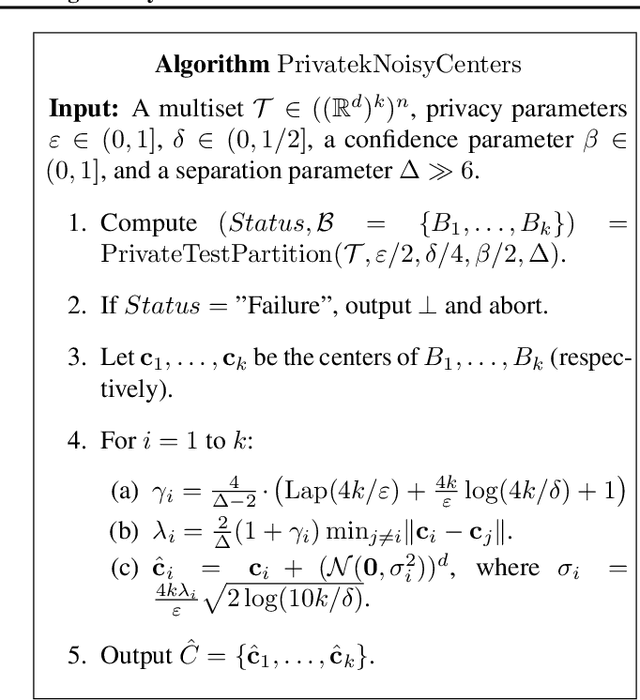

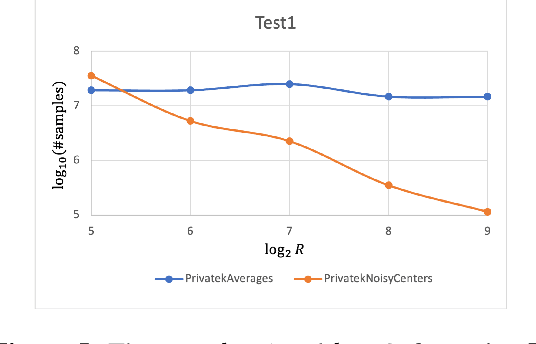
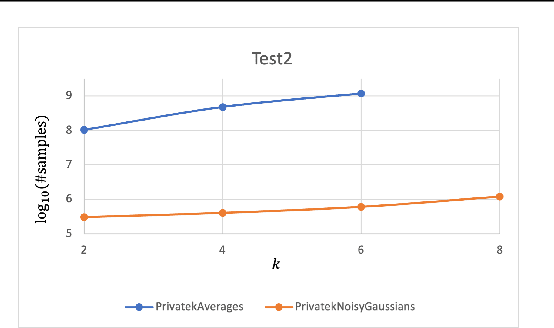
Clustering is a fundamental problem in data analysis. In differentially private clustering, the goal is to identify $k$ cluster centers without disclosing information on individual data points. Despite significant research progress, the problem had so far resisted practical solutions. In this work we aim at providing simple implementable differentially private clustering algorithms that provide utility when the data is "easy," e.g., when there exists a significant separation between the clusters. We propose a framework that allows us to apply non-private clustering algorithms to the easy instances and privately combine the results. We are able to get improved sample complexity bounds in some cases of Gaussian mixtures and $k$-means. We complement our theoretical analysis with an empirical evaluation on synthetic data.
A Scientific Information Extraction Dataset for Nature Inspired Engineering
May 15, 2020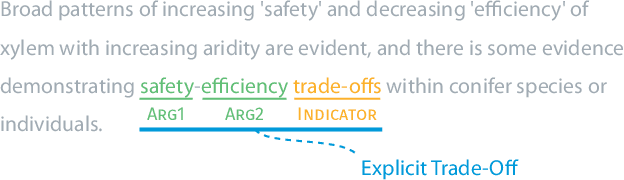
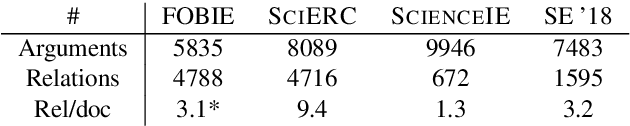

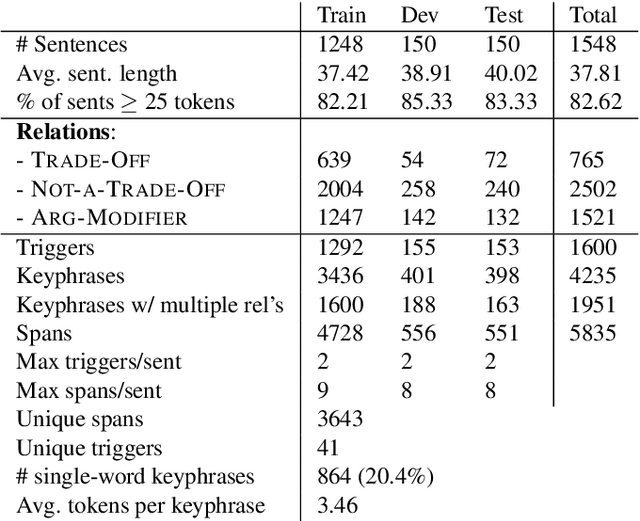
Nature has inspired various ground-breaking technological developments in applications ranging from robotics to aerospace engineering and the manufacturing of medical devices. However, accessing the information captured in scientific biology texts is a time-consuming and hard task that requires domain-specific knowledge. Improving access for outsiders can help interdisciplinary research like Nature Inspired Engineering. This paper describes a dataset of 1,500 manually-annotated sentences that express domain-independent relations between central concepts in a scientific biology text, such as trade-offs and correlations. The arguments of these relations can be Multi Word Expressions and have been annotated with modifying phrases to form non-projective graphs. The dataset allows for training and evaluating Relation Extraction algorithms that aim for coarse-grained typing of scientific biological documents, enabling a high-level filter for engineers.
Learning Population-level Shape Statistics and Anatomy Segmentation From Images: A Joint Deep Learning Model
Jan 10, 2022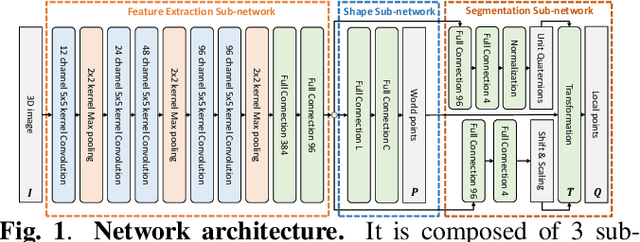
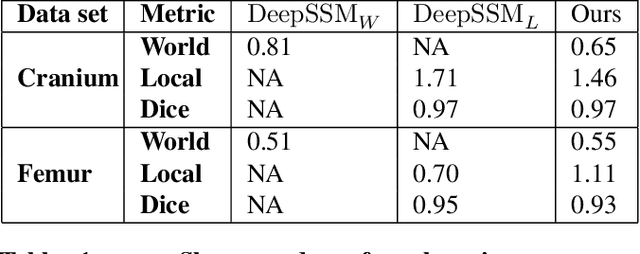
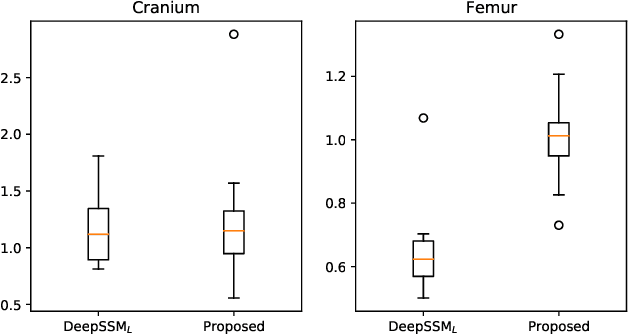
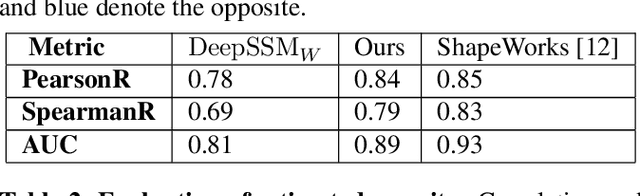
Statistical shape modeling is an essential tool for the quantitative analysis of anatomical populations. Point distribution models (PDMs) represent the anatomical surface via a dense set of correspondences, an intuitive and easy-to-use shape representation for subsequent applications. These correspondences are exhibited in two coordinate spaces: the local coordinates describing the geometrical features of each individual anatomical surface and the world coordinates representing the population-level statistical shape information after removing global alignment differences across samples in the given cohort. We propose a deep-learning-based framework that simultaneously learns these two coordinate spaces directly from the volumetric images. The proposed joint model serves a dual purpose; the world correspondences can directly be used for shape analysis applications, circumventing the heavy pre-processing and segmentation involved in traditional PDM models. Additionally, the local correspondences can be used for anatomy segmentation. We demonstrate the efficacy of this joint model for both shape modeling applications on two datasets and its utility in inferring the anatomical surface.
The Computational Drug Repositioning without Negative Sampling
Nov 29, 2021
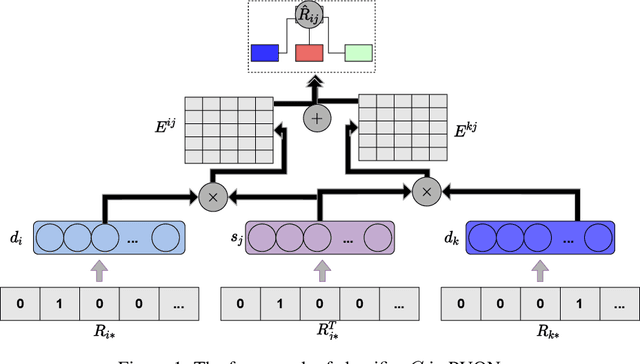
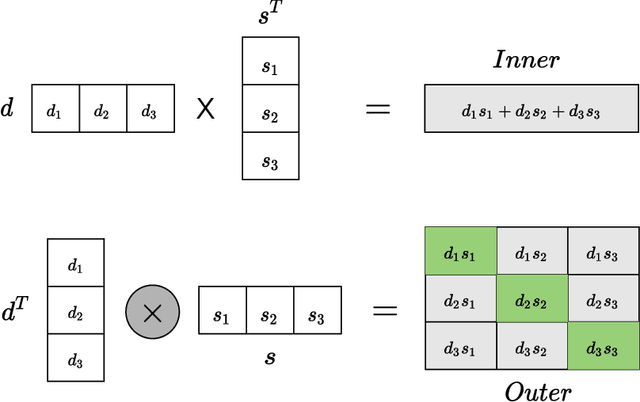
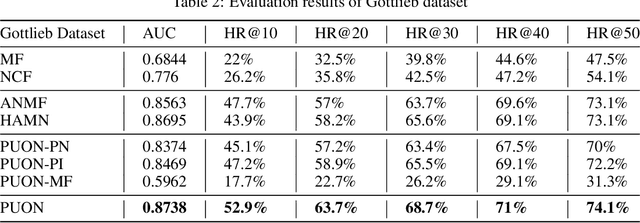
Computational drug repositioning technology is an effective tool to accelerate drug development. Although this technique has been widely used and successful in recent decades, many existing models still suffer from multiple drawbacks such as the massive number of unvalidated drug-disease associations and inner product in the matrix factorization model. The limitations of these works are mainly due to the following two reasons: first, previous works used negative sampling techniques to treat unvalidated drug-disease associations as negative samples, which is invalid in real-world settings; Second, the inner product lacks modeling on the crossover information between dimensions of the latent factor. In this paper, we propose a novel PUON framework for addressing the above deficiencies, which models the joint distribution of drug-disease associations using validated and unvalidated drug-disease associations without employing negative sampling techniques. The PUON also modeled the cross-information of the latent factor of drugs and diseases using the outer product operation. For a comprehensive comparison, we considered 7 popular baselines. Extensive experiments in two real-world datasets showed that PUON achieved the best performance based on 6 popular evaluation metrics.
Leveraging Social Influence based on Users Activity Centers for Point-of-Interest Recommendation
Jan 10, 2022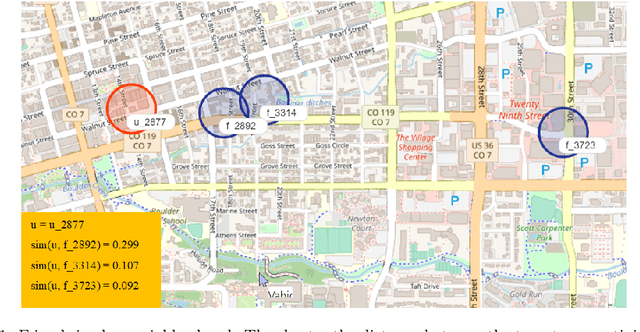
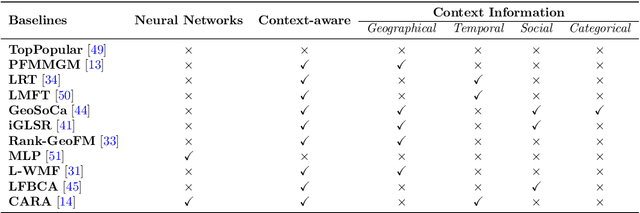
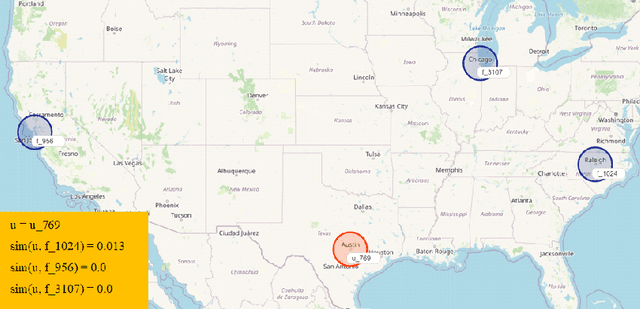
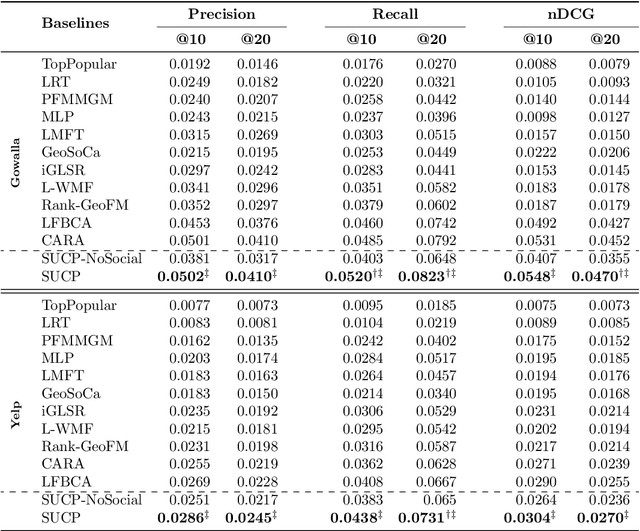
Recommender Systems (RSs) aim to model and predict the user preference while interacting with items, such as Points of Interest (POIs). These systems face several challenges, such as data sparsity, limiting their effectiveness. In this paper, we address this problem by incorporating social, geographical, and temporal information into the Matrix Factorization (MF) technique. To this end, we model social influence based on two factors: similarities between users in terms of common check-ins and the friendships between them. We introduce two levels of friendship based on explicit friendship networks and high check-in overlap between users. We base our friendship algorithm on users' geographical activity centers. The results show that our proposed model outperforms the state-of-the-art on two real-world datasets. More specifically, our ablation study shows that the social model improves the performance of our proposed POI recommendation system by 31% and 14% on the Gowalla and Yelp datasets in terms of Precision@10, respectively.
Learning with less labels in Digital Pathology via Scribble Supervision from natural images
Jan 07, 2022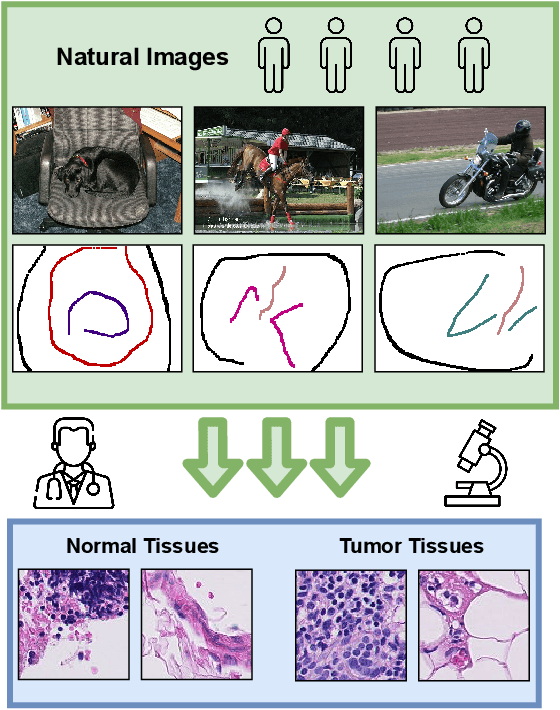
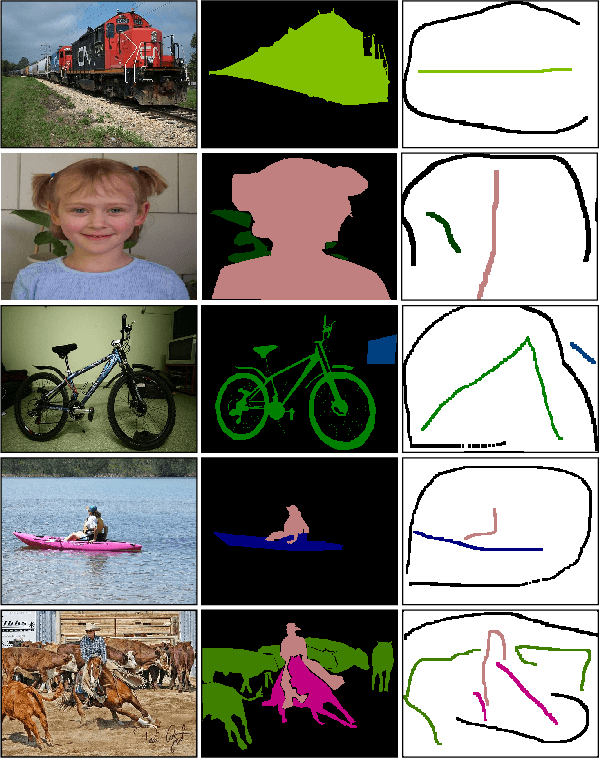
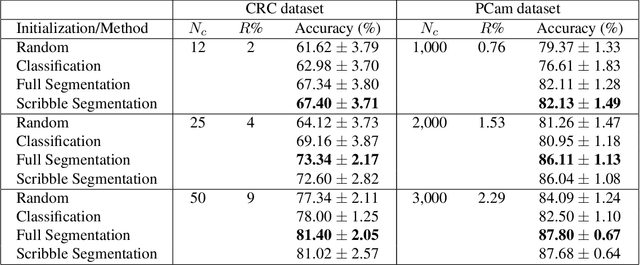
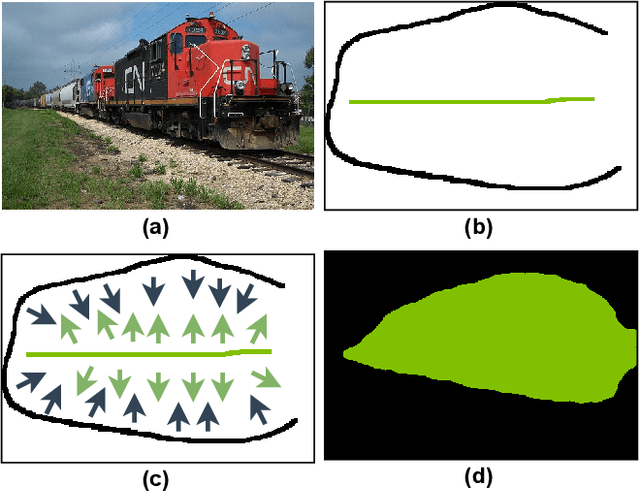

A critical challenge of training deep learning models in the Digital Pathology (DP) domain is the high annotation cost by medical experts. One way to tackle this issue is via transfer learning from the natural image domain (NI), where the annotation cost is considerably cheaper. Cross-domain transfer learning from NI to DP is shown to be successful via class labels~\cite{teh2020learning}. One potential weakness of relying on class labels is the lack of spatial information, which can be obtained from spatial labels such as full pixel-wise segmentation labels and scribble labels. We demonstrate that scribble labels from NI domain can boost the performance of DP models on two cancer classification datasets (Patch Camelyon Breast Cancer and Colorectal Cancer dataset). Furthermore, we show that models trained with scribble labels yield the same performance boost as full pixel-wise segmentation labels despite being significantly easier and faster to collect.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge