"Information": models, code, and papers
Local Topology Inference of Mobile Robotic Networks under Formation Control
Apr 30, 2022
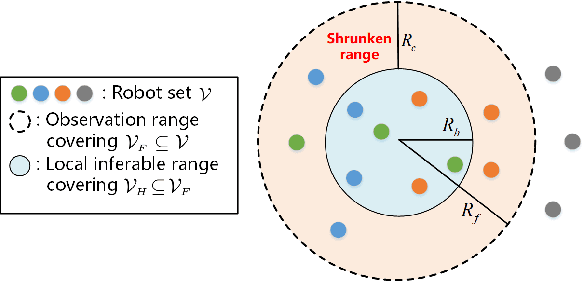
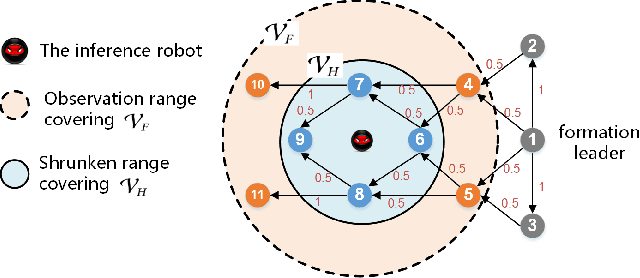
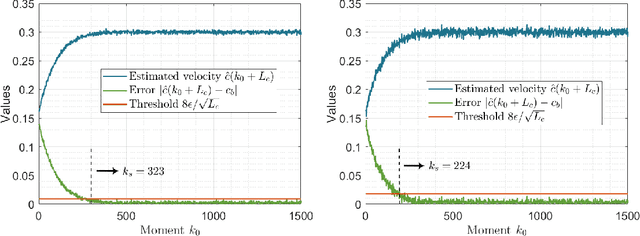
The interaction topology is critical for efficient cooperation of mobile robotic networks (MRNs). We focus on the local topology inference problem of MRNs under formation control, where an inference robot with limited observation range can manoeuvre among the formation robots. This problem faces new challenges brought by the highly coupled influence of unobservable formation robots, inaccessible formation inputs, and unknown interaction range. The novel idea here is to advocate a range-shrink strategy to perfectly avoid the influence of unobservable robots while filtering the input. To that end, we develop consecutive algorithms to determine a feasible constant robot subset from the changing robot set within the observation range, and estimate the formation input and the interaction range. Then, an ordinary least squares based local topology estimator is designed with the previously inferred information. Resorting to the concentration measure, we prove the convergence rate and accuracy of the proposed estimator, taking the estimation errors of previous steps into account. Extensions on nonidentical observation slots and more complicated scenarios are also analyzed. Comprehensive simulation tests and method comparisons corroborate the theoretical findings.
Fast ABC-Boost: A Unified Framework for Selecting the Base Class in Multi-Class Classification
May 22, 2022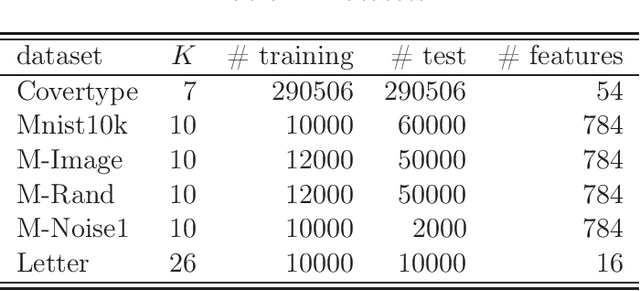
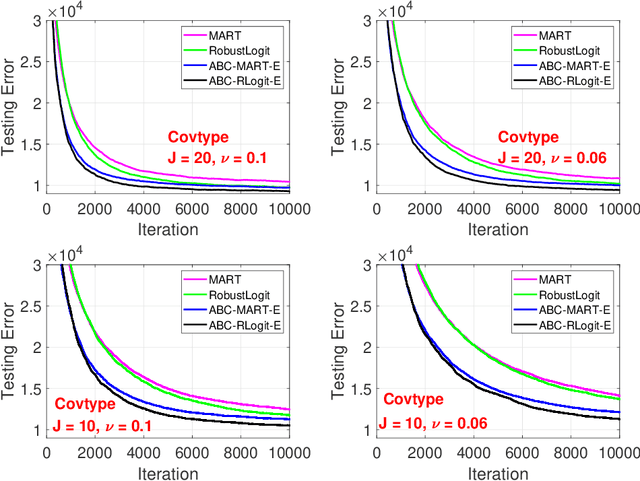
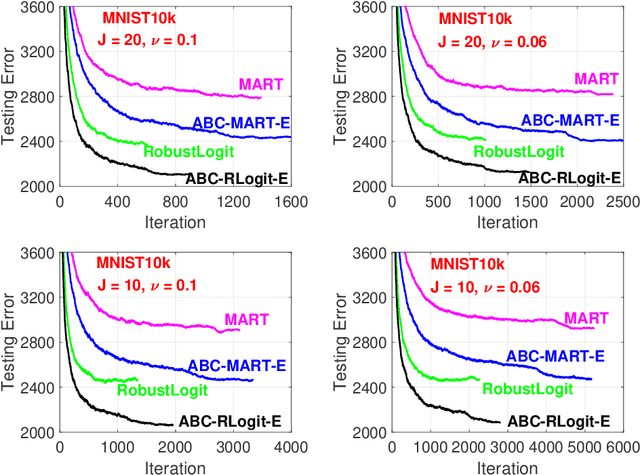
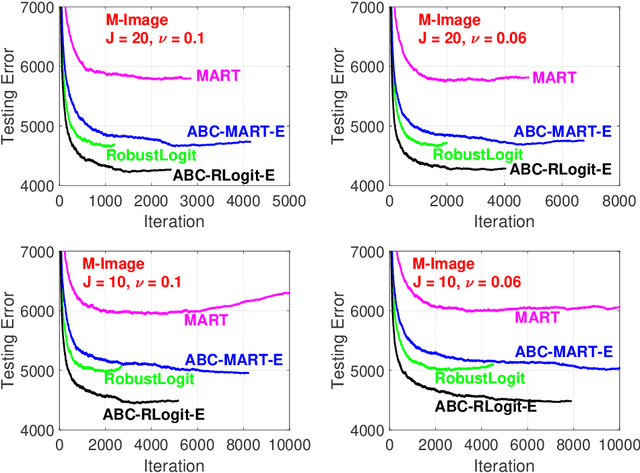
The work in ICML'09 showed that the derivatives of the classical multi-class logistic regression loss function could be re-written in terms of a pre-chosen "base class" and applied the new derivatives in the popular boosting framework. In order to make use of the new derivatives, one must have a strategy to identify/choose the base class at each boosting iteration. The idea of "adaptive base class boost" (ABC-Boost) in ICML'09, adopted a computationally expensive "exhaustive search" strategy for the base class at each iteration. It has been well demonstrated that ABC-Boost, when integrated with trees, can achieve substantial improvements in many multi-class classification tasks. Furthermore, the work in UAI'10 derived the explicit second-order tree split gain formula which typically improved the classification accuracy considerably, compared with using only the fist-order information for tree-splitting, for both multi-class and binary-class classification tasks. In this paper, we develop a unified framework for effectively selecting the base class by introducing a series of ideas to improve the computational efficiency of ABC-Boost. Our framework has parameters $(s,g,w)$. At each boosting iteration, we only search for the "$s$-worst classes" (instead of all classes) to determine the base class. We also allow a "gap" $g$ when conducting the search. That is, we only search for the base class at every $g+1$ iterations. We furthermore allow a "warm up" stage by only starting the search after $w$ boosting iterations. The parameters $s$, $g$, $w$, can be viewed as tunable parameters and certain combinations of $(s,g,w)$ may even lead to better test accuracy than the "exhaustive search" strategy. Overall, our proposed framework provides a robust and reliable scheme for implementing ABC-Boost in practice.
Recovering Quantitative Models of Human Information Processing with Differentiable Architecture Search
Mar 25, 2021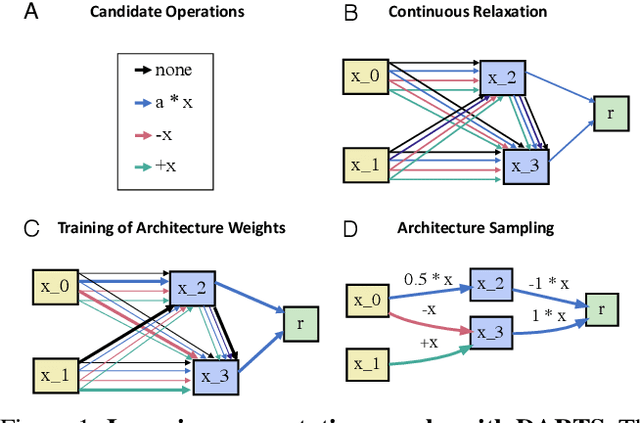
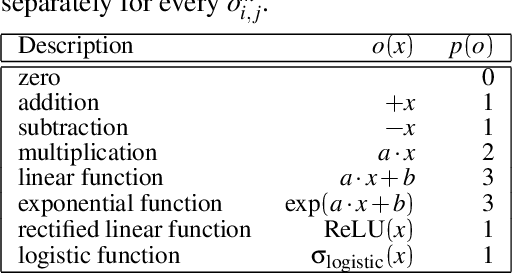
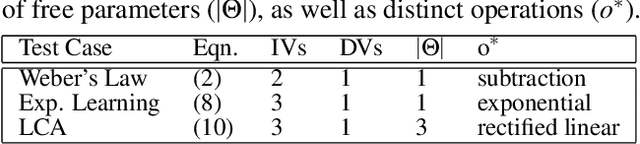


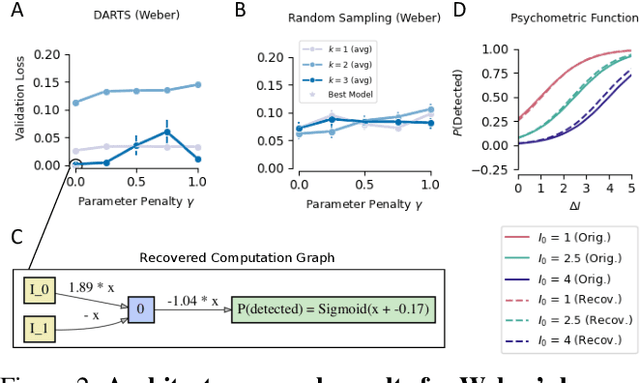
The integration of behavioral phenomena into mechanistic models of cognitive function is a fundamental staple of cognitive science. Yet, researchers are beginning to accumulate increasing amounts of data without having the temporal or monetary resources to integrate these data into scientific theories. We seek to overcome these limitations by incorporating existing machine learning techniques into an open-source pipeline for the automated construction of quantitative models. This pipeline leverages the use of neural architecture search to automate the discovery of interpretable model architectures, and automatic differentiation to automate the fitting of model parameters to data. We evaluate the utility of these methods based on their ability to recover quantitative models of human information processing from synthetic data. We find that these methods are capable of recovering basic quantitative motifs from models of psychophysics, learning and decision making. We also highlight weaknesses of this framework, and discuss future directions for their mitigation.
BEIR: A Heterogenous Benchmark for Zero-shot Evaluation of Information Retrieval Models
Apr 28, 2021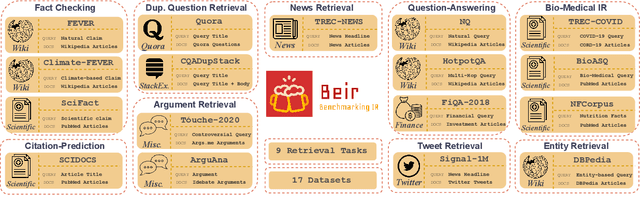
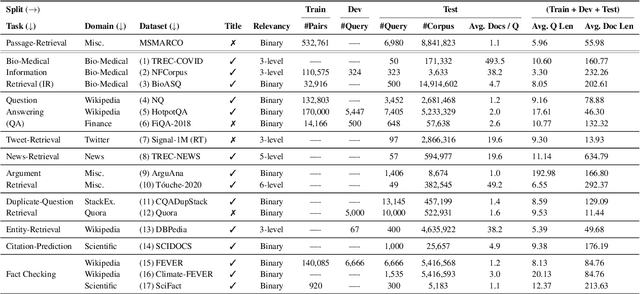
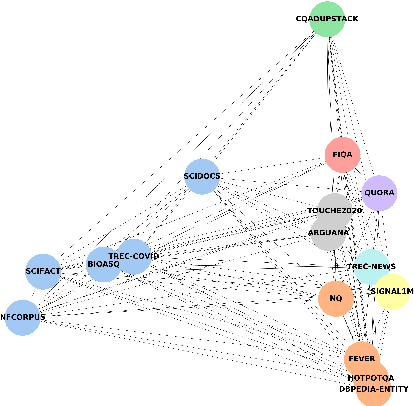
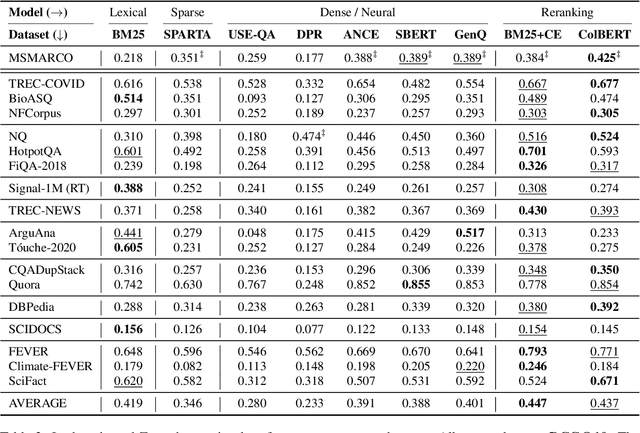
Neural IR models have often been studied in homogeneous and narrow settings, which has considerably limited insights into their generalization capabilities. To address this, and to allow researchers to more broadly establish the effectiveness of their models, we introduce BEIR (Benchmarking IR), a heterogeneous benchmark for information retrieval. We leverage a careful selection of 17 datasets for evaluation spanning diverse retrieval tasks including open-domain datasets as well as narrow expert domains. We study the effectiveness of nine state-of-the-art retrieval models in a zero-shot evaluation setup on BEIR, finding that performing well consistently across all datasets is challenging. Our results show BM25 is a robust baseline and Reranking-based models overall achieve the best zero-shot performances, however, at high computational costs. In contrast, Dense-retrieval models are computationally more efficient but often underperform other approaches, highlighting the considerable room for improvement in their generalization capabilities. In this work, we extensively analyze different retrieval models and provide several suggestions that we believe may be useful for future work. BEIR datasets and code are available at https://github.com/UKPLab/beir.
Biomedical Information Extraction for Disease Gene Prioritization
Nov 12, 2020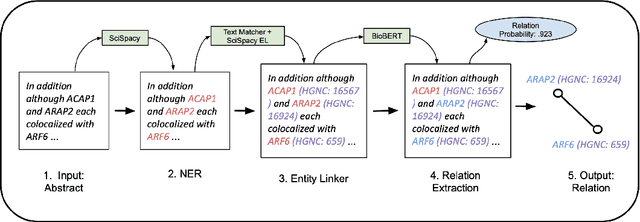
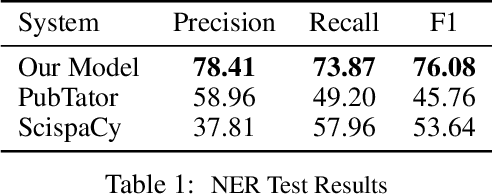
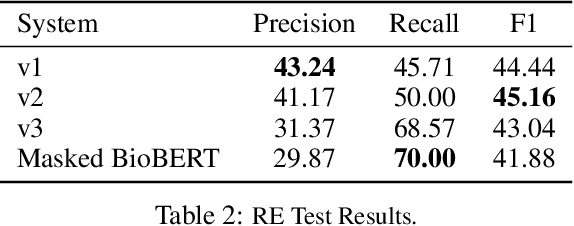
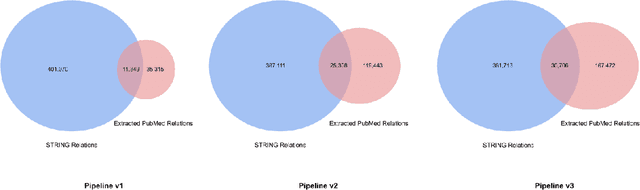
We introduce a biomedical information extraction (IE) pipeline that extracts biological relationships from text and demonstrate that its components, such as named entity recognition (NER) and relation extraction (RE), outperform state-of-the-art in BioNLP. We apply it to tens of millions of PubMed abstracts to extract protein-protein interactions (PPIs) and augment these extractions to a biomedical knowledge graph that already contains PPIs extracted from STRING, the leading structured PPI database. We show that, despite already containing PPIs from an established structured source, augmenting our own IE-based extractions to the graph allows us to predict novel disease-gene associations with a 20% relative increase in hit@30, an important step towards developing drug targets for uncured diseases.
Generative De Novo Protein Design with Global Context
Apr 21, 2022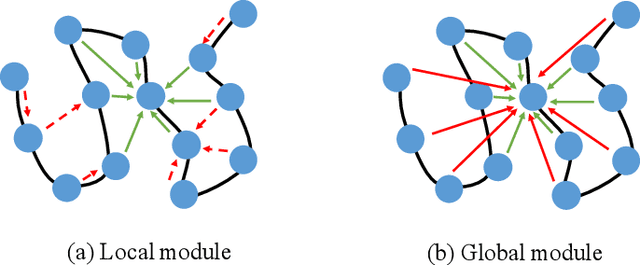
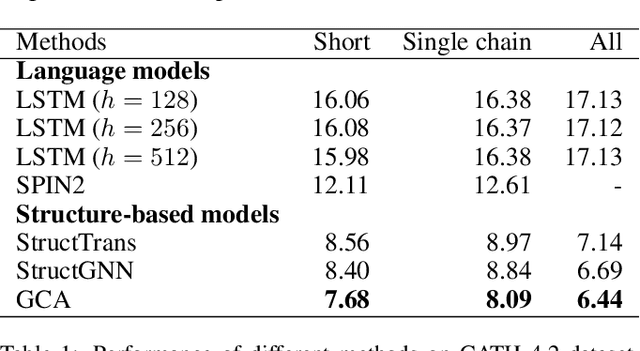
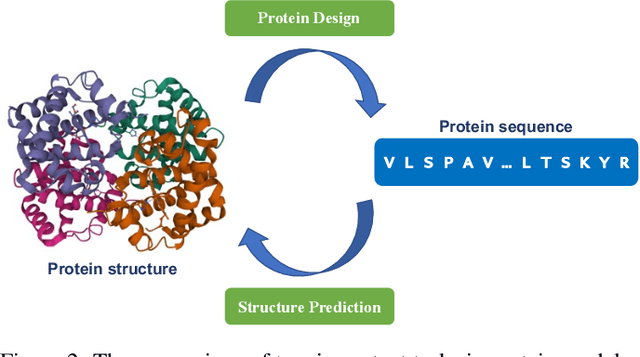
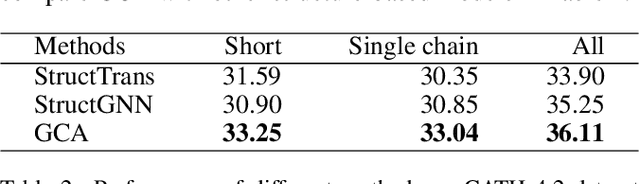
The linear sequence of amino acids determines protein structure and function. Protein design, known as the inverse of protein structure prediction, aims to obtain a novel protein sequence that will fold into the defined structure. Recent works on computational protein design have studied designing sequences for the desired backbone structure with local positional information and achieved competitive performance. However, similar local environments in different backbone structures may result in different amino acids, indicating that protein structure's global context matters. Thus, we propose the Global-Context Aware generative de novo protein design method (GCA), consisting of local and global modules. While local modules focus on relationships between neighbor amino acids, global modules explicitly capture non-local contexts. Experimental results demonstrate that the proposed GCA method outperforms state-of-the-arts on de novo protein design. Our code and pretrained model will be released.
Learning to Ground Decentralized Multi-Agent Communication with Contrastive Learning
Mar 07, 2022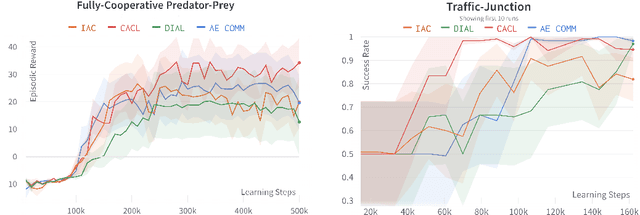
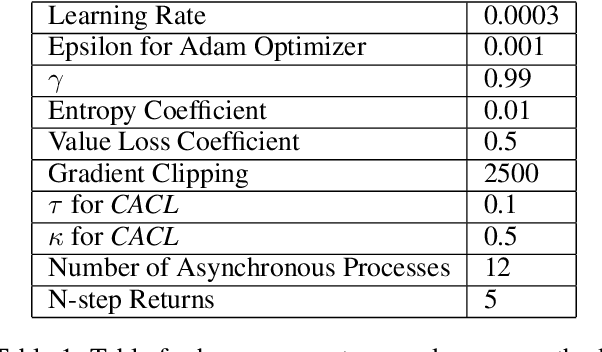
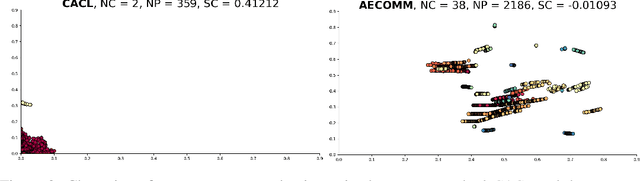
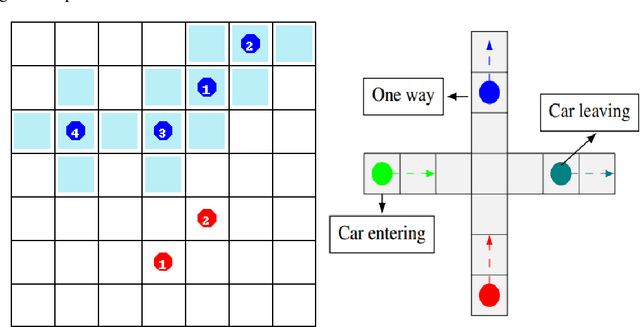
For communication to happen successfully, a common language is required between agents to understand information communicated by one another. Inducing the emergence of a common language has been a difficult challenge to multi-agent learning systems. In this work, we introduce an alternative perspective to the communicative messages sent between agents, considering them as different incomplete views of the environment state. Based on this perspective, we propose a simple approach to induce the emergence of a common language by maximizing the mutual information between messages of a given trajectory in a self-supervised manner. By evaluating our method in communication-essential environments, we empirically show how our method leads to better learning performance and speed, and learns a more consistent common language than existing methods, without introducing additional learning parameters.
Deep Reinforcement Learning for Data-Driven Adaptive Scanning in Ptychography
Mar 29, 2022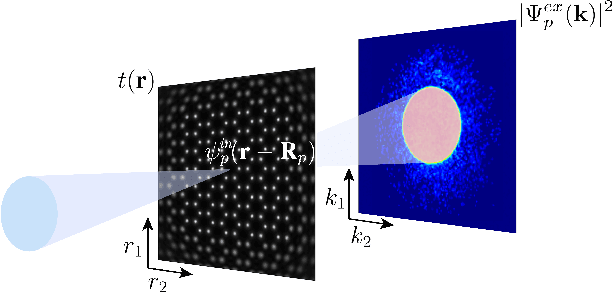
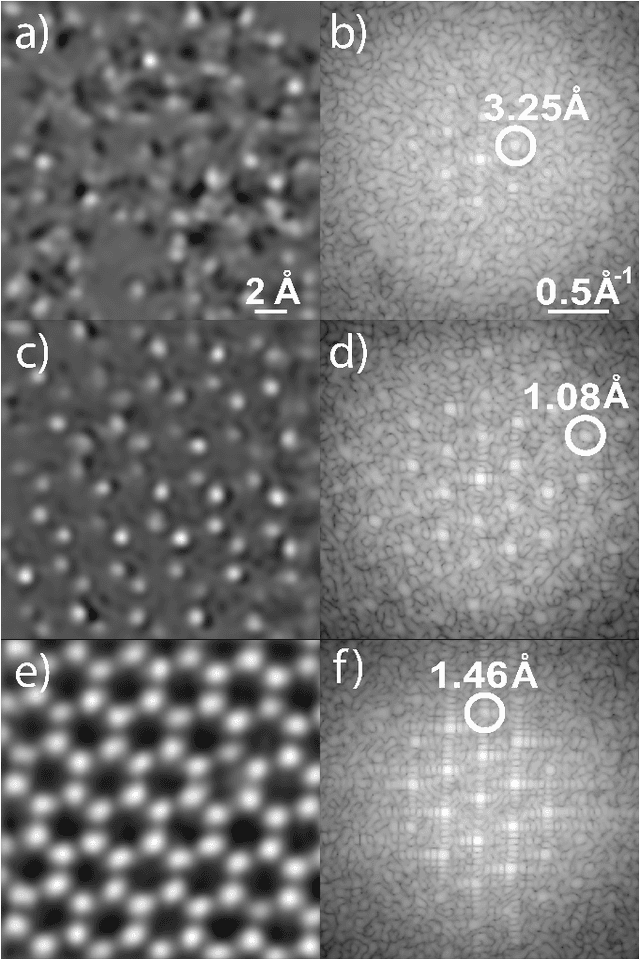
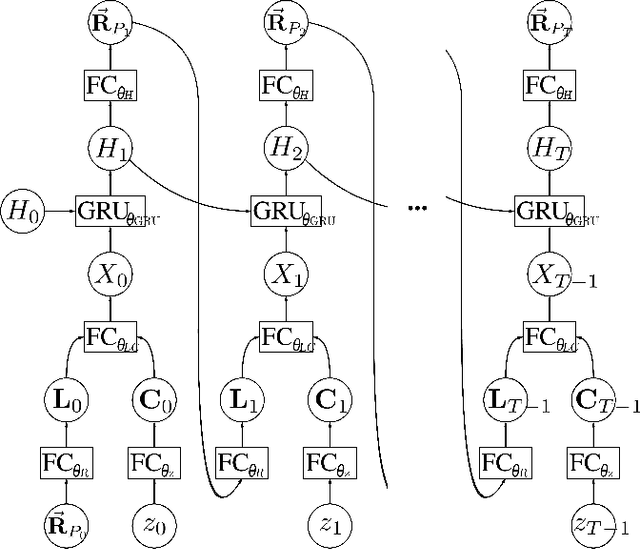
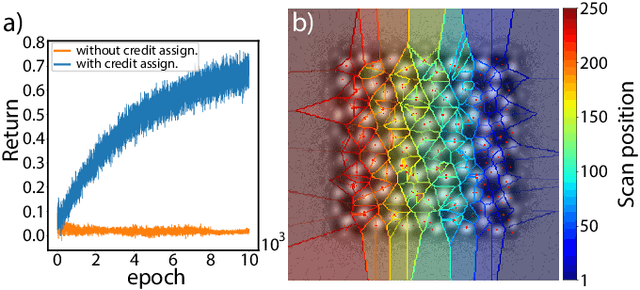

We present a method that lowers the dose required for a ptychographic reconstruction by adaptively scanning the specimen, thereby providing the required spatial information redundancy in the regions of highest importance. The proposed method is built upon a deep learning model that is trained by reinforcement learning (RL), using prior knowledge of the specimen structure from training data sets. We show that equivalent low-dose experiments using adaptive scanning outperform conventional ptychography experiments in terms of reconstruction resolution.
Unsupervised Learning of 3D Semantic Keypoints with Mutual Reconstruction
Mar 19, 2022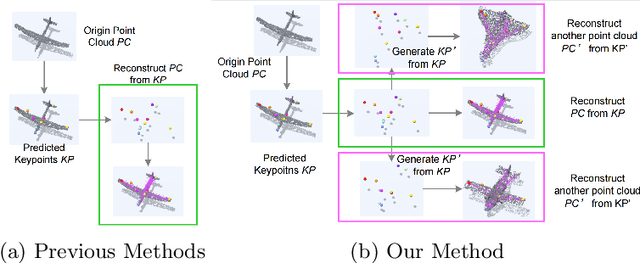
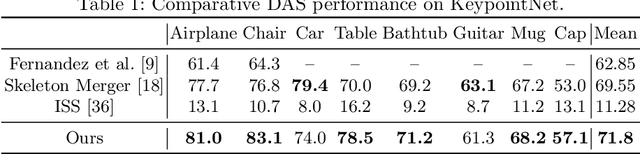
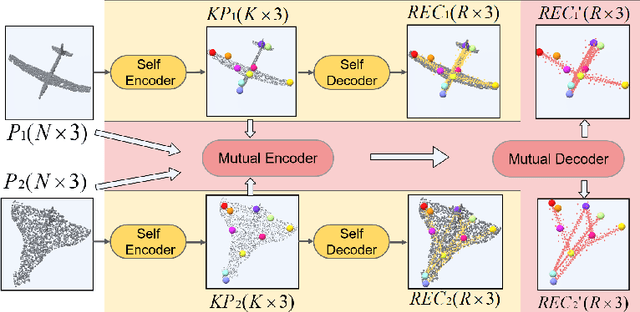
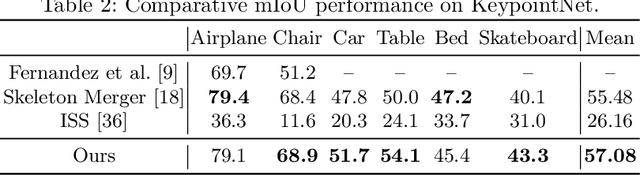
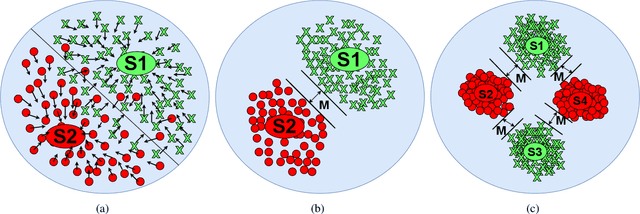
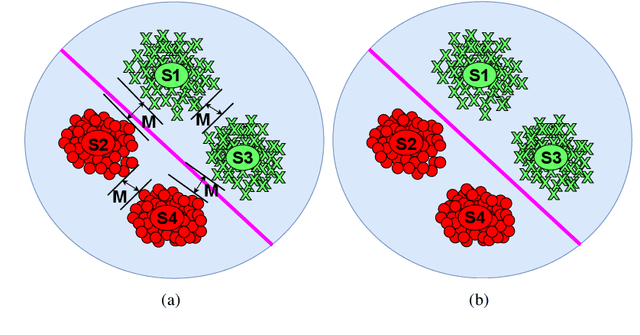
Semantic 3D keypoints are category-level semantic consistent points on 3D objects. Detecting 3D semantic keypoints is a foundation for a number of 3D vision tasks but remains challenging, due to the ambiguity of semantic information, especially when the objects are represented by unordered 3D point clouds. Existing unsupervised methods tend to generate category-level keypoints in implicit manners, making it difficult to extract high-level information, such as semantic labels and topology. From a novel mutual reconstruction perspective, we present an unsupervised method to generate consistent semantic keypoints from point clouds explicitly. To achieve this, the proposed model predicts keypoints that not only reconstruct the object itself but also reconstruct other instances in the same category. To the best of our knowledge, the proposed method is the first to mine 3D semantic consistent keypoints from a mutual reconstruction view. Experiments under various evaluation metrics as well as comparisons with the state-of-the-arts demonstrate the efficacy of our new solution to mining semantic consistent keypoints with mutual reconstruction.
Biceph-Net: A robust and lightweight framework for the diagnosis of Alzheimer's disease using 2D-MRI scans and deep similarity learning
Mar 23, 2022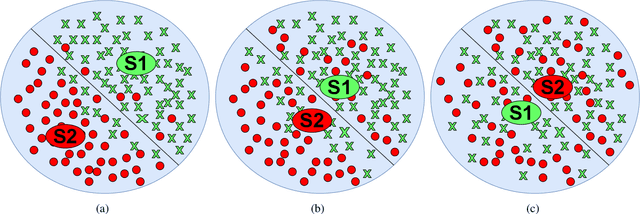
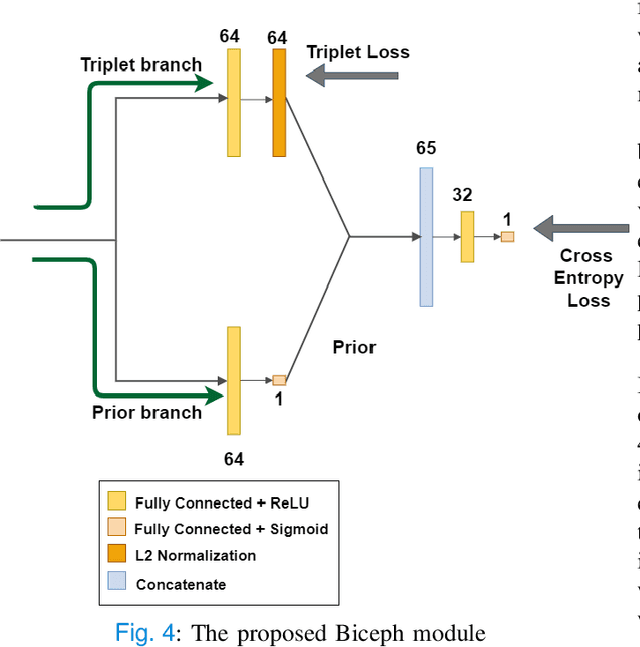
Alzheimer's Disease (AD) is a neurodegenerative disease that is one of the significant causes of death in the elderly population. Many deep learning techniques have been proposed to diagnose AD using Magnetic Resonance Imaging (MRI) scans. Predicting AD using 2D slices extracted from 3D MRI scans is challenging as the inter-slice information gets lost. To this end, we propose a novel and lightweight framework termed 'Biceph-Net' for AD diagnosis using 2D MRI scans that model both the intra-slice and inter-slice information. Biceph-Net has been experimentally shown to perform similar to other Spatio-temporal neural networks while being computationally more efficient. Biceph-Net is also superior in performance compared to vanilla 2D convolutional neural networks (CNN) for AD diagnosis using 2D MRI slices. Biceph-Net also has an inbuilt neighbourhood-based model interpretation feature that can be exploited to understand the classification decision taken by the network. Biceph-Net experimentally achieves a test accuracy of 100% in the classification of Cognitively Normal (CN) vs AD, 98.16% for Mild Cognitive Impairment (MCI) vs AD, and 97.80% for CN vs MCI vs AD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge