"Information": models, code, and papers
Alignahead: Online Cross-Layer Knowledge Extraction on Graph Neural Networks
May 05, 2022
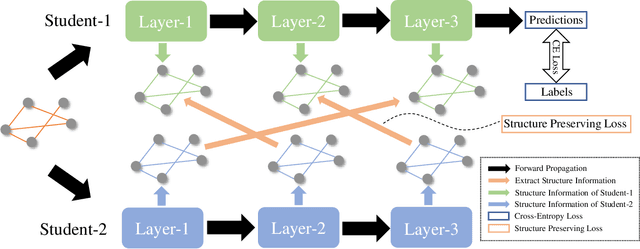
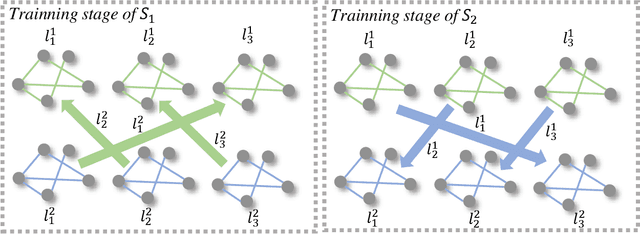
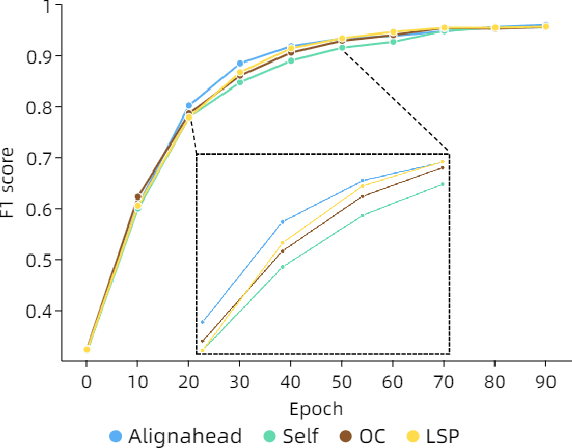
Existing knowledge distillation methods on graph neural networks (GNNs) are almost offline, where the student model extracts knowledge from a powerful teacher model to improve its performance. However, a pre-trained teacher model is not always accessible due to training cost, privacy, etc. In this paper, we propose a novel online knowledge distillation framework to resolve this problem. Specifically, each student GNN model learns the extracted local structure from another simultaneously trained counterpart in an alternating training procedure. We further develop a cross-layer distillation strategy by aligning ahead one student layer with the layer in different depth of another student model, which theoretically makes the structure information spread over all layers. Experimental results on five datasets including PPI, Coauthor-CS/Physics and Amazon-Computer/Photo demonstrate that the student performance is consistently boosted in our collaborative training framework without the supervision of a pre-trained teacher model. In addition, we also find that our alignahead technique can accelerate the model convergence speed and its effectiveness can be generally improved by increasing the student numbers in training. Code is available: https://github.com/GuoJY-eatsTG/Alignahead
Boosting Semi-supervised Image Segmentation with Global and Local Mutual Information Regularization
Mar 08, 2021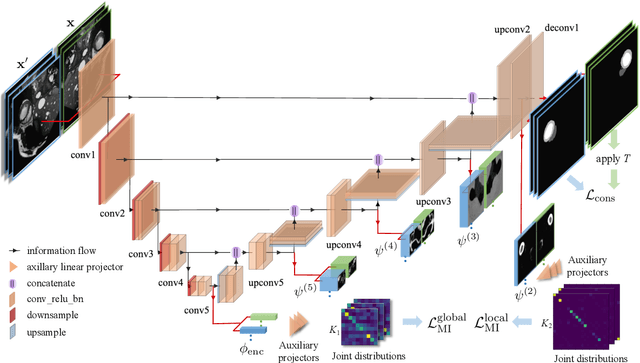
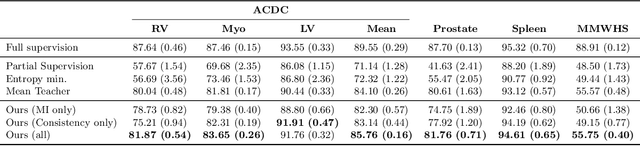
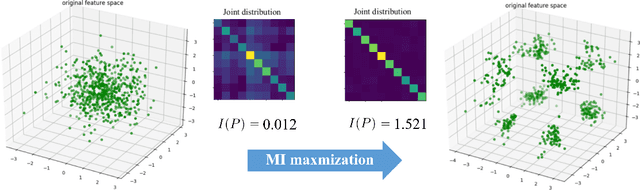
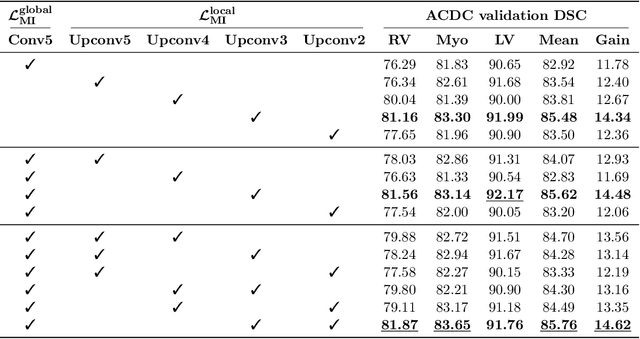
The scarcity of labeled data often impedes the application of deep learning to the segmentation of medical images. Semi-supervised learning seeks to overcome this limitation by leveraging unlabeled examples in the learning process. In this paper, we present a novel semi-supervised segmentation method that leverages mutual information (MI) on categorical distributions to achieve both global representation invariance and local smoothness. In this method, we maximize the MI for intermediate feature embeddings that are taken from both the encoder and decoder of a segmentation network. We first propose a global MI loss constraining the encoder to learn an image representation that is invariant to geometric transformations. Instead of resorting to computationally-expensive techniques for estimating the MI on continuous feature embeddings, we use projection heads to map them to a discrete cluster assignment where MI can be computed efficiently. Our method also includes a local MI loss to promote spatial consistency in the feature maps of the decoder and provide a smoother segmentation. Since mutual information does not require a strict ordering of clusters in two different assignments, we incorporate a final consistency regularization loss on the output which helps align the cluster labels throughout the network. We evaluate the method on three challenging publicly-available datasets for medical image segmentation. Experimental results show our method to outperform recently-proposed approaches for semi-supervised segmentation and provide an accuracy near to full supervision while training with very few annotated images
i-Code: An Integrative and Composable Multimodal Learning Framework
May 05, 2022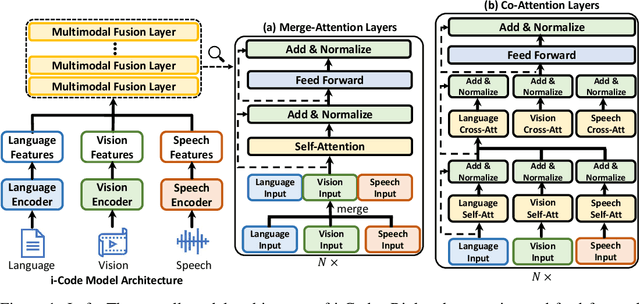
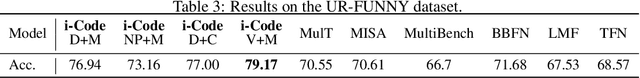
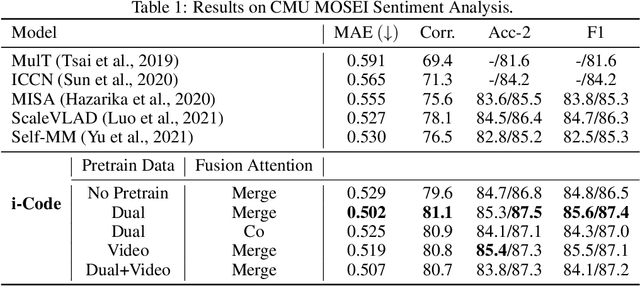
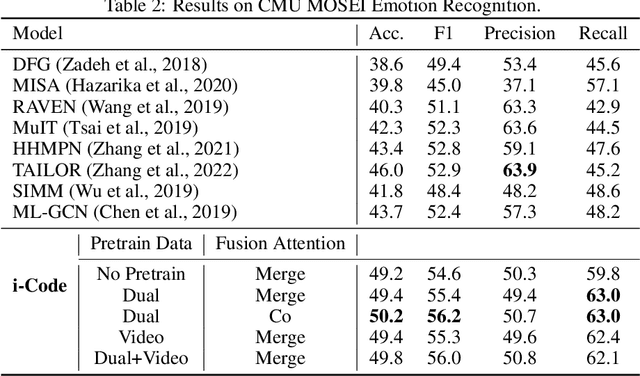

Human intelligence is multimodal; we integrate visual, linguistic, and acoustic signals to maintain a holistic worldview. Most current pretraining methods, however, are limited to one or two modalities. We present i-Code, a self-supervised pretraining framework where users may flexibly combine the modalities of vision, speech, and language into unified and general-purpose vector representations. In this framework, data from each modality are first given to pretrained single-modality encoders. The encoder outputs are then integrated with a multimodal fusion network, which uses novel attention mechanisms and other architectural innovations to effectively combine information from the different modalities. The entire system is pretrained end-to-end with new objectives including masked modality unit modeling and cross-modality contrastive learning. Unlike previous research using only video for pretraining, the i-Code framework can dynamically process single, dual, and triple-modality data during training and inference, flexibly projecting different combinations of modalities into a single representation space. Experimental results demonstrate how i-Code can outperform state-of-the-art techniques on five video understanding tasks and the GLUE NLP benchmark, improving by as much as 11% and demonstrating the power of integrative multimodal pretraining.
Multimodal Machine Learning in Precision Health
Apr 10, 2022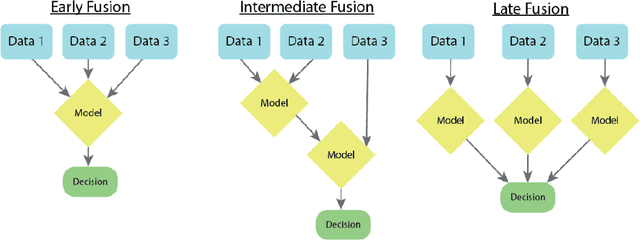
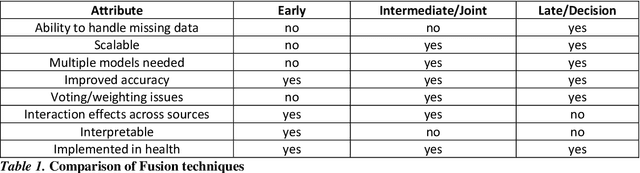

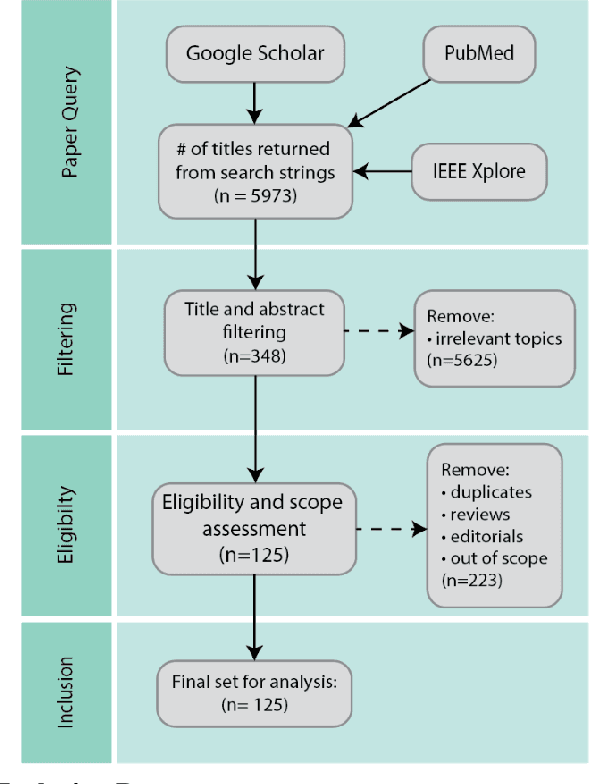
As machine learning and artificial intelligence are more frequently being leveraged to tackle problems in the health sector, there has been increased interest in utilizing them in clinical decision-support. This has historically been the case in single modal data such as electronic health record data. Attempts to improve prediction and resemble the multimodal nature of clinical expert decision-making this has been met in the computational field of machine learning by a fusion of disparate data. This review was conducted to summarize this field and identify topics ripe for future research. We conducted this review in accordance with the PRISMA (Preferred Reporting Items for Systematic reviews and Meta-Analyses) extension for Scoping Reviews to characterize multi-modal data fusion in health. We used a combination of content analysis and literature searches to establish search strings and databases of PubMed, Google Scholar, and IEEEXplore from 2011 to 2021. A final set of 125 articles were included in the analysis. The most common health areas utilizing multi-modal methods were neurology and oncology. However, there exist a wide breadth of current applications. The most common form of information fusion was early fusion. Notably, there was an improvement in predictive performance performing heterogeneous data fusion. Lacking from the papers were clear clinical deployment strategies and pursuit of FDA-approved tools. These findings provide a map of the current literature on multimodal data fusion as applied to health diagnosis/prognosis problems. Multi-modal machine learning, while more robust in its estimations over unimodal methods, has drawbacks in its scalability and the time-consuming nature of information concatenation.
Classification of Hyperspectral Images Using SVM with Shape-adaptive Reconstruction and Smoothed Total Variation
Apr 14, 2022
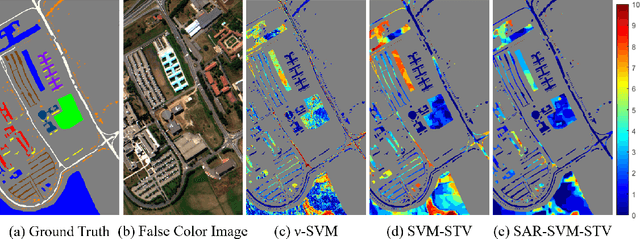
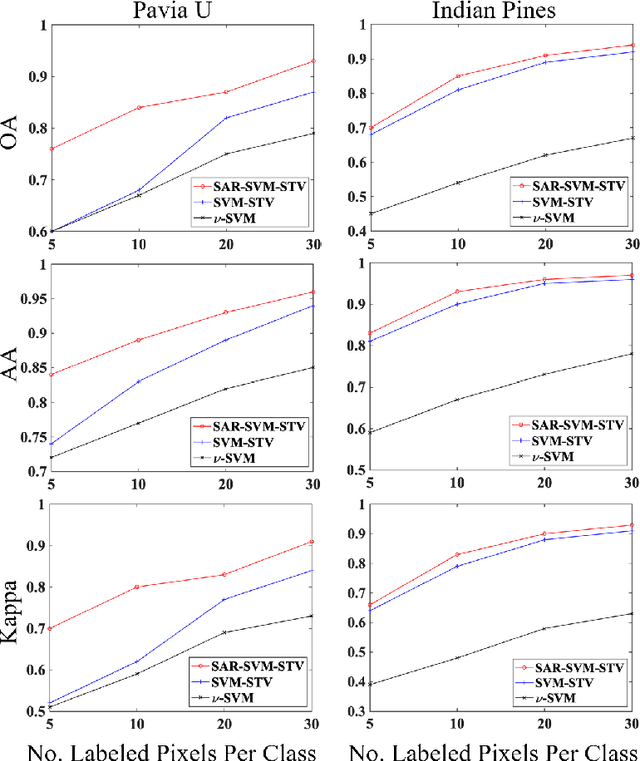
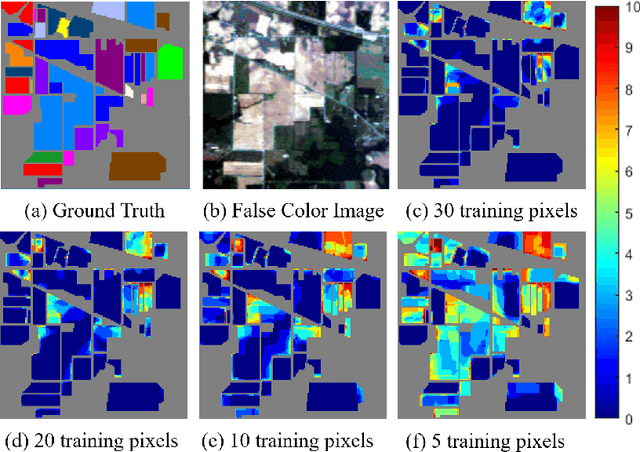

In this work, a novel algorithm called SVM with Shape-adaptive Reconstruction and Smoothed Total Variation (SaR-SVM-STV) is introduced to classify hyperspectral images, which makes full use of spatial and spectral information. The Shape-adaptive Reconstruction (SaR) is introduced to preprocess each pixel based on the Pearson Correlation between pixels in its shape-adaptive (SA) region. Support Vector Machines (SVMs) are trained to estimate the pixel-wise probability maps of each class. Then the Smoothed Total Variation (STV) model is applied to denoise and generate the final classification map. Experiments show that SaR-SVM-STV outperforms the SVM-STV method with a few training labels, demonstrating the significance of reconstructing hyperspectral images before classification.
Evidence-aware Fake News Detection with Graph Neural Networks
Feb 08, 2022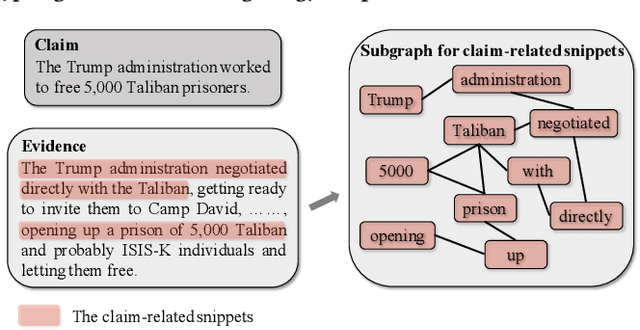
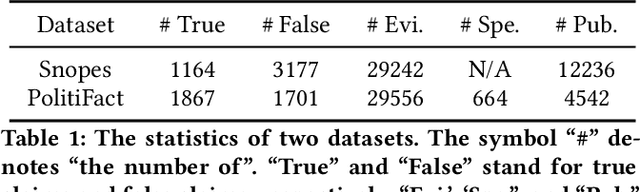
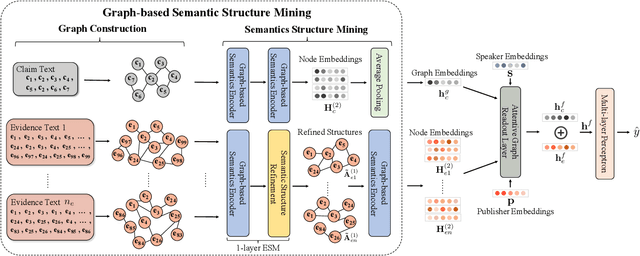
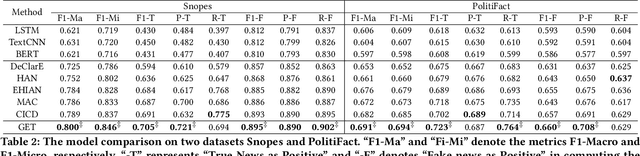
The prevalence and perniciousness of fake news has been a critical issue on the Internet, which stimulates the development of automatic fake news detection in turn. In this paper, we focus on the evidence-based fake news detection, where several evidences are utilized to probe the veracity of news (i.e., a claim). Most previous methods first employ sequential models to embed the semantic information and then capture the claim-evidence interaction based on different attention mechanisms. Despite their effectiveness, they still suffer from two main weaknesses. Firstly, due to the inherent drawbacks of sequential models, they fail to integrate the relevant information that is scattered far apart in evidences for veracity checking. Secondly, they neglect much redundant information contained in evidences that may be useless or even harmful. To solve these problems, we propose a unified Graph-based sEmantic sTructure mining framework, namely GET in short. Specifically, different from the existing work that treats claims and evidences as sequences, we model them as graph-structured data and capture the long-distance semantic dependency among dispersed relevant snippets via neighborhood propagation. After obtaining contextual semantic information, our model reduces information redundancy by performing graph structure learning. Finally, the fine-grained semantic representations are fed into the downstream claim-evidence interaction module for predictions. Comprehensive experiments have demonstrated the superiority of GET over the state-of-the-arts.
Gene Function Prediction with Gene Interaction Networks: A Context Graph Kernel Approach
Apr 22, 2022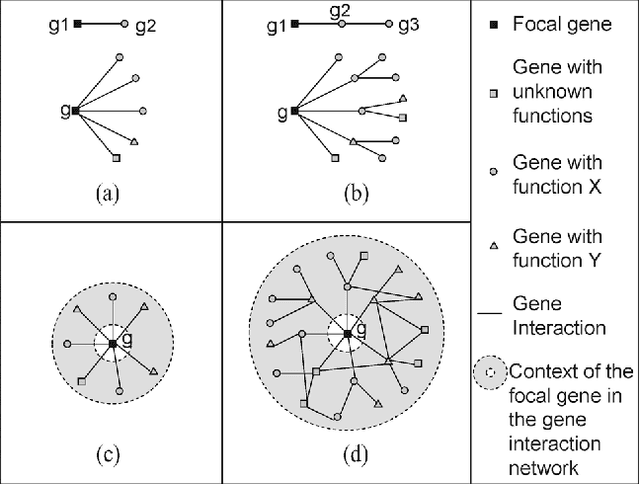
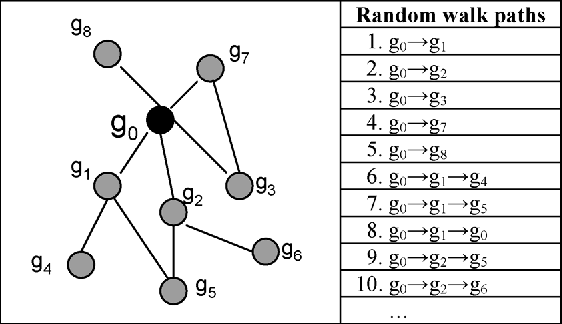
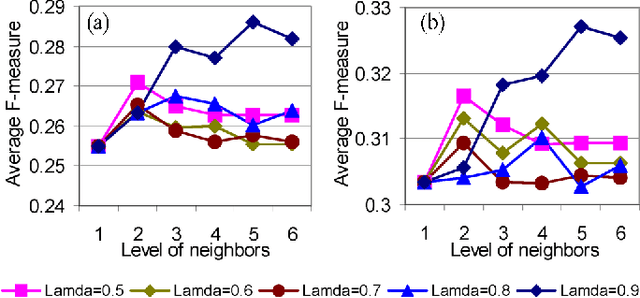
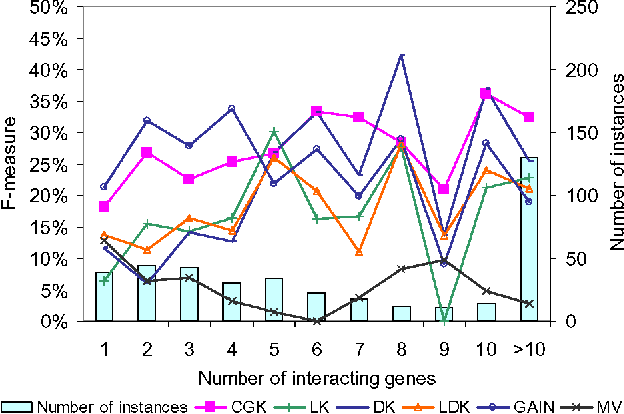
Predicting gene functions is a challenge for biologists in the post genomic era. Interactions among genes and their products compose networks that can be used to infer gene functions. Most previous studies adopt a linkage assumption, i.e., they assume that gene interactions indicate functional similarities between connected genes. In this study, we propose to use a gene's context graph, i.e., the gene interaction network associated with the focal gene, to infer its functions. In a kernel-based machine-learning framework, we design a context graph kernel to capture the information in context graphs. Our experimental study on a testbed of p53-related genes demonstrates the advantage of using indirect gene interactions and shows the empirical superiority of the proposed approach over linkage-assumption-based methods, such as the algorithm to minimize inconsistent connected genes and diffusion kernels.
Low Resolution Information Also Matters: Learning Multi-Resolution Representations for Person Re-Identification
May 26, 2021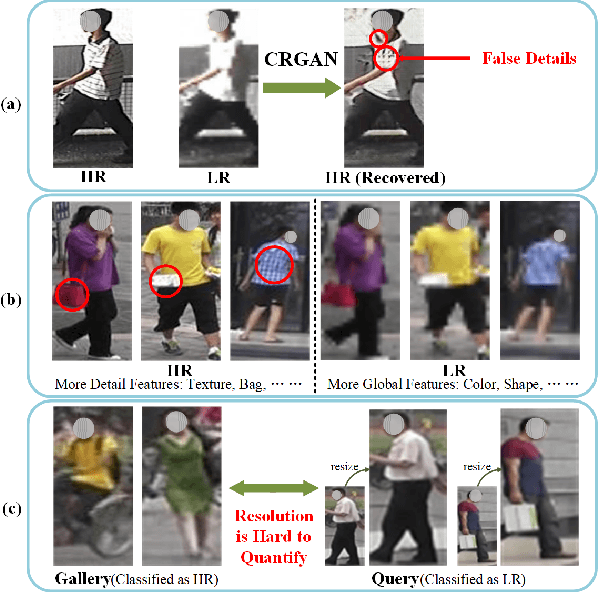
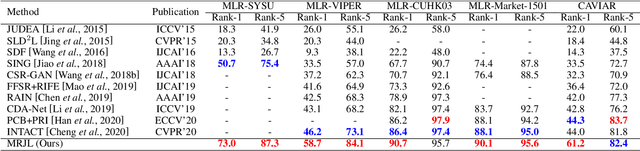
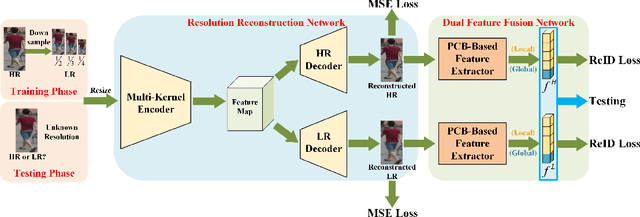
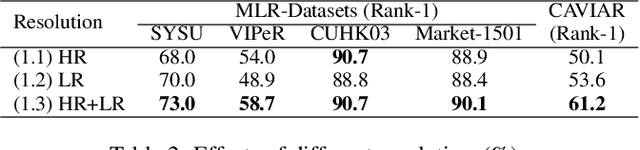
As a prevailing task in video surveillance and forensics field, person re-identification (re-ID) aims to match person images captured from non-overlapped cameras. In unconstrained scenarios, person images often suffer from the resolution mismatch problem, i.e., \emph{Cross-Resolution Person Re-ID}. To overcome this problem, most existing methods restore low resolution (LR) images to high resolution (HR) by super-resolution (SR). However, they only focus on the HR feature extraction and ignore the valid information from original LR images. In this work, we explore the influence of resolutions on feature extraction and develop a novel method for cross-resolution person re-ID called \emph{\textbf{M}ulti-Resolution \textbf{R}epresentations \textbf{J}oint \textbf{L}earning} (\textbf{MRJL}). Our method consists of a Resolution Reconstruction Network (RRN) and a Dual Feature Fusion Network (DFFN). The RRN uses an input image to construct a HR version and a LR version with an encoder and two decoders, while the DFFN adopts a dual-branch structure to generate person representations from multi-resolution images. Comprehensive experiments on five benchmarks verify the superiority of the proposed MRJL over the relevent state-of-the-art methods.
Information Content in Neuronal Calcium Spike Trains: Entropy Rate Estimation based on Empirical Probabilities
Feb 01, 2021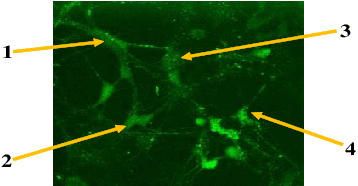
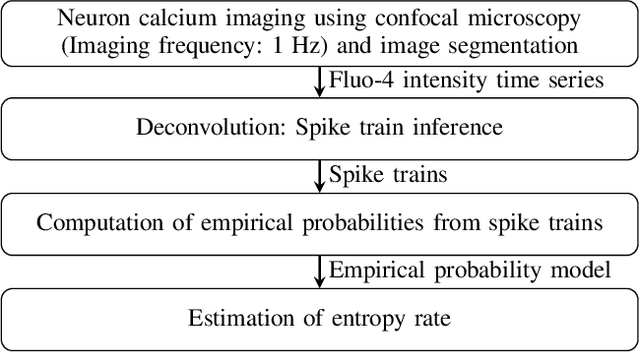

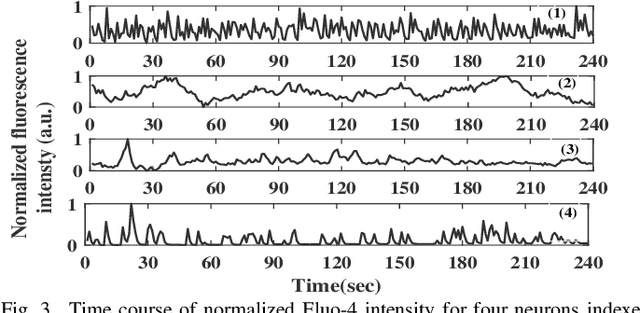
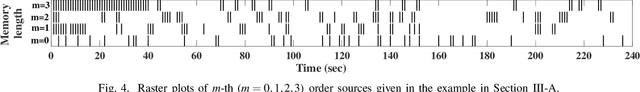
Quantification of information content and its temporal variation in intracellular calcium spike trains in neurons helps one understand functions such as memory, learning, and cognition. Such quantification could also reveal pathological signaling perturbation that potentially leads to devastating neurodegenerative conditions including Parkinson's, Alzheimer's, and Huntington's diseases. Accordingly, estimation of entropy rate, an information-theoretic measure of information content, assumes primary significance. However, such estimation in the present context is challenging because, while entropy rate is traditionally defined asymptotically for long blocks under the assumption of stationarity, neurons are known to encode information in short intervals and the associated spike trains often exhibit nonstationarity. Against this backdrop, we propose an entropy rate estimator based on empirical probabilities that operates within windows, short enough to ensure approximate stationarity. Specifically, our estimator, parameterized by the length of encoding contexts, attempts to model the underlying memory structures in neuronal spike trains. In an example Markov process, we compared the performance of the proposed method with that of versions of the Lempel-Ziv algorithm as well as with that of a certain stationary distribution method and found the former to exhibit higher accuracy levels and faster convergence. Also, in experimentally recorded calcium responses of four hippocampal neurons, the proposed method showed faster convergence. Significantly, our technique detected structural heterogeneity in the underlying process memory in the responses of the aforementioned neurons. We believe that the proposed method facilitates large-scale studies of such heterogeneity, which could in turn identify signatures of various diseases in terms of entropy rate estimates.
Towards a Comprehensive Solution for a Vision-based Digitized Neurological Examination
May 15, 2022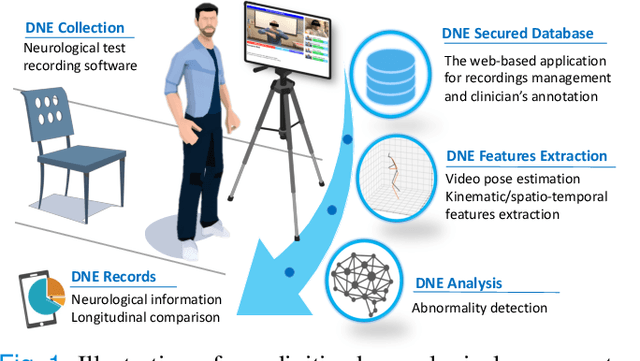
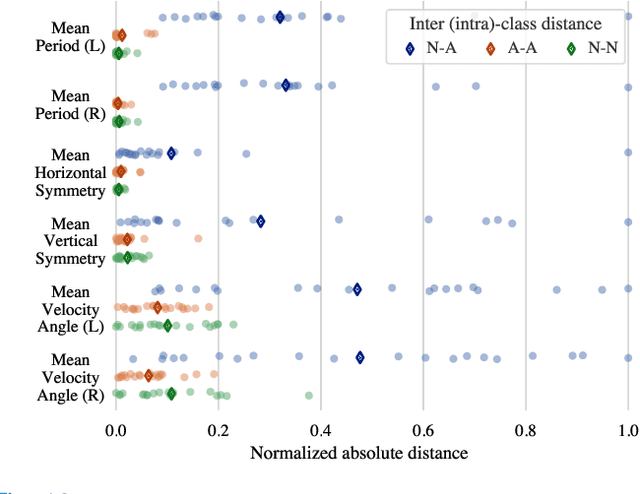
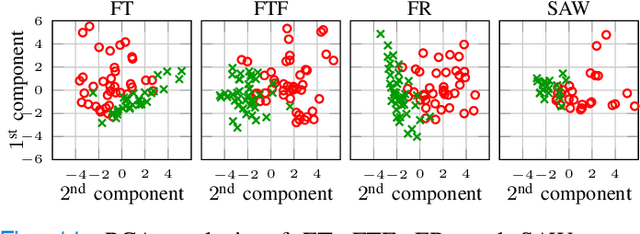
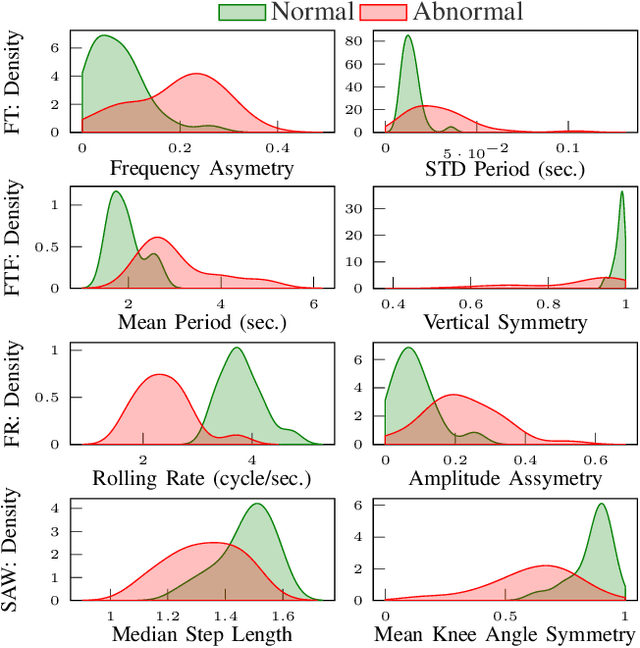
The ability to use digitally recorded and quantified neurological exam information is important to help healthcare systems deliver better care, in-person and via telehealth, as they compensate for a growing shortage of neurologists. Current neurological digital biomarker pipelines, however, are narrowed down to a specific neurological exam component or applied for assessing specific conditions. In this paper, we propose an accessible vision-based exam and documentation solution called Digitized Neurological Examination (DNE) to expand exam biomarker recording options and clinical applications using a smartphone/tablet. Through our DNE software, healthcare providers in clinical settings and people at home are enabled to video capture an examination while performing instructed neurological tests, including finger tapping, finger to finger, forearm roll, and stand-up and walk. Our modular design of the DNE software supports integrations of additional tests. The DNE extracts from the recorded examinations the 2D/3D human-body pose and quantifies kinematic and spatio-temporal features. The features are clinically relevant and allow clinicians to document and observe the quantified movements and the changes of these metrics over time. A web server and a user interface for recordings viewing and feature visualizations are available. DNE was evaluated on a collected dataset of 21 subjects containing normal and simulated-impaired movements. The overall accuracy of DNE is demonstrated by classifying the recorded movements using various machine learning models. Our tests show an accuracy beyond 90% for upper-limb tests and 80% for the stand-up and walk tests.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge