"Information": models, code, and papers
Exploring Dual Encoder Architectures for Question Answering
Apr 14, 2022
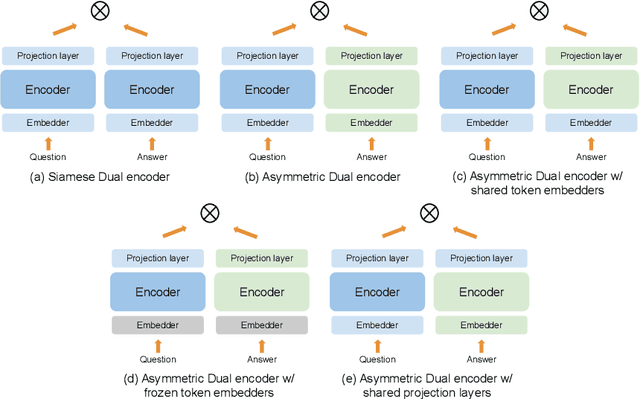
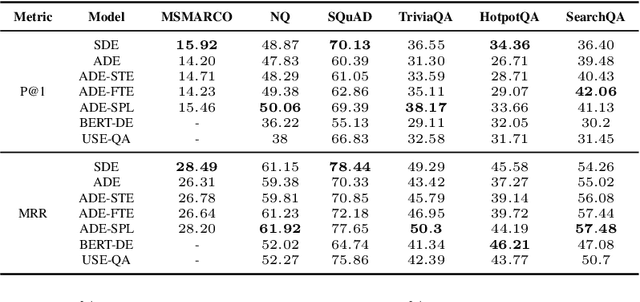
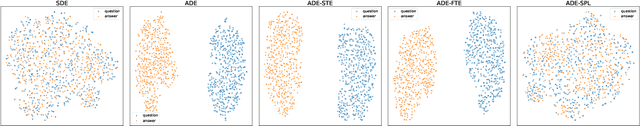
Dual encoders have been used for question-answering (QA) and information retrieval (IR) tasks with good results. There are two major types of dual encoders, Siamese Dual Encoders (SDE), with parameters shared across two encoders, and Asymmetric Dual Encoder (ADE), with two distinctly parameterized encoders. In this work, we explore the dual encoder architectures for QA retrieval tasks. By evaluating on MS MARCO and the MultiReQA benchmark, we show that SDE performs significantly better than ADE. We further propose three different improved versions of ADEs. Based on the evaluation of QA retrieval tasks and direct analysis of the embeddings, we demonstrate that sharing parameters in projection layers would enable ADEs to perform competitively with SDEs.
Boosting Semi-supervised Image Segmentation with Global and Local Mutual Information Regularization
Mar 08, 2021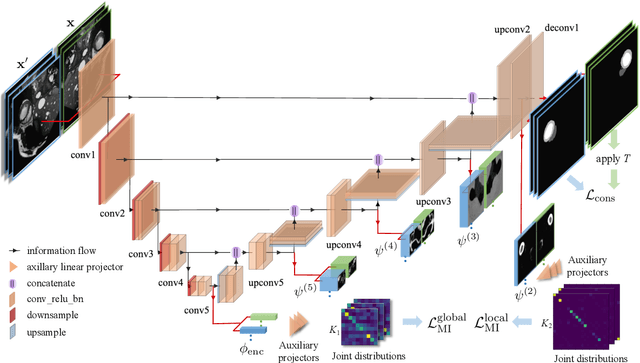
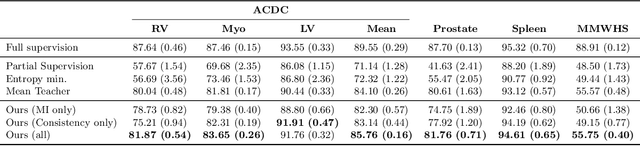
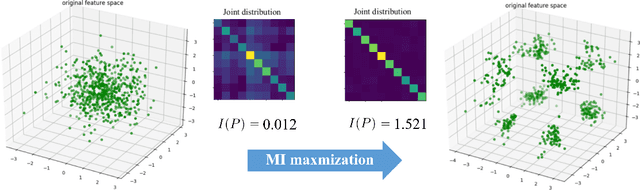
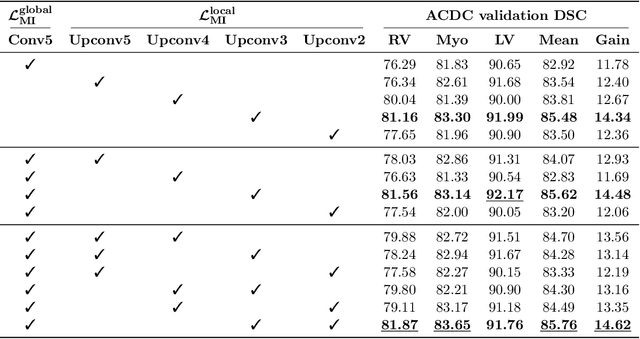
The scarcity of labeled data often impedes the application of deep learning to the segmentation of medical images. Semi-supervised learning seeks to overcome this limitation by leveraging unlabeled examples in the learning process. In this paper, we present a novel semi-supervised segmentation method that leverages mutual information (MI) on categorical distributions to achieve both global representation invariance and local smoothness. In this method, we maximize the MI for intermediate feature embeddings that are taken from both the encoder and decoder of a segmentation network. We first propose a global MI loss constraining the encoder to learn an image representation that is invariant to geometric transformations. Instead of resorting to computationally-expensive techniques for estimating the MI on continuous feature embeddings, we use projection heads to map them to a discrete cluster assignment where MI can be computed efficiently. Our method also includes a local MI loss to promote spatial consistency in the feature maps of the decoder and provide a smoother segmentation. Since mutual information does not require a strict ordering of clusters in two different assignments, we incorporate a final consistency regularization loss on the output which helps align the cluster labels throughout the network. We evaluate the method on three challenging publicly-available datasets for medical image segmentation. Experimental results show our method to outperform recently-proposed approaches for semi-supervised segmentation and provide an accuracy near to full supervision while training with very few annotated images
Multi-modal Graph Learning for Disease Prediction
Mar 11, 2022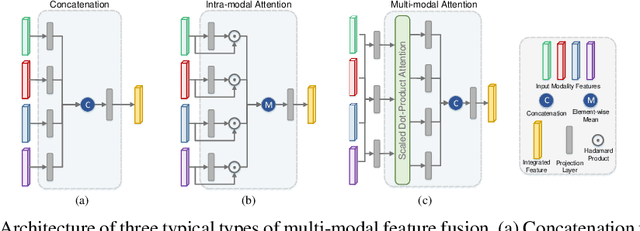
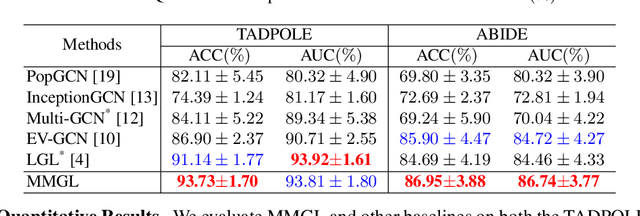
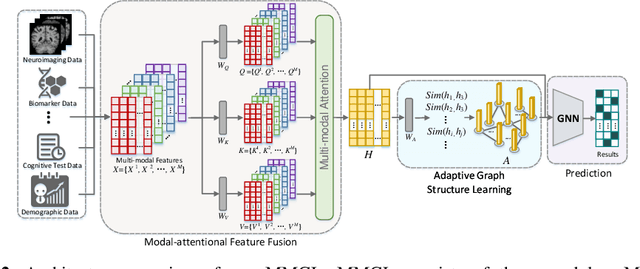
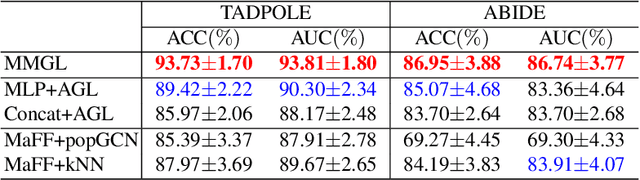
Benefiting from the powerful expressive capability of graphs, graph-based approaches have been popularly applied to handle multi-modal medical data and achieved impressive performance in various biomedical applications. For disease prediction tasks, most existing graph-based methods tend to define the graph manually based on specified modality (e.g., demographic information), and then integrated other modalities to obtain the patient representation by Graph Representation Learning (GRL). However, constructing an appropriate graph in advance is not a simple matter for these methods. Meanwhile, the complex correlation between modalities is ignored. These factors inevitably yield the inadequacy of providing sufficient information about the patient's condition for a reliable diagnosis. To this end, we propose an end-to-end Multi-modal Graph Learning framework (MMGL) for disease prediction with multi-modality. To effectively exploit the rich information across multi-modality associated with the disease, modality-aware representation learning is proposed to aggregate the features of each modality by leveraging the correlation and complementarity between the modalities. Furthermore, instead of defining the graph manually, the latent graph structure is captured through an effective way of adaptive graph learning. It could be jointly optimized with the prediction model, thus revealing the intrinsic connections among samples. Our model is also applicable to the scenario of inductive learning for those unseen data. An extensive group of experiments on two disease prediction tasks demonstrates that the proposed MMGL achieves more favorable performance. The code of MMGL is available at \url{https://github.com/SsGood/MMGL}.
Rate-Distortion Theoretic Generalization Bounds for Stochastic Learning Algorithms
Mar 04, 2022Understanding generalization in modern machine learning settings has been one of the major challenges in statistical learning theory. In this context, recent years have witnessed the development of various generalization bounds suggesting different complexity notions such as the mutual information between the data sample and the algorithm output, compressibility of the hypothesis space, and the fractal dimension of the hypothesis space. While these bounds have illuminated the problem at hand from different angles, their suggested complexity notions might appear seemingly unrelated, thereby restricting their high-level impact. In this study, we prove novel generalization bounds through the lens of rate-distortion theory, and explicitly relate the concepts of mutual information, compressibility, and fractal dimensions in a single mathematical framework. Our approach consists of (i) defining a generalized notion of compressibility by using source coding concepts, and (ii) showing that the `compression error rate' can be linked to the generalization error both in expectation and with high probability. We show that in the `lossless compression' setting, we recover and improve existing mutual information-based bounds, whereas a `lossy compression' scheme allows us to link generalization to the rate-distortion dimension -- a particular notion of fractal dimension. Our results bring a more unified perspective on generalization and open up several future research directions.
Conditional $β$-VAE for De Novo Molecular Generation
May 01, 2022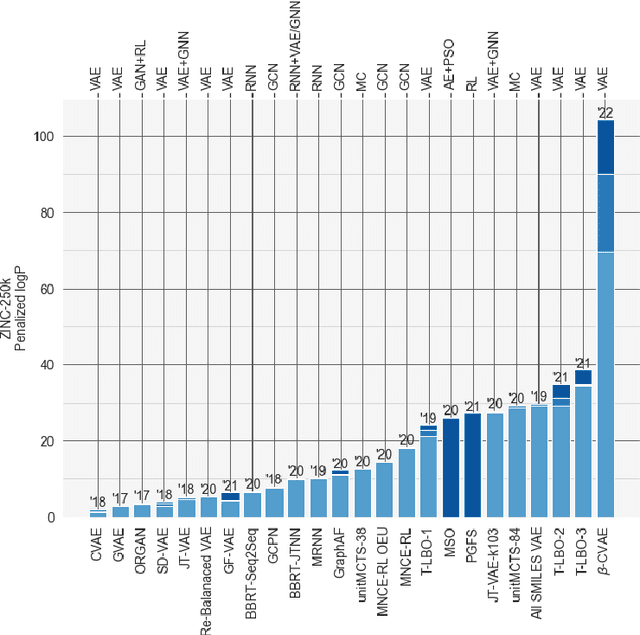
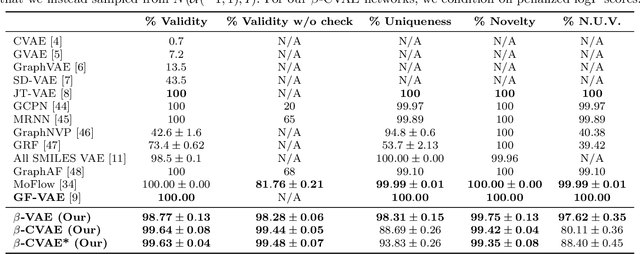
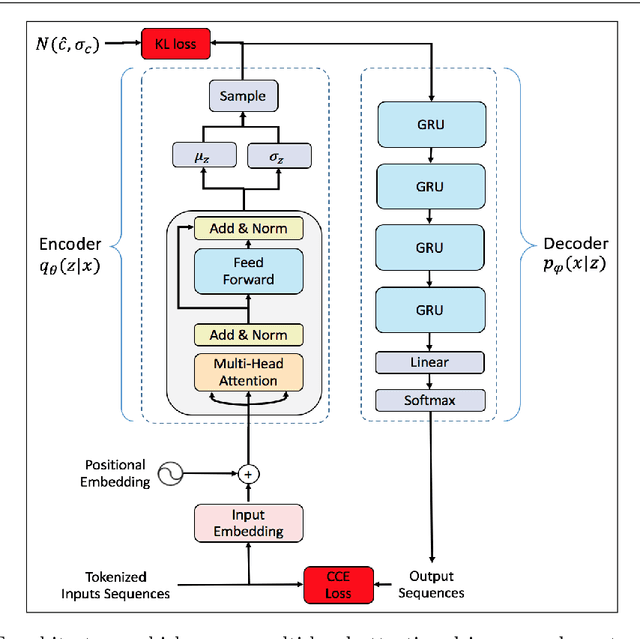
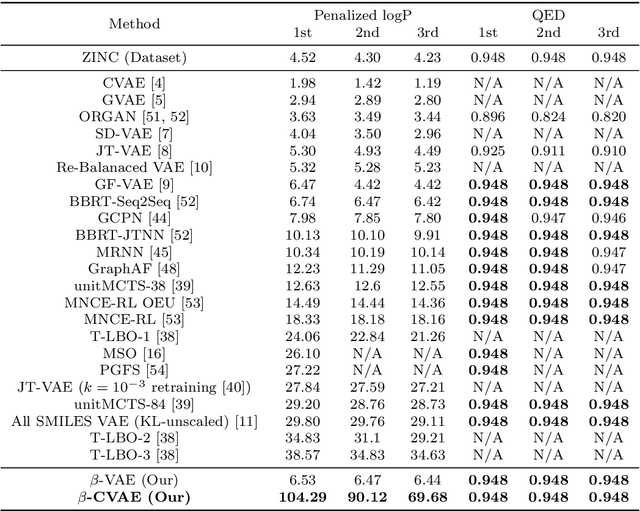
Deep learning has significantly advanced and accelerated de novo molecular generation. Generative networks, namely Variational Autoencoders (VAEs) can not only randomly generate new molecules, but also alter molecular structures to optimize specific chemical properties which are pivotal for drug-discovery. While VAEs have been proposed and researched in the past for pharmaceutical applications, they possess deficiencies which limit their ability to both optimize properties and decode syntactically valid molecules. We present a recurrent, conditional $\beta$-VAE which disentangles the latent space to enhance post hoc molecule optimization. We create a mutual information driven training protocol and data augmentations to both increase molecular validity and promote longer sequence generation. We demonstrate the efficacy of our framework on the ZINC-250k dataset, achieving SOTA unconstrained optimization results on the penalized LogP (pLogP) and QED scores, while also matching current SOTA results for validity, novelty and uniqueness scores for random generation. We match the current SOTA on QED for top-3 molecules at 0.948, while setting a new SOTA for pLogP optimization at 104.29, 90.12, 69.68 and demonstrating improved results on the constrained optimization task.
CAR: Class-aware Regularizations for Semantic Segmentation
Mar 14, 2022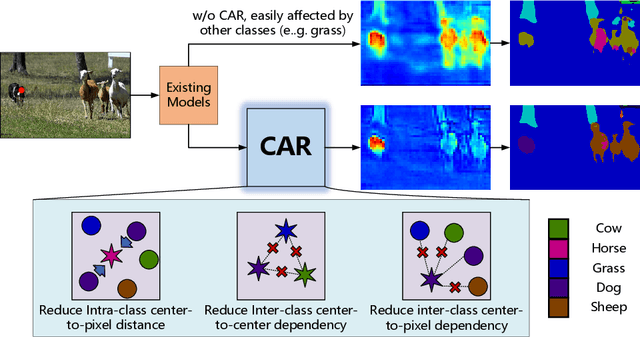
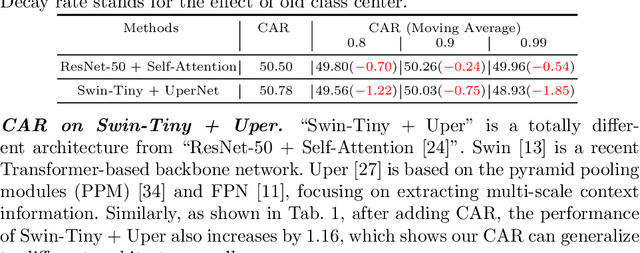
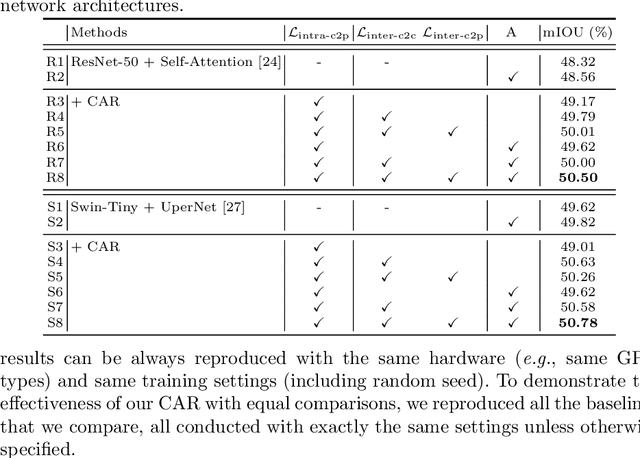
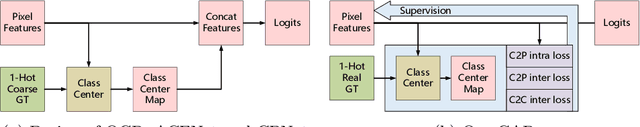

Recent segmentation methods, such as OCR and CPNet, utilizing "class level" information in addition to pixel features, have achieved notable success for boosting the accuracy of existing network modules. However, the extracted class-level information was simply concatenated to pixel features, without explicitly being exploited for better pixel representation learning. Moreover, these approaches learn soft class centers based on coarse mask prediction, which is prone to error accumulation. In this paper, aiming to use class level information more effectively, we propose a universal Class-Aware Regularization (CAR) approach to optimize the intra-class variance and inter-class distance during feature learning, motivated by the fact that humans can recognize an object by itself no matter which other objects it appears with. Three novel loss functions are proposed. The first loss function encourages more compact class representations within each class, the second directly maximizes the distance between different class centers, and the third further pushes the distance between inter-class centers and pixels. Furthermore, the class center in our approach is directly generated from ground truth instead of from the error-prone coarse prediction. Our method can be easily applied to most existing segmentation models during training, including OCR and CPNet, and can largely improve their accuracy at no additional inference overhead. Extensive experiments and ablation studies conducted on multiple benchmark datasets demonstrate that the proposed CAR can boost the accuracy of all baseline models by up to 2.23% mIOU with superior generalization ability. The complete code is available at https://github.com/edwardyehuang/CAR.
Online Graph Learning from Social Interactions
Mar 11, 2022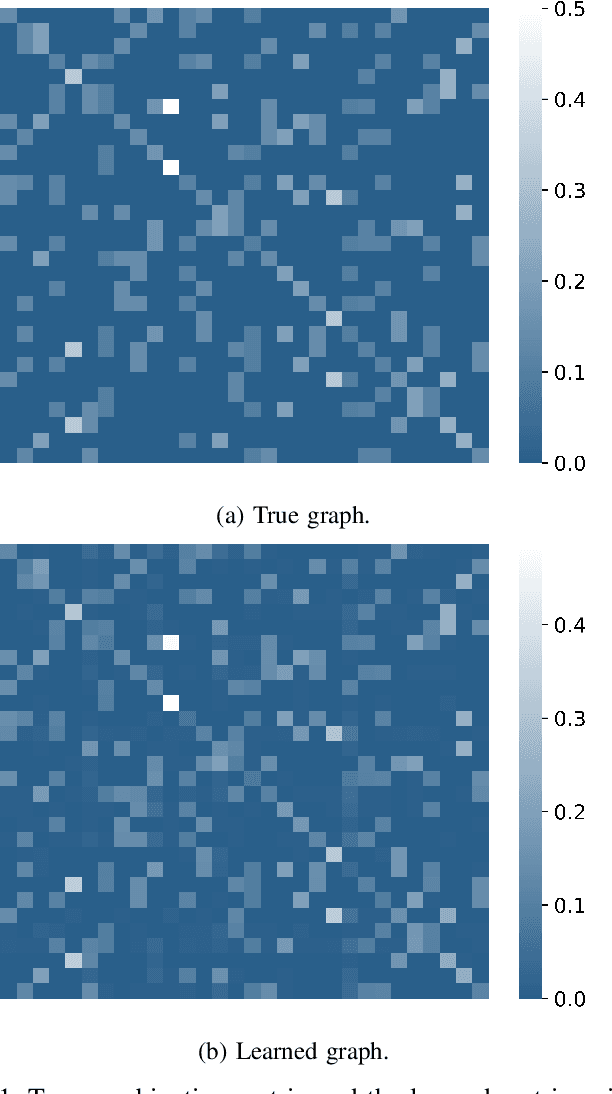
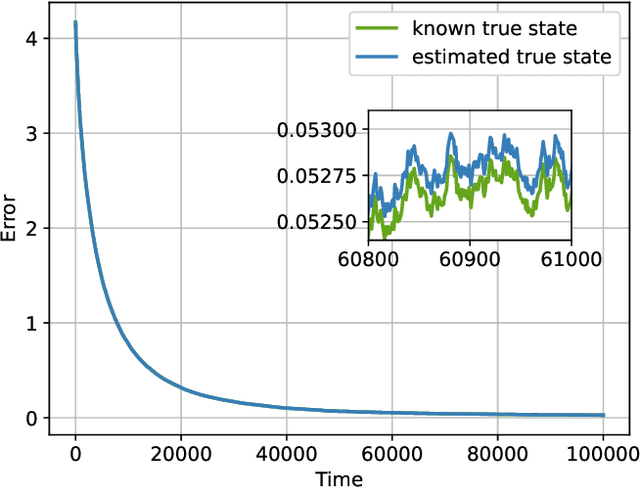
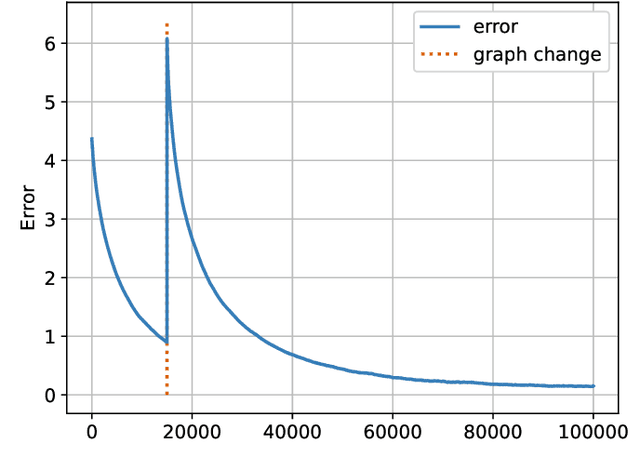
Social learning algorithms provide models for the formation of opinions over social networks resulting from local reasoning and peer-to-peer exchanges. Interactions occur over an underlying graph topology, which describes the flow of information and relative influence between pairs of agents. For a given graph topology, these algorithms allow for the prediction of formed opinions. In this work, we study the inverse problem. Given a social learning model and observations of the evolution of beliefs over time, we aim at identifying the underlying graph topology. The learned graph allows for the inference of pairwise influence between agents, the overall influence agents have over the behavior of the network, as well as the flow of information through the social network. The proposed algorithm is online in nature and can adapt dynamically to changes in the graph topology or the true hypothesis.
A Summary of the ALQAC 2021 Competition
Apr 25, 2022
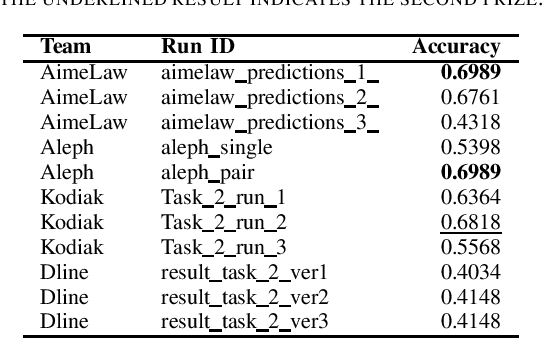
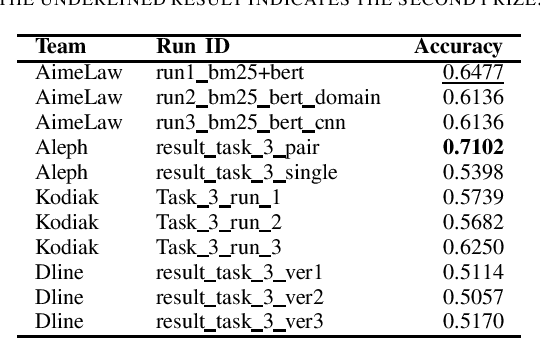


We summarize the evaluation of the first Automated Legal Question Answering Competition (ALQAC 2021). The competition this year contains three tasks, which aims at processing the statute law document, which are Legal Text Information Retrieval (Task 1), Legal Text Entailment Prediction (Task 2), and Legal Text Question Answering (Task 3). The final goal of these tasks is to build a system that can automatically determine whether a particular statement is lawful. There is no limit to the approaches of the participating teams. This year, there are 5 teams participating in Task 1, 6 teams participating in Task 2, and 5 teams participating in Task 3. There are in total 36 runs submitted to the organizer. In this paper, we summarize each team's approaches, official results, and some discussion about the competition. Only results of the teams who successfully submit their approach description paper are reported in this paper.
NeuralTree: A 256-Channel 0.227uJ/class Versatile Neural Activity Classification and Closed-Loop Neuromodulation SoC
May 21, 2022



Closed-loop neural interfaces with on-chip machine learning can detect and suppress disease symptoms in neurological disorders or restore lost functions in paralyzed patients. While high-density neural recording can provide rich neural activity information for accurate disease-state detection, existing systems have low channel count and poor scalability, which could limit their therapeutic efficacy. This work presents a highly scalable and versatile closed-loop neural interface SoC that can overcome these limitations. A 256-channel time-division multiplexed (TDM) front-end with a two-step fast-settling mixed-signal DC servo loop (DSL) is proposed to record high-spatial-resolution neural activity and perform channel-selective brain-state inference. A tree-structured neural network (NeuralTree) classification processor extracts a rich set of neural biomarkers in a patient- and disease-specific manner. Trained with an energy-aware learning algorithm, the NeuralTree classifier detects the symptoms of underlying disorders (e.g., epilepsy and movement disorders) at an optimal energy-accuracy trade-off. A 16-channel high-voltage (HV) compliant neurostimulator closes the therapeutic loop by delivering charge-balanced biphasic current pulses to the brain. The proposed SoC was fabricated in 65nm CMOS and achieved a 0.227uJ/class energy efficiency in a compact area of 0.014mm^2/channel. The SoC was extensively verified on human electroencephalography (EEG) and intracranial EEG (iEEG) epilepsy datasets, obtaining 95.6%/94% sensitivity and 96.8%/96.9% specificity, respectively. In-vivo neural recordings using soft uECoG arrays and multi-domain biomarker extraction were further performed on a rat model of epilepsy. In addition, for the first time in literature, on-chip classification of rest-state tremor in Parkinson's disease from human local field potentials (LFPs) was demonstrated.
Information Content in Neuronal Calcium Spike Trains: Entropy Rate Estimation based on Empirical Probabilities
Feb 01, 2021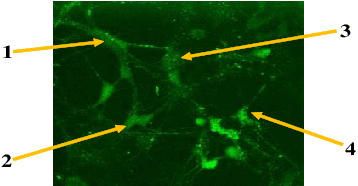
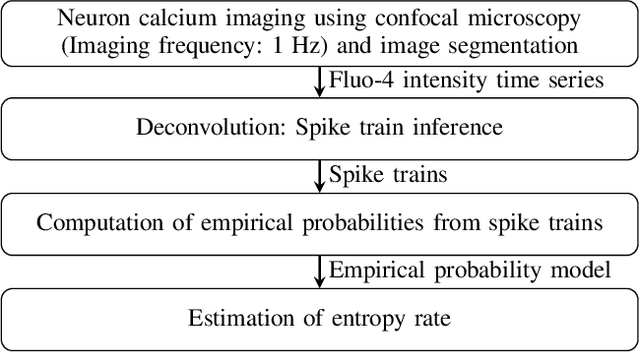

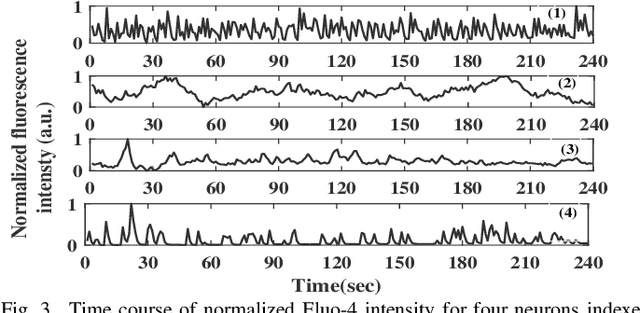
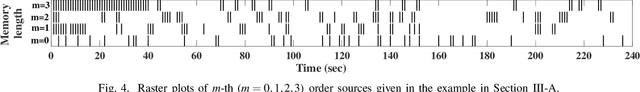
Quantification of information content and its temporal variation in intracellular calcium spike trains in neurons helps one understand functions such as memory, learning, and cognition. Such quantification could also reveal pathological signaling perturbation that potentially leads to devastating neurodegenerative conditions including Parkinson's, Alzheimer's, and Huntington's diseases. Accordingly, estimation of entropy rate, an information-theoretic measure of information content, assumes primary significance. However, such estimation in the present context is challenging because, while entropy rate is traditionally defined asymptotically for long blocks under the assumption of stationarity, neurons are known to encode information in short intervals and the associated spike trains often exhibit nonstationarity. Against this backdrop, we propose an entropy rate estimator based on empirical probabilities that operates within windows, short enough to ensure approximate stationarity. Specifically, our estimator, parameterized by the length of encoding contexts, attempts to model the underlying memory structures in neuronal spike trains. In an example Markov process, we compared the performance of the proposed method with that of versions of the Lempel-Ziv algorithm as well as with that of a certain stationary distribution method and found the former to exhibit higher accuracy levels and faster convergence. Also, in experimentally recorded calcium responses of four hippocampal neurons, the proposed method showed faster convergence. Significantly, our technique detected structural heterogeneity in the underlying process memory in the responses of the aforementioned neurons. We believe that the proposed method facilitates large-scale studies of such heterogeneity, which could in turn identify signatures of various diseases in terms of entropy rate estimates.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge