"Information": models, code, and papers
Decomposed Mutual Information Estimation for Contrastive Representation Learning
Jun 25, 2021

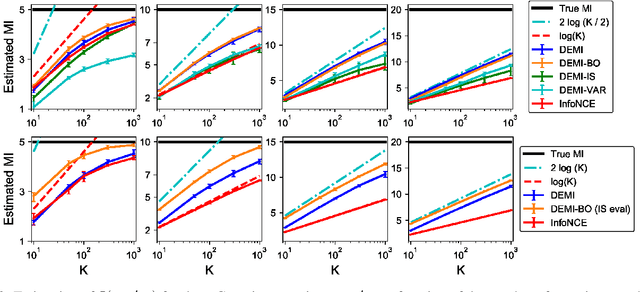
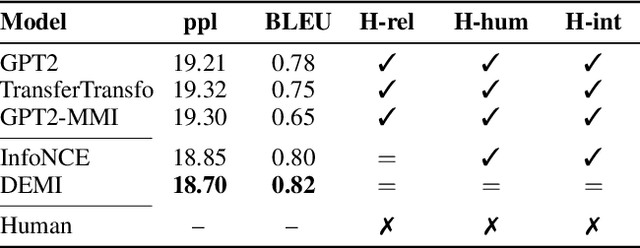
Recent contrastive representation learning methods rely on estimating mutual information (MI) between multiple views of an underlying context. E.g., we can derive multiple views of a given image by applying data augmentation, or we can split a sequence into views comprising the past and future of some step in the sequence. Contrastive lower bounds on MI are easy to optimize, but have a strong underestimation bias when estimating large amounts of MI. We propose decomposing the full MI estimation problem into a sum of smaller estimation problems by splitting one of the views into progressively more informed subviews and by applying the chain rule on MI between the decomposed views. This expression contains a sum of unconditional and conditional MI terms, each measuring modest chunks of the total MI, which facilitates approximation via contrastive bounds. To maximize the sum, we formulate a contrastive lower bound on the conditional MI which can be approximated efficiently. We refer to our general approach as Decomposed Estimation of Mutual Information (DEMI). We show that DEMI can capture a larger amount of MI than standard non-decomposed contrastive bounds in a synthetic setting, and learns better representations in a vision domain and for dialogue generation.
Adaptive GLCM sampling for transformer-based COVID-19 detection on CT
Jul 04, 2022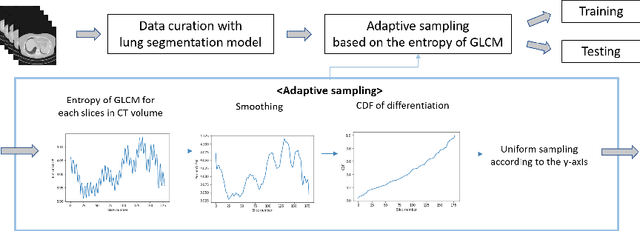
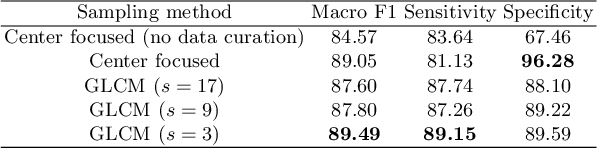
The world has suffered from COVID-19 (SARS-CoV-2) for the last two years, causing much damage and change in people's daily lives. Thus, automated detection of COVID-19 utilizing deep learning on chest computed tomography (CT) scans became promising, which helps correct diagnosis efficiently. Recently, transformer-based COVID-19 detection method on CT is proposed to utilize 3D information in CT volume. However, its sampling method for selecting slices is not optimal. To leverage rich 3D information in CT volume, we propose a transformer-based COVID-19 detection using a novel data curation and adaptive sampling method using gray level co-occurrence matrices (GLCM). To train the model which consists of CNN layer, followed by transformer architecture, we first executed data curation based on lung segmentation and utilized the entropy of GLCM value of every slice in CT volumes to select important slices for the prediction. The experimental results show that the proposed method improve the detection performance with large margin without much difficult modification to the model.
An Efficient Multi-Scale Fusion Network for 3D Organ at Risk (OAR) Segmentation
Aug 15, 2022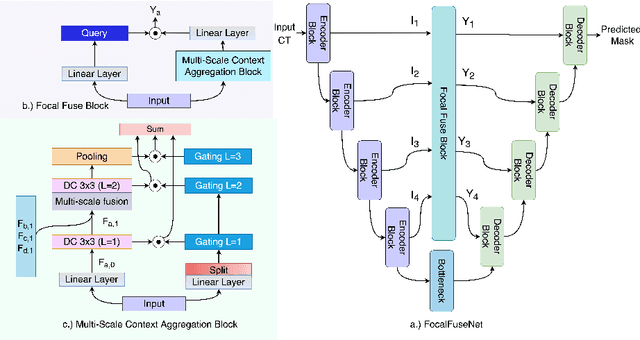

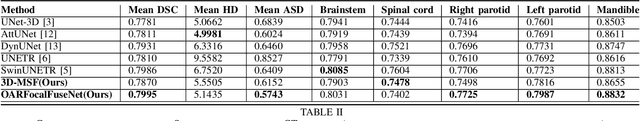
Accurate segmentation of organs-at-risks (OARs) is a precursor for optimizing radiation therapy planning. Existing deep learning-based multi-scale fusion architectures have demonstrated a tremendous capacity for 2D medical image segmentation. The key to their success is aggregating global context and maintaining high resolution representations. However, when translated into 3D segmentation problems, existing multi-scale fusion architectures might underperform due to their heavy computation overhead and substantial data diet. To address this issue, we propose a new OAR segmentation framework, called OARFocalFuseNet, which fuses multi-scale features and employs focal modulation for capturing global-local context across multiple scales. Each resolution stream is enriched with features from different resolution scales, and multi-scale information is aggregated to model diverse contextual ranges. As a result, feature representations are further boosted. The comprehensive comparisons in our experimental setup with OAR segmentation as well as multi-organ segmentation show that our proposed OARFocalFuseNet outperforms the recent state-of-the-art methods on publicly available OpenKBP datasets and Synapse multi-organ segmentation. Both of the proposed methods (3D-MSF and OARFocalFuseNet) showed promising performance in terms of standard evaluation metrics. Our best performing method (OARFocalFuseNet) obtained a dice coefficient of 0.7995 and hausdorff distance of 5.1435 on OpenKBP datasets and dice coefficient of 0.8137 on Synapse multi-organ segmentation dataset.
MIR-VIO: Mutual Information Residual-based Visual Inertial Odometry with UWB Fusion for Robust Localization
Sep 08, 2021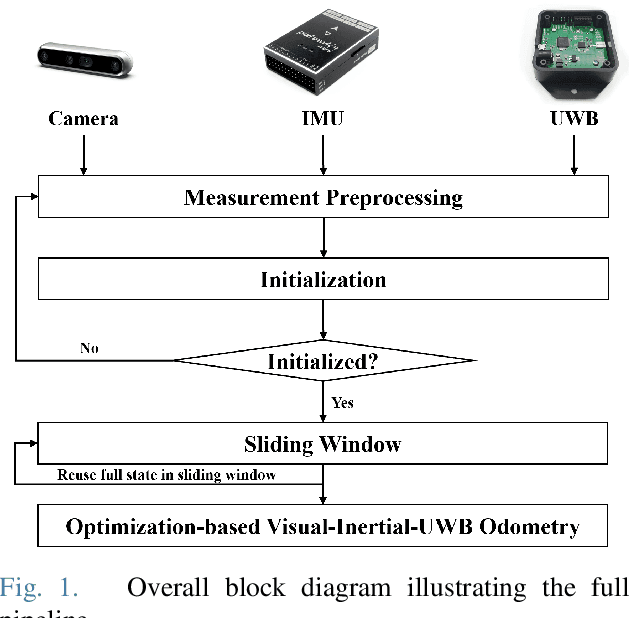
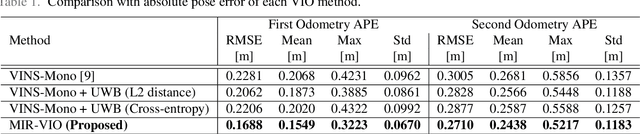
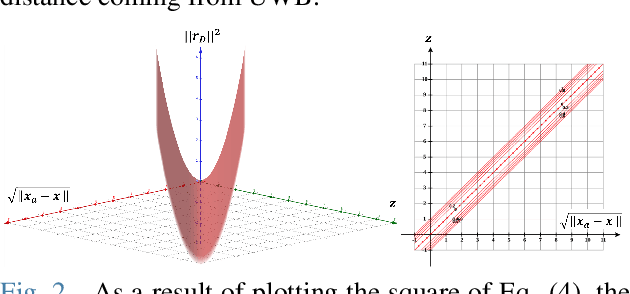
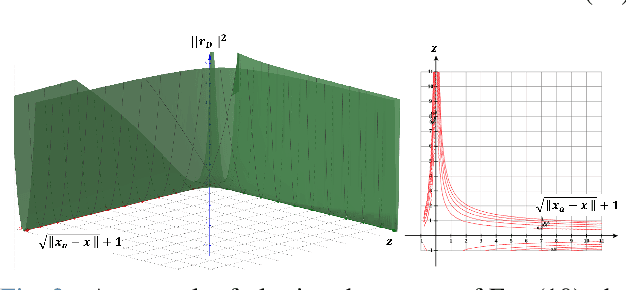
For many years, there has been an impressive progress on visual odometry applied to mobile robots and drones. However, the visual perception is still in the spotlight as a challenging field because the vision sensor has some problems in obtaining correct scale information with a monocular camera and also is vulnerable to a situation in which illumination is changed. In this paper, UWB sensor fusion is proposed in the visual inertial odometry algorithm as a solution to mitigate this problem. We designed a cost function based on mutual information considering the UWB. Considering the characteristic of the UWB signal model, where the uncertainty increases as the distance between the UWB anchor and the tag increases, we introduced a new residual term to the cost function. When the experiment was conducted in an indoor environment with the above methodology, the initialization problem in an environment with few feature points was solved through the UWB sensor fusion, and localization became robust. And when the residual term using the concept of mutual information was used, the most robust odometry could be obtained.
ChrSNet: Chromosome Straightening using Self-attention Guided Networks
Jul 01, 2022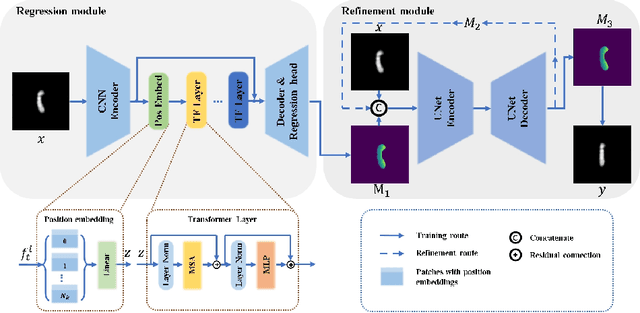
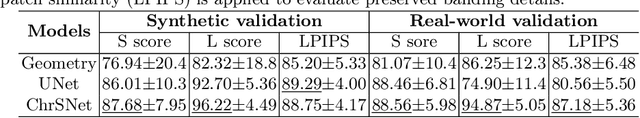
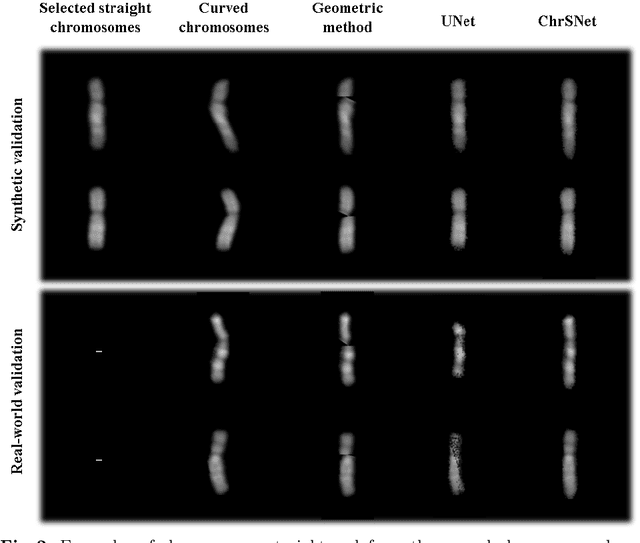
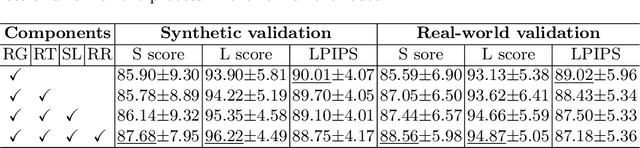
Karyotyping is an important procedure to assess the possible existence of chromosomal abnormalities. However, because of the non-rigid nature, chromosomes are usually heavily curved in microscopic images and such deformed shapes hinder the chromosome analysis for cytogeneticists. In this paper, we present a self-attention guided framework to erase the curvature of chromosomes. The proposed framework extracts spatial information and local textures to preserve banding patterns in a regression module. With complementary information from the bent chromosome, a refinement module is designed to further improve fine details. In addition, we propose two dedicated geometric constraints to maintain the length and restore the distortion of chromosomes. To train our framework, we create a synthetic dataset where curved chromosomes are generated from the real-world straight chromosomes by grid-deformation. Quantitative and qualitative experiments are conducted on synthetic and real-world data. Experimental results show that our proposed method can effectively straighten bent chromosomes while keeping banding details and length.
Highlight Specular Reflection Separation based on Tensor Low-rank and Sparse Decomposition Using Polarimetric Cues
Jul 07, 2022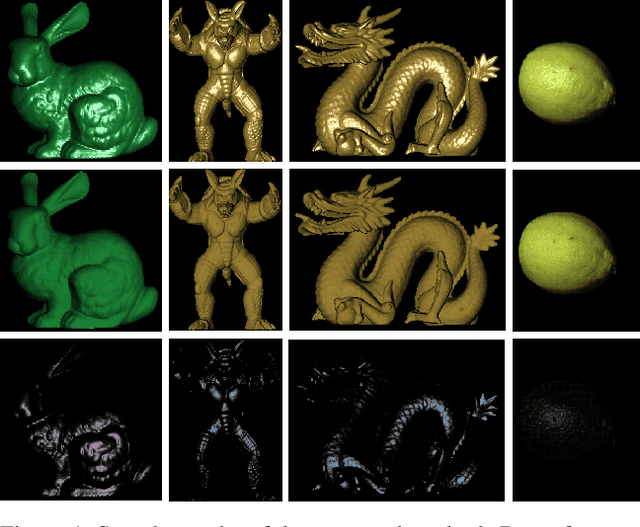
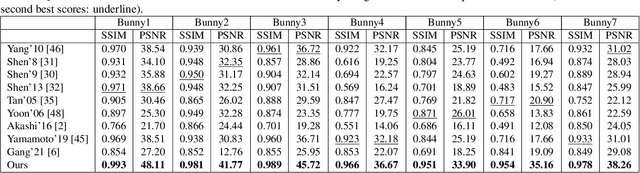
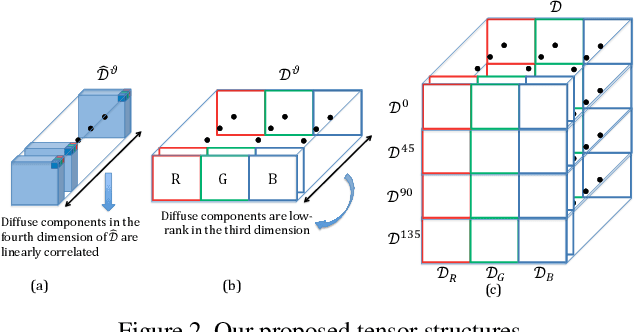
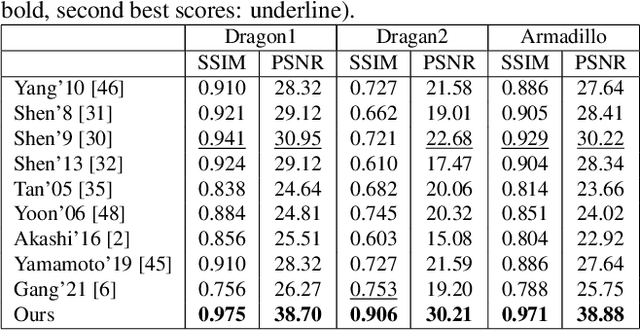
This paper is concerned with specular reflection removal based on tensor low-rank decomposition framework with the help of polarization information. Our method is motivated by the observation that the specular highlight of an image is sparsely distributed while the remaining diffuse reflection can be well approximated by a linear combination of several distinct colors using a low-rank and sparse decomposition framework. Unlike current solutions, our tensor low-rank decomposition keeps the spatial structure of specular and diffuse information which enables us to recover the diffuse image under strong specular reflection or in saturated regions. We further define and impose a new polarization regularization term as constraint on color channels. This regularization boosts the performance of the method to recover an accurate diffuse image by handling the color distortion, a common problem of chromaticity-based methods, especially in case of strong specular reflection. Through comprehensive experiments on both synthetic and real polarization images, we demonstrate that our method is able to significantly improve the accuracy of highlight specular removal, and outperform the competitive methods to recover the diffuse image, especially in regions of strong specular reflection or in saturated areas.
Smoothness Analysis for Probabilistic Programs with Application to Optimised Variational Inference
Aug 22, 2022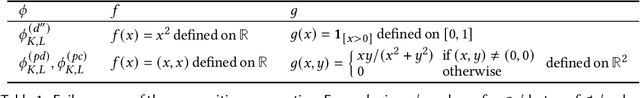

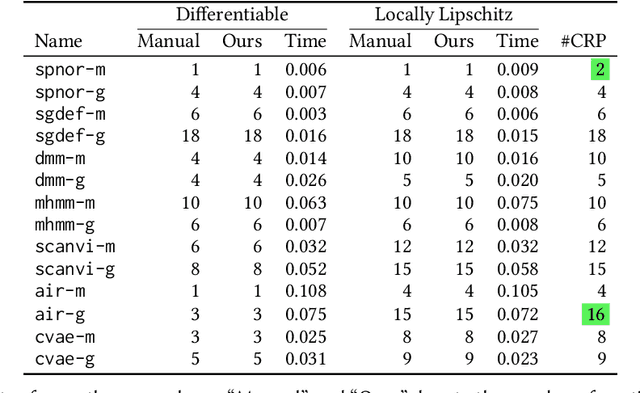
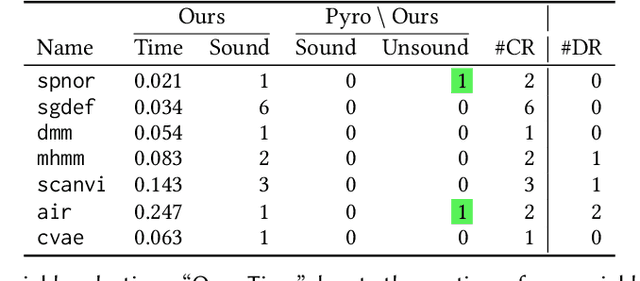
We present a static analysis for discovering differentiable or more generally smooth parts of a given probabilistic program, and show how the analysis can be used to improve the pathwise gradient estimator, one of the most popular methods for posterior inference and model learning. Our improvement increases the scope of the estimator from differentiable models to non-differentiable ones without requiring manual intervention of the user; the improved estimator automatically identifies differentiable parts of a given probabilistic program using our static analysis, and applies the pathwise gradient estimator to the identified parts while using a more general but less efficient estimator, called score estimator, for the rest of the program. Our analysis has a surprisingly subtle soundness argument, partly due to the misbehaviours of some target smoothness properties when viewed from the perspective of program analysis designers. For instance, some smoothness properties are not preserved by function composition, and this makes it difficult to analyse sequential composition soundly without heavily sacrificing precision. We formulate five assumptions on a target smoothness property, prove the soundness of our analysis under those assumptions, and show that our leading examples satisfy these assumptions. We also show that by using information from our analysis, our improved gradient estimator satisfies an important differentiability requirement and thus, under a mild regularity condition, computes the correct estimate on average, i.e., it returns an unbiased estimate. Our experiments with representative probabilistic programs in the Pyro language show that our static analysis is capable of identifying smooth parts of those programs accurately, and making our improved pathwise gradient estimator exploit all the opportunities for high performance in those programs.
PalQuant: Accelerating High-precision Networks on Low-precision Accelerators
Aug 03, 2022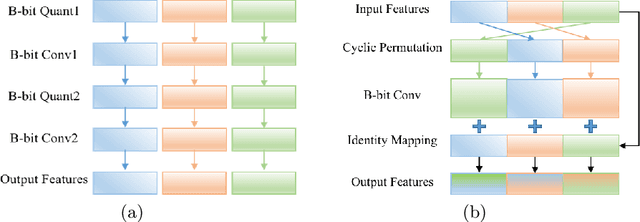
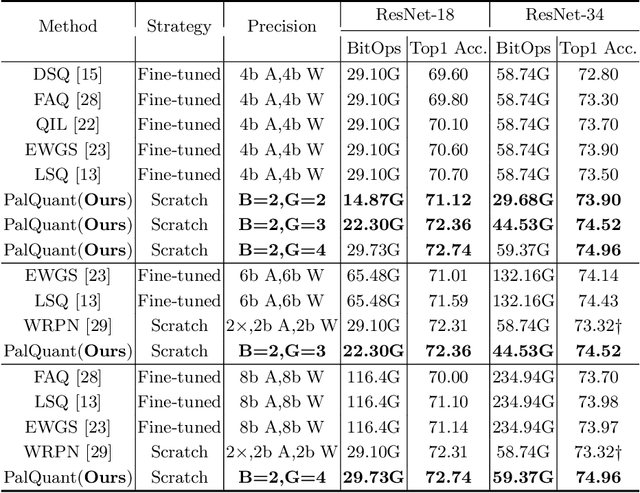
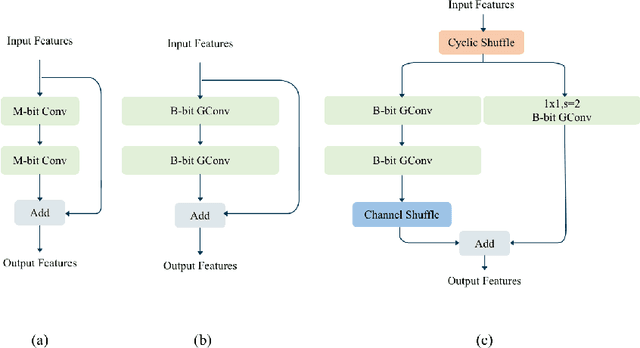
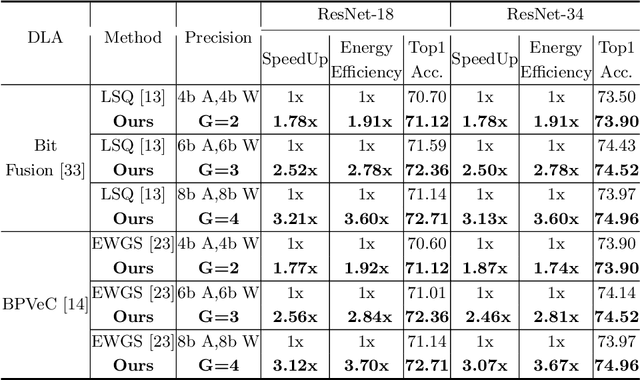
Recently low-precision deep learning accelerators (DLAs) have become popular due to their advantages in chip area and energy consumption, yet the low-precision quantized models on these DLAs bring in severe accuracy degradation. One way to achieve both high accuracy and efficient inference is to deploy high-precision neural networks on low-precision DLAs, which is rarely studied. In this paper, we propose the PArallel Low-precision Quantization (PalQuant) method that approximates high-precision computations via learning parallel low-precision representations from scratch. In addition, we present a novel cyclic shuffle module to boost the cross-group information communication between parallel low-precision groups. Extensive experiments demonstrate that PalQuant has superior performance to state-of-the-art quantization methods in both accuracy and inference speed, e.g., for ResNet-18 network quantization, PalQuant can obtain 0.52\% higher accuracy and 1.78$\times$ speedup simultaneously over their 4-bit counter-part on a state-of-the-art 2-bit accelerator. Code is available at \url{https://github.com/huqinghao/PalQuant}.
Developing an NLP-based Recommender System for the Ethical, Legal, and Social Implications of Synthetic Biology
Jul 10, 2022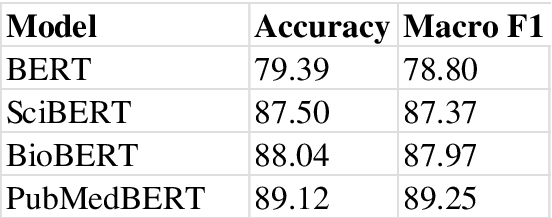
Synthetic biology is an emerging field that involves the engineering and re-design of organisms for purposes such as food security, health, and environmental protection. As such, it poses numerous ethical, legal, and social implications (ELSI) for researchers and policy makers. Various efforts to ensure socially responsible synthetic biology are underway. Policy making is one regulatory avenue, and other initiatives have sought to embed social scientists and ethicists on synthetic biology projects. However, given the nascency of synthetic biology, the number of heterogeneous domains it spans, and the open nature of many ethical questions, it has proven challenging to establish widespread concrete policies, and including social scientists and ethicists on synthetic biology teams has met with mixed success. This text proposes a different approach, asking instead is it possible to develop a well-performing recommender model based upon natural language processing (NLP) to connect synthetic biologists with information on the ELSI of their specific research? This recommender was developed as part of a larger project building a Synthetic Biology Knowledge System (SBKS) to accelerate discovery and exploration of the synthetic biology design space. Our approach aims to distill for synthetic biologists relevant ethical and social scientific information and embed it into synthetic biology research workflows.
TFCNs: A CNN-Transformer Hybrid Network for Medical Image Segmentation
Jul 07, 2022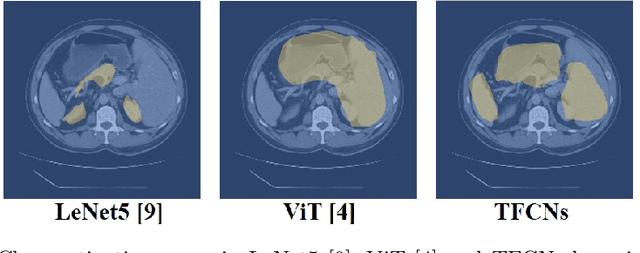
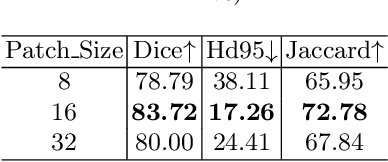
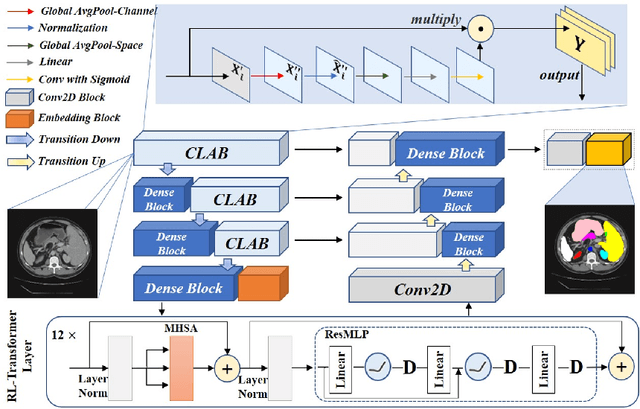
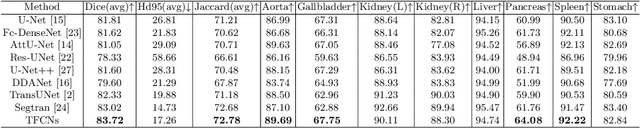
Medical image segmentation is one of the most fundamental tasks concerning medical information analysis. Various solutions have been proposed so far, including many deep learning-based techniques, such as U-Net, FC-DenseNet, etc. However, high-precision medical image segmentation remains a highly challenging task due to the existence of inherent magnification and distortion in medical images as well as the presence of lesions with similar density to normal tissues. In this paper, we propose TFCNs (Transformers for Fully Convolutional denseNets) to tackle the problem by introducing ResLinear-Transformer (RL-Transformer) and Convolutional Linear Attention Block (CLAB) to FC-DenseNet. TFCNs is not only able to utilize more latent information from the CT images for feature extraction, but also can capture and disseminate semantic features and filter non-semantic features more effectively through the CLAB module. Our experimental results show that TFCNs can achieve state-of-the-art performance with dice scores of 83.72\% on the Synapse dataset. In addition, we evaluate the robustness of TFCNs for lesion area effects on the COVID-19 public datasets. The Python code will be made publicly available on https://github.com/HUANGLIZI/TFCNs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge