"Information": models, code, and papers
Globally Gated Deep Linear Networks
Oct 31, 2022
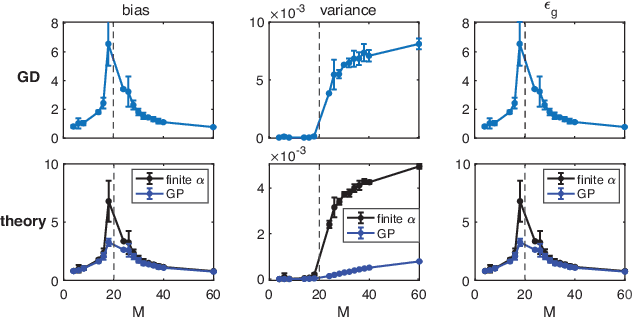
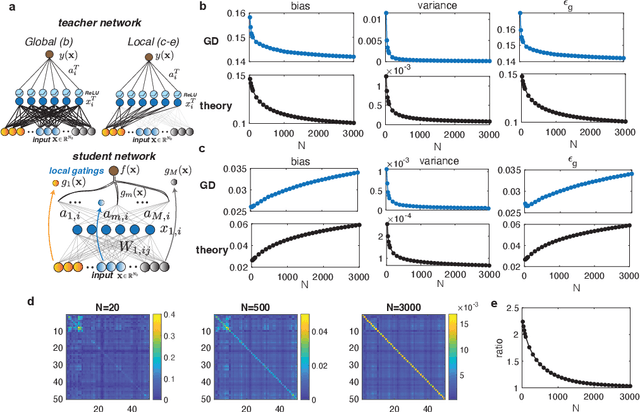
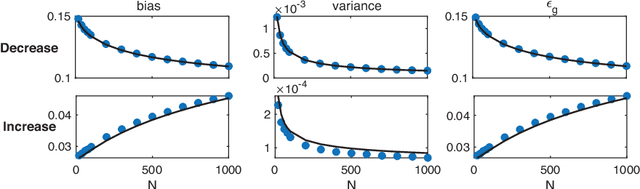
Recently proposed Gated Linear Networks present a tractable nonlinear network architecture, and exhibit interesting capabilities such as learning with local error signals and reduced forgetting in sequential learning. In this work, we introduce a novel gating architecture, named Globally Gated Deep Linear Networks (GGDLNs) where gating units are shared among all processing units in each layer, thereby decoupling the architectures of the nonlinear but unlearned gatings and the learned linear processing motifs. We derive exact equations for the generalization properties in these networks in the finite-width thermodynamic limit, defined by $P,N\rightarrow\infty, P/N\sim O(1)$, where P and N are the training sample size and the network width respectively. We find that the statistics of the network predictor can be expressed in terms of kernels that undergo shape renormalization through a data-dependent matrix compared to the GP kernels. Our theory accurately captures the behavior of finite width GGDLNs trained with gradient descent dynamics. We show that kernel shape renormalization gives rise to rich generalization properties w.r.t. network width, depth and L2 regularization amplitude. Interestingly, networks with sufficient gating units behave similarly to standard ReLU networks. Although gatings in the model do not participate in supervised learning, we show the utility of unsupervised learning of the gating parameters. Additionally, our theory allows the evaluation of the network's ability for learning multiple tasks by incorporating task-relevant information into the gating units. In summary, our work is the first exact theoretical solution of learning in a family of nonlinear networks with finite width. The rich and diverse behavior of the GGDLNs suggests that they are helpful analytically tractable models of learning single and multiple tasks, in finite-width nonlinear deep networks.
Learning Multi-Modal Brain Tumor Segmentation from Privileged Semi-Paired MRI Images with Curriculum Disentanglement Learning
Aug 26, 2022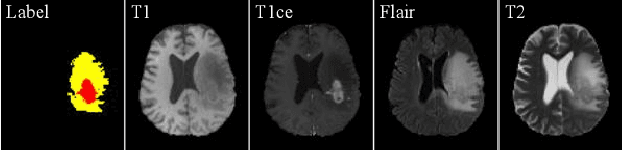
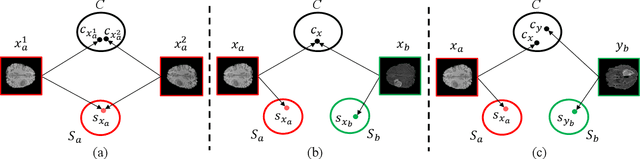
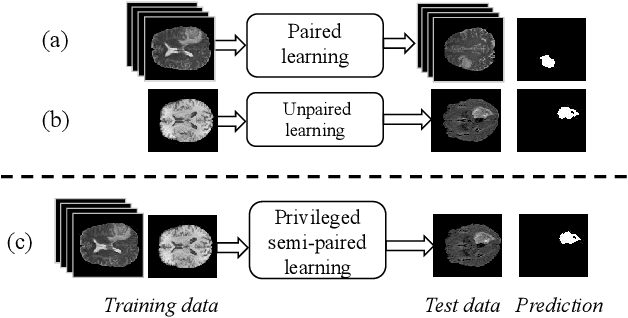
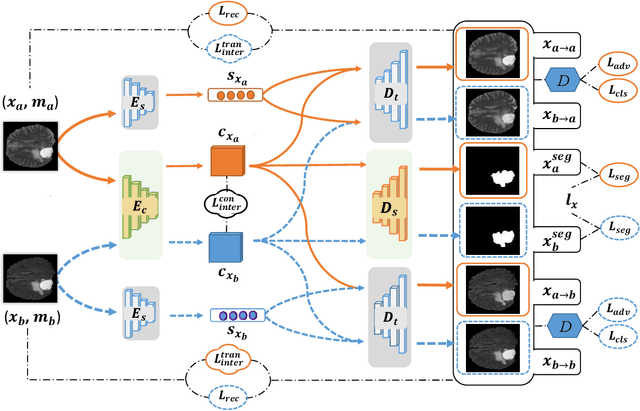
Due to the difficulties of obtaining multimodal paired images in clinical practice, recent studies propose to train brain tumor segmentation models with unpaired images and capture complementary information through modality translation. However, these models cannot fully exploit the complementary information from different modalities. In this work, we thus present a novel two-step (intra-modality and inter-modality) curriculum disentanglement learning framework to effectively utilize privileged semi-paired images, i.e. limited paired images that are only available in training, for brain tumor segmentation. Specifically, in the first step, we propose to conduct reconstruction and segmentation with augmented intra-modality style-consistent images. In the second step, the model jointly performs reconstruction, unsupervised/supervised translation, and segmentation for both unpaired and paired inter-modality images. A content consistency loss and a supervised translation loss are proposed to leverage complementary information from different modalities in this step. Through these two steps, our method effectively extracts modality-specific style codes describing the attenuation of tissue features and image contrast, and modality-invariant content codes containing anatomical and functional information from the input images. Experiments on three brain tumor segmentation tasks show that our model outperforms competing segmentation models based on unpaired images.
Deeply Supervised Layer Selective Attention Network: Towards Label-Efficient Learning for Medical Image Classification
Sep 28, 2022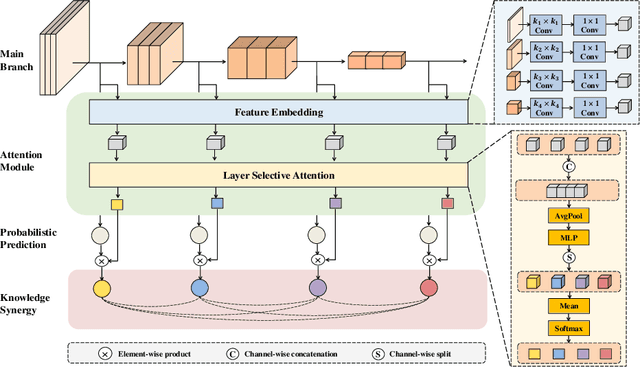

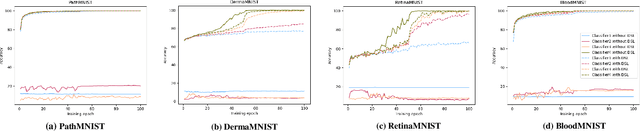
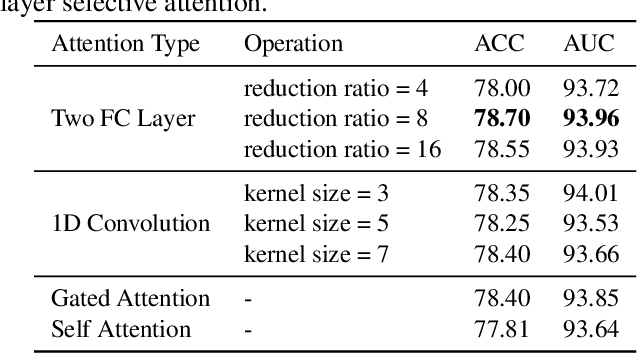
Labeling medical images depends on professional knowledge, making it difficult to acquire large amount of annotated medical images with high quality in a short time. Thus, making good use of limited labeled samples in a small dataset to build a high-performance model is the key to medical image classification problem. In this paper, we propose a deeply supervised Layer Selective Attention Network (LSANet), which comprehensively uses label information in feature-level and prediction-level supervision. For feature-level supervision, in order to better fuse the low-level features and high-level features, we propose a novel visual attention module, Layer Selective Attention (LSA), to focus on the feature selection of different layers. LSA introduces a weight allocation scheme which can dynamically adjust the weighting factor of each auxiliary branch during the whole training process to further enhance deeply supervised learning and ensure its generalization. For prediction-level supervision, we adopt the knowledge synergy strategy to promote hierarchical information interactions among all supervision branches via pairwise knowledge matching. Using the public dataset, MedMNIST, which is a large-scale benchmark for biomedical image classification covering diverse medical specialties, we evaluate LSANet on multiple mainstream CNN architectures and various visual attention modules. The experimental results show the substantial improvements of our proposed method over its corresponding counterparts, demonstrating that LSANet can provide a promising solution for label-efficient learning in the field of medical image classification.
FontTransformer: Few-shot High-resolution Chinese Glyph Image Synthesis via Stacked Transformers
Oct 13, 2022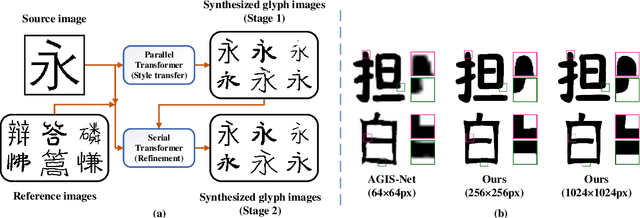
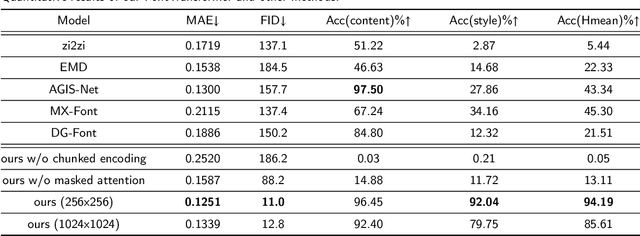
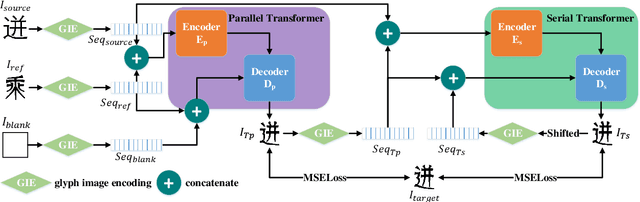
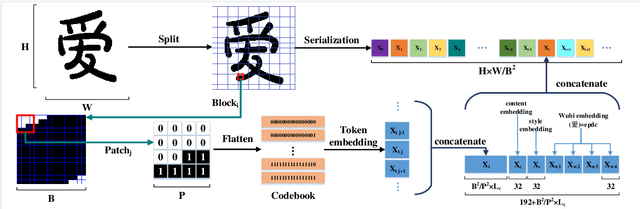
Automatic generation of high-quality Chinese fonts from a few online training samples is a challenging task, especially when the amount of samples is very small. Existing few-shot font generation methods can only synthesize low-resolution glyph images that often possess incorrect topological structures or/and incomplete strokes. To address the problem, this paper proposes FontTransformer, a novel few-shot learning model, for high-resolution Chinese glyph image synthesis by using stacked Transformers. The key idea is to apply the parallel Transformer to avoid the accumulation of prediction errors and utilize the serial Transformer to enhance the quality of synthesized strokes. Meanwhile, we also design a novel encoding scheme to feed more glyph information and prior knowledge to our model, which further enables the generation of high-resolution and visually-pleasing glyph images. Both qualitative and quantitative experimental results demonstrate the superiority of our method compared to other existing approaches in the few-shot Chinese font synthesis task.
Multi-Task Learning for Joint Semantic Role and Proto-Role Labeling
Oct 13, 2022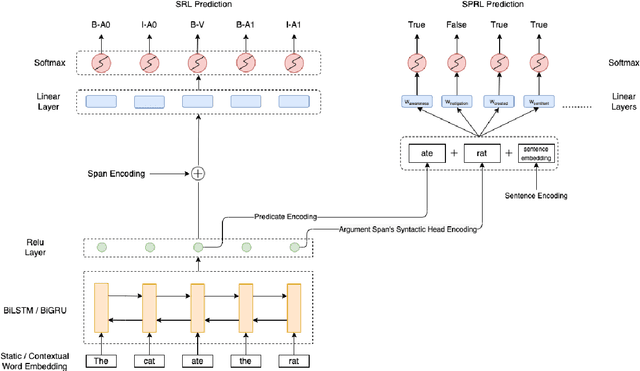
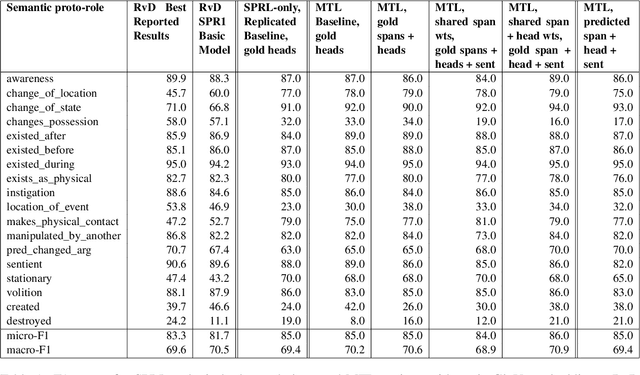
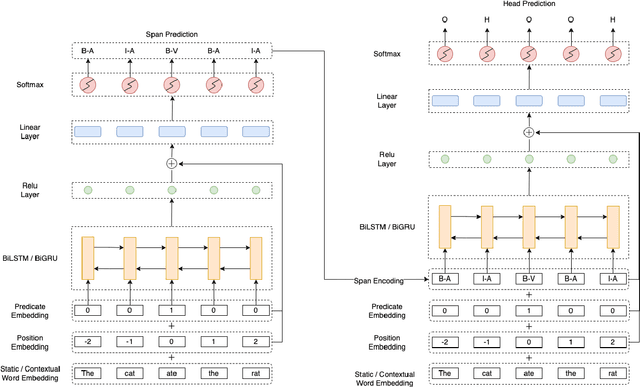
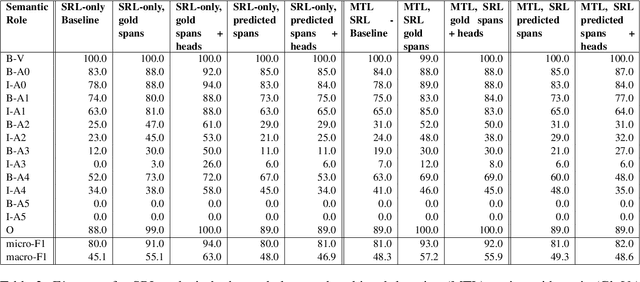
We put forward an end-to-end multi-step machine learning model which jointly labels semantic roles and the proto-roles of Dowty (1991), given a sentence and the predicates therein. Our best architecture first learns argument spans followed by learning the argument's syntactic heads. This information is shared with the next steps for predicting the semantic roles and proto-roles. We also experiment with transfer learning from argument and head prediction to role and proto-role labeling. We compare using static and contextual embeddings for words, arguments, and sentences. Unlike previous work, our model does not require pre-training or fine-tuning on additional tasks, beyond using off-the-shelf (static or contextual) embeddings and supervision. It also does not require argument spans, their semantic roles, and/or their gold syntactic heads as additional input, because it learns to predict all these during training. Our multi-task learning model raises the state-of-the-art predictions for most proto-roles.
Shape Preserving Facial Landmarks with Graph Attention Networks
Oct 13, 2022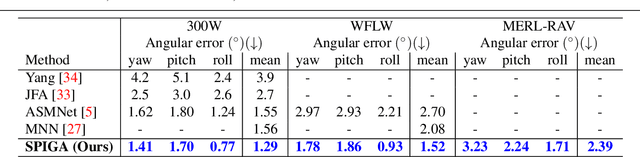
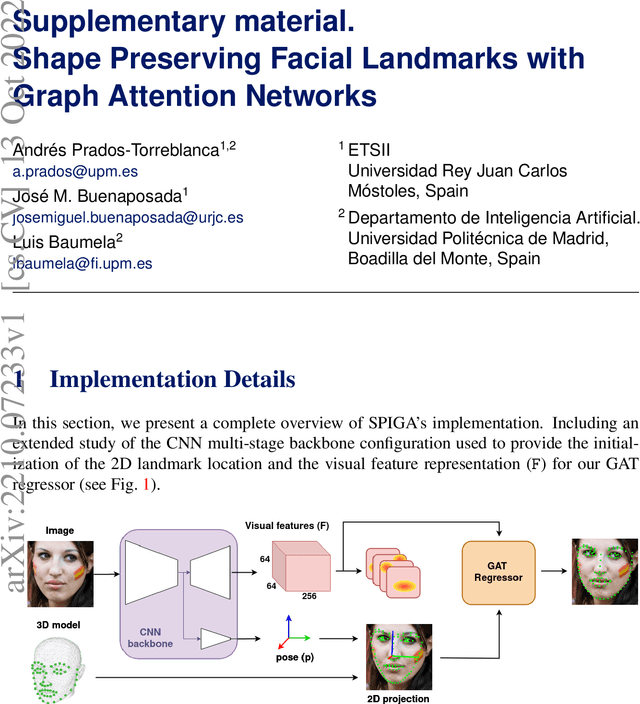

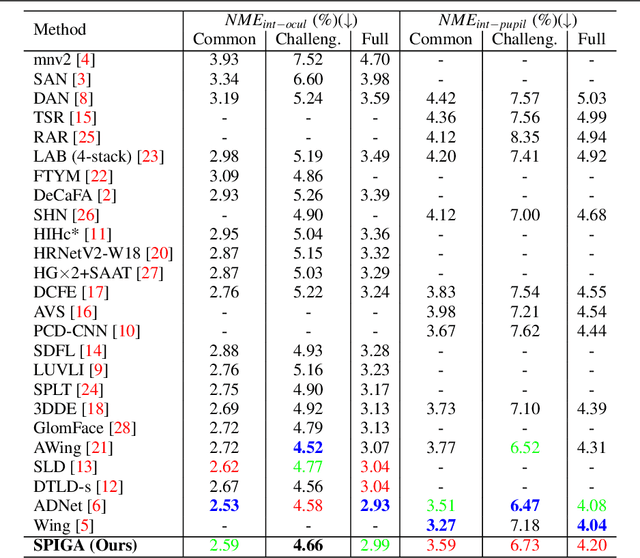
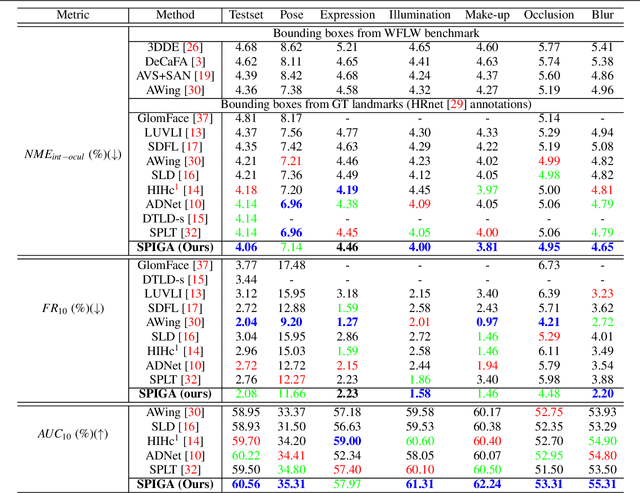
Top-performing landmark estimation algorithms are based on exploiting the excellent ability of large convolutional neural networks (CNNs) to represent local appearance. However, it is well known that they can only learn weak spatial relationships. To address this problem, we propose a model based on the combination of a CNN with a cascade of Graph Attention Network regressors. To this end, we introduce an encoding that jointly represents the appearance and location of facial landmarks and an attention mechanism to weigh the information according to its reliability. This is combined with a multi-task approach to initialize the location of graph nodes and a coarse-to-fine landmark description scheme. Our experiments confirm that the proposed model learns a global representation of the structure of the face, achieving top performance in popular benchmarks on head pose and landmark estimation. The improvement provided by our model is most significant in situations involving large changes in the local appearance of landmarks.
Outlier-Robust Group Inference via Gradient Space Clustering
Oct 13, 2022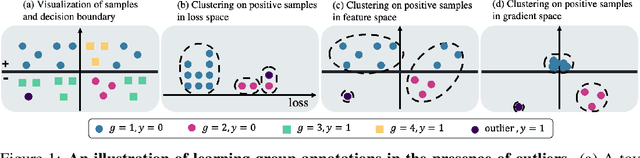
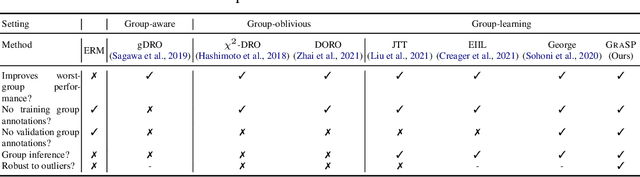
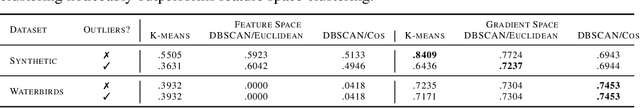
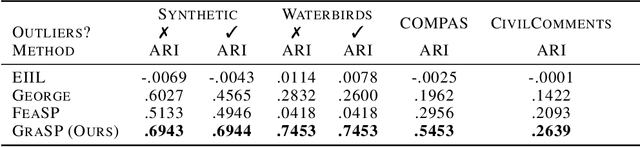
Traditional machine learning models focus on achieving good performance on the overall training distribution, but they often underperform on minority groups. Existing methods can improve the worst-group performance, but they can have several limitations: (i) they require group annotations, which are often expensive and sometimes infeasible to obtain, and/or (ii) they are sensitive to outliers. Most related works fail to solve these two issues simultaneously as they focus on conflicting perspectives of minority groups and outliers. We address the problem of learning group annotations in the presence of outliers by clustering the data in the space of gradients of the model parameters. We show that data in the gradient space has a simpler structure while preserving information about minority groups and outliers, making it suitable for standard clustering methods like DBSCAN. Extensive experiments demonstrate that our method significantly outperforms state-of-the-art both in terms of group identification and downstream worst-group performance.
RTFormer: Efficient Design for Real-Time Semantic Segmentation with Transformer
Oct 13, 2022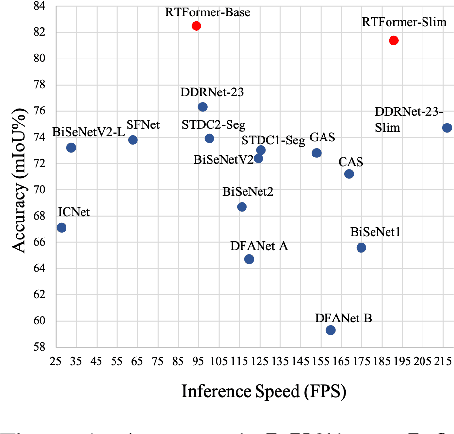
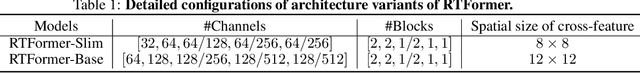
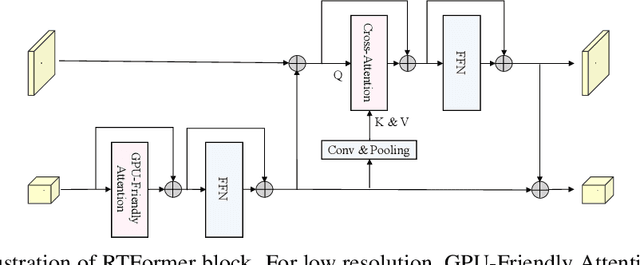
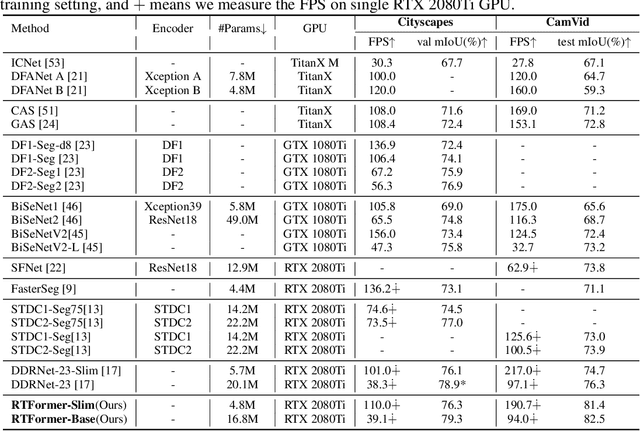
Recently, transformer-based networks have shown impressive results in semantic segmentation. Yet for real-time semantic segmentation, pure CNN-based approaches still dominate in this field, due to the time-consuming computation mechanism of transformer. We propose RTFormer, an efficient dual-resolution transformer for real-time semantic segmenation, which achieves better trade-off between performance and efficiency than CNN-based models. To achieve high inference efficiency on GPU-like devices, our RTFormer leverages GPU-Friendly Attention with linear complexity and discards the multi-head mechanism. Besides, we find that cross-resolution attention is more efficient to gather global context information for high-resolution branch by spreading the high level knowledge learned from low-resolution branch. Extensive experiments on mainstream benchmarks demonstrate the effectiveness of our proposed RTFormer, it achieves state-of-the-art on Cityscapes, CamVid and COCOStuff, and shows promising results on ADE20K. Code is available at PaddleSeg: https://github.com/PaddlePaddle/PaddleSeg.
Multi-Modality Cardiac Image Computing: A Survey
Aug 26, 2022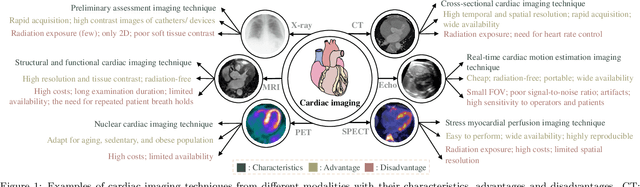
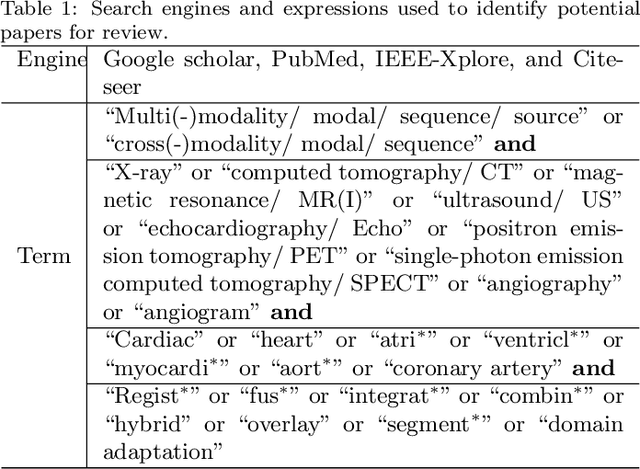
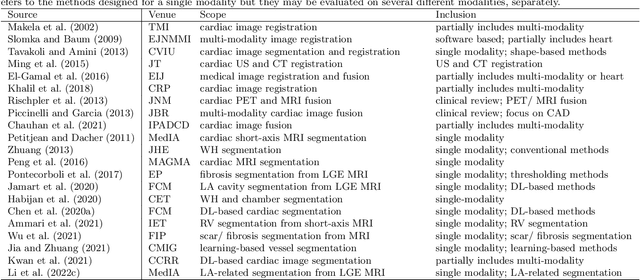

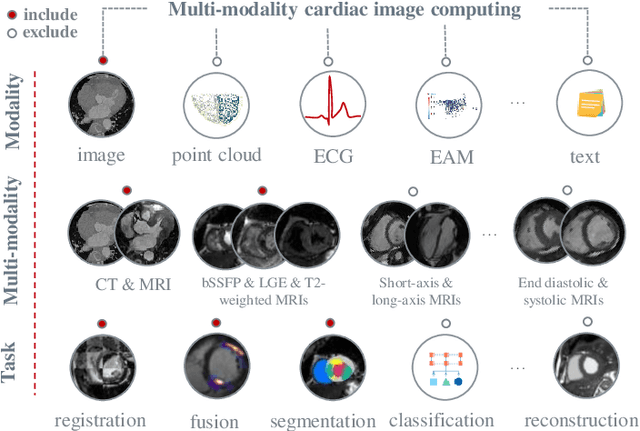

Multi-modality cardiac imaging plays a key role in the management of patients with cardiovascular diseases. It allows a combination of complementary anatomical, morphological and functional information, increases diagnosis accuracy, and improves the efficacy of cardiovascular interventions and clinical outcomes. Fully-automated processing and quantitative analysis of multi-modality cardiac images could have a direct impact on clinical research and evidence-based patient management. However, these require overcoming significant challenges including inter-modality misalignment and finding optimal methods to integrate information from different modalities. This paper aims to provide a comprehensive review of multi-modality imaging in cardiology, the computing methods, the validation strategies, the related clinical workflows and future perspectives. For the computing methodologies, we have a favored focus on the three tasks, i.e., registration, fusion and segmentation, which generally involve multi-modality imaging data, \textit{either combining information from different modalities or transferring information across modalities}. The review highlights that multi-modality cardiac imaging data has the potential of wide applicability in the clinic, such as trans-aortic valve implantation guidance, myocardial viability assessment, and catheter ablation therapy and its patient selection. Nevertheless, many challenges remain unsolved, such as missing modality, combination of imaging and non-imaging data, and uniform analysis and representation of different modalities. There is also work to do in defining how the well-developed techniques fit in clinical workflows and how much additional and relevant information they introduce. These problems are likely to continue to be an active field of research and the questions to be answered in the future.
Statistical shape representations for temporal registration of plant components in 3D
Sep 23, 2022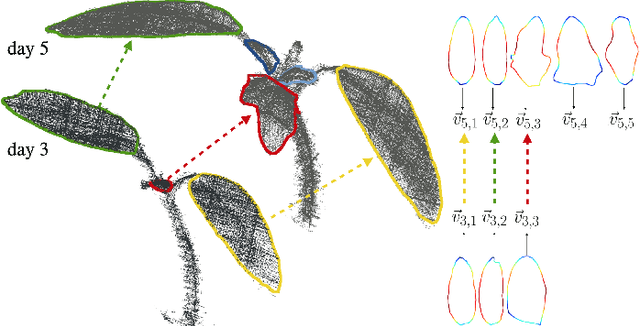
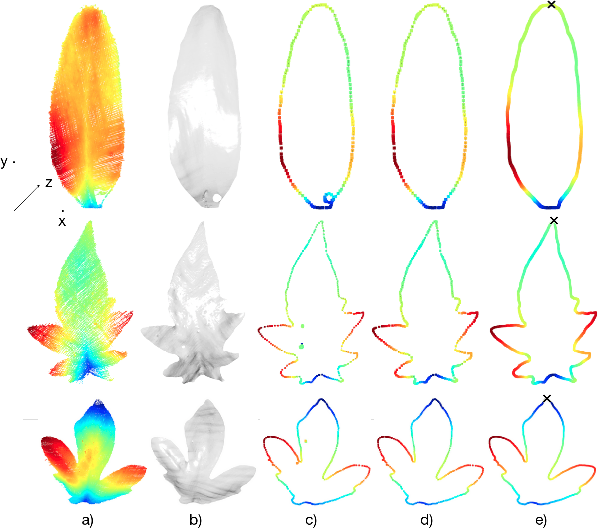
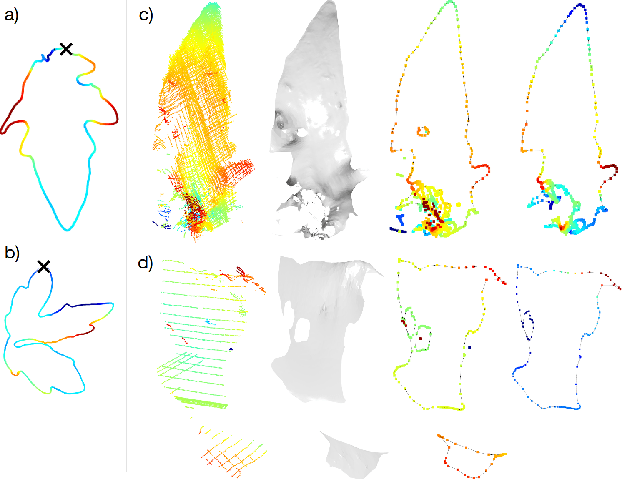
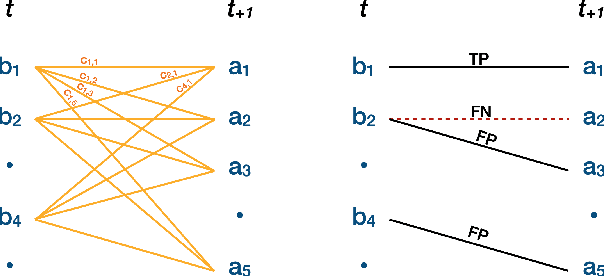
Plants are dynamic organisms. Understanding temporal variations in vegetation is an essential problem for all robots in the wild. However, associating repeated 3D scans of plants across time is challenging. A key step in this process is re-identifying and tracking the same individual plant components over time. Previously, this has been achieved by comparing their global spatial or topological location. In this work, we demonstrate how using shape features improves temporal organ matching. We present a landmark-free shape compression algorithm, which allows for the extraction of 3D shape features of leaves, characterises leaf shape and curvature efficiently in few parameters, and makes the association of individual leaves in feature space possible. The approach combines 3D contour extraction and further compression using Principal Component Analysis (PCA) to produce a shape space encoding, which is entirely learned from data and retains information about edge contours and 3D curvature. Our evaluation on temporal scan sequences of tomato plants shows, that incorporating shape features improves temporal leaf-matching. A combination of shape, location, and rotation information proves most informative for recognition of leaves over time and yields a true positive rate of 75%, a 15% improvement on sate-of-the-art methods. This is essential for robotic crop monitoring, which enables whole-of-lifecycle phenotyping.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge