"Information": models, code, and papers
A plug-in graph neural network to boost temporal sensitivity in fMRI analysis
Jan 01, 2023
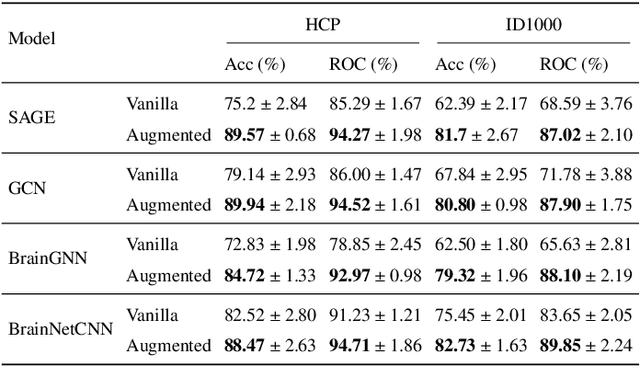
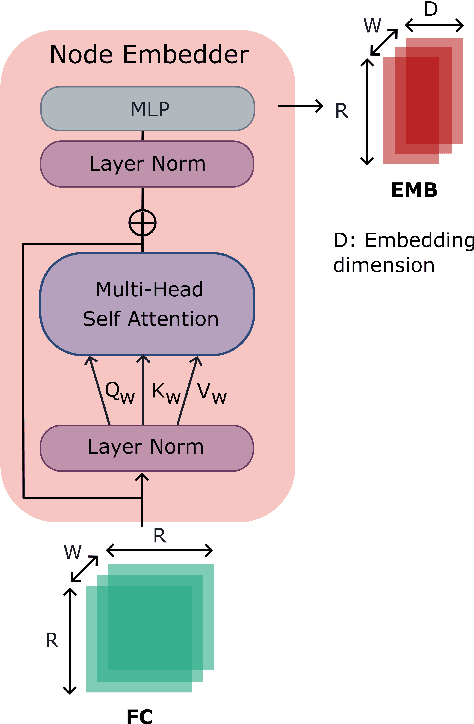
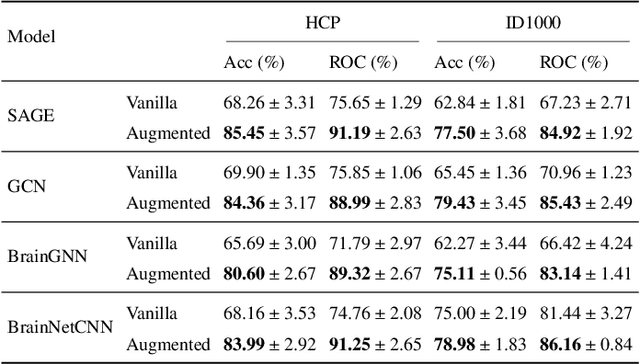
Learning-based methods have recently enabled performance leaps in analysis of high-dimensional functional MRI (fMRI) time series. Deep learning models that receive as input functional connectivity (FC) features among brain regions have been commonly adopted in the literature. However, many models focus on temporally static FC features across a scan, reducing sensitivity to dynamic features of brain activity. Here, we describe a plug-in graph neural network that can be flexibly integrated into a main learning-based fMRI model to boost its temporal sensitivity. Receiving brain regions as nodes and blood-oxygen-level-dependent (BOLD) signals as node inputs, the proposed GraphCorr method leverages a node embedder module based on a transformer encoder to capture temporally-windowed latent representations of BOLD signals. GraphCorr also leverages a lag filter module to account for delayed interactions across nodes by computing cross-correlation of windowed BOLD signals across a range of time lags. Information captured by the two modules is fused via a message passing algorithm executed on the graph, and enhanced node features are then computed at the output. These enhanced features are used to drive a subsequent learning-based model to analyze fMRI time series with elevated sensitivity. Comprehensive demonstrations on two public datasets indicate improved classification performance and interpretability for several state-of-the-art graphical and convolutional methods that employ GraphCorr-derived feature representations of fMRI time series as their input.
NeuroExplainer: Fine-Grained Attention Decoding to Uncover Cortical Development Patterns of Preterm Infants
Jan 01, 2023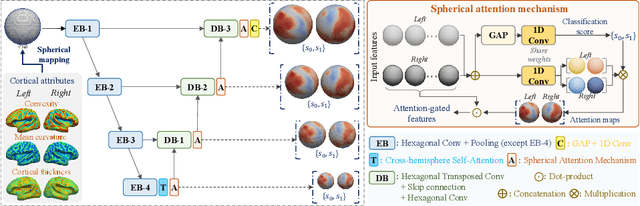
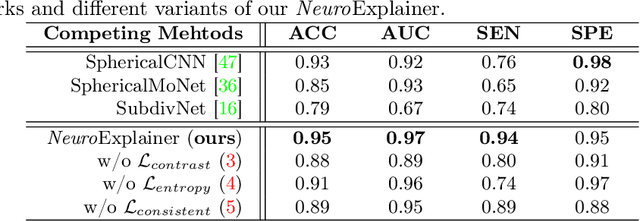
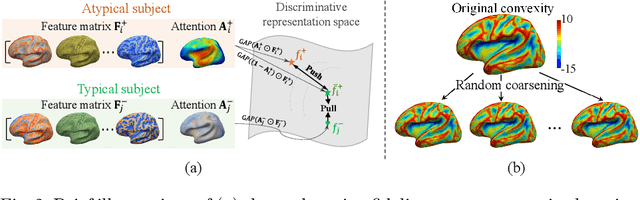
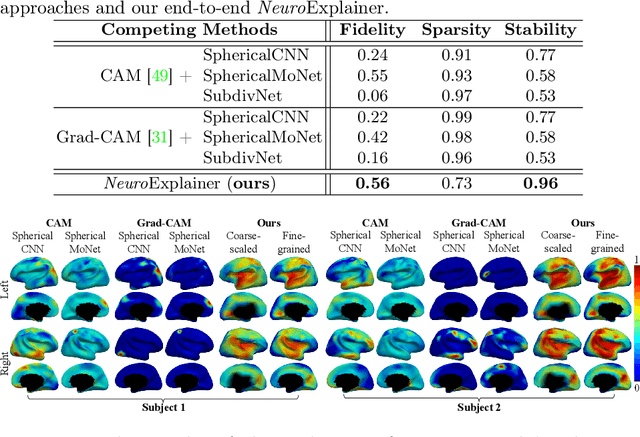
Deploying reliable deep learning techniques in interdisciplinary applications needs learned models to output accurate and ({even more importantly}) explainable predictions. Existing approaches typically explicate network outputs in a post-hoc fashion, under an implicit assumption that faithful explanations come from accurate predictions/classifications. We have an opposite claim that explanations boost (or even determine) classification. That is, end-to-end learning of explanation factors to augment discriminative representation extraction could be a more intuitive strategy to inversely assure fine-grained explainability, e.g., in those neuroimaging and neuroscience studies with high-dimensional data containing noisy, redundant, and task-irrelevant information. In this paper, we propose such an explainable geometric deep network dubbed as NeuroExplainer, with applications to uncover altered infant cortical development patterns associated with preterm birth. Given fundamental cortical attributes as network input, our NeuroExplainer adopts a hierarchical attention-decoding framework to learn fine-grained attentions and respective discriminative representations to accurately recognize preterm infants from term-born infants at term-equivalent age. NeuroExplainer learns the hierarchical attention-decoding modules under subject-level weak supervision coupled with targeted regularizers deduced from domain knowledge regarding brain development. These prior-guided constraints implicitly maximizes the explainability metrics (i.e., fidelity, sparsity, and stability) in network training, driving the learned network to output detailed explanations and accurate classifications. Experimental results on the public dHCP benchmark suggest that NeuroExplainer led to quantitatively reliable explanation results that are qualitatively consistent with representative neuroimaging studies.
MTNeuro: A Benchmark for Evaluating Representations of Brain Structure Across Multiple Levels of Abstraction
Jan 01, 2023
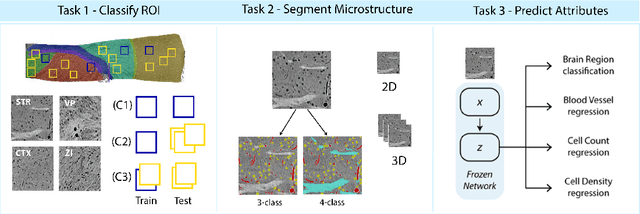
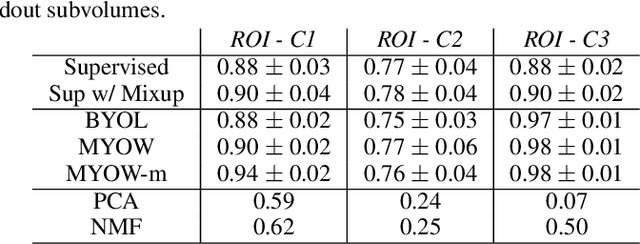

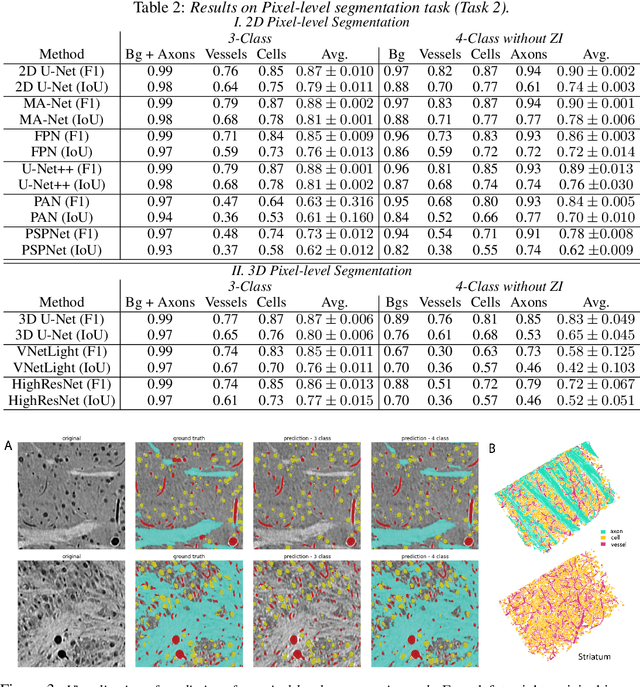
There are multiple scales of abstraction from which we can describe the same image, depending on whether we are focusing on fine-grained details or a more global attribute of the image. In brain mapping, learning to automatically parse images to build representations of both small-scale features (e.g., the presence of cells or blood vessels) and global properties of an image (e.g., which brain region the image comes from) is a crucial and open challenge. However, most existing datasets and benchmarks for neuroanatomy consider only a single downstream task at a time. To bridge this gap, we introduce a new dataset, annotations, and multiple downstream tasks that provide diverse ways to readout information about brain structure and architecture from the same image. Our multi-task neuroimaging benchmark (MTNeuro) is built on volumetric, micrometer-resolution X-ray microtomography images spanning a large thalamocortical section of mouse brain, encompassing multiple cortical and subcortical regions. We generated a number of different prediction challenges and evaluated several supervised and self-supervised models for brain-region prediction and pixel-level semantic segmentation of microstructures. Our experiments not only highlight the rich heterogeneity of this dataset, but also provide insights into how self-supervised approaches can be used to learn representations that capture multiple attributes of a single image and perform well on a variety of downstream tasks. Datasets, code, and pre-trained baseline models are provided at: https://mtneuro.github.io/ .
Removing Objects From Neural Radiance Fields
Dec 22, 2022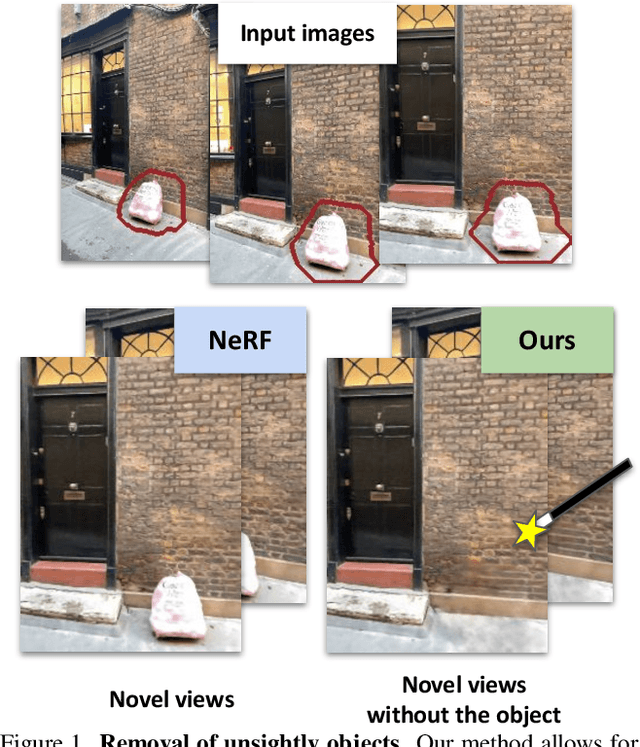
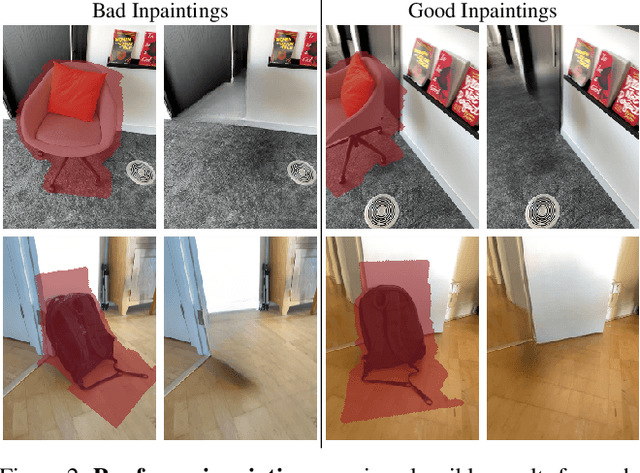
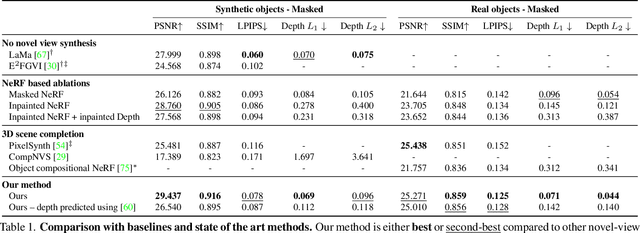

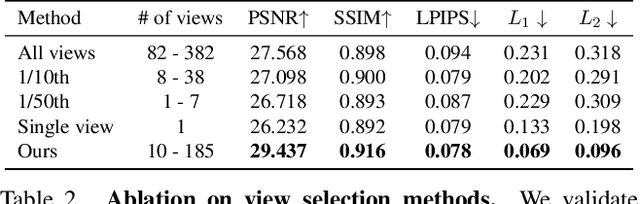
Neural Radiance Fields (NeRFs) are emerging as a ubiquitous scene representation that allows for novel view synthesis. Increasingly, NeRFs will be shareable with other people. Before sharing a NeRF, though, it might be desirable to remove personal information or unsightly objects. Such removal is not easily achieved with the current NeRF editing frameworks. We propose a framework to remove objects from a NeRF representation created from an RGB-D sequence. Our NeRF inpainting method leverages recent work in 2D image inpainting and is guided by a user-provided mask. Our algorithm is underpinned by a confidence based view selection procedure. It chooses which of the individual 2D inpainted images to use in the creation of the NeRF, so that the resulting inpainted NeRF is 3D consistent. We show that our method for NeRF editing is effective for synthesizing plausible inpaintings in a multi-view coherent manner. We validate our approach using a new and still-challenging dataset for the task of NeRF inpainting.
Over-the-Air Federated Learning with Enhanced Privacy
Dec 22, 2022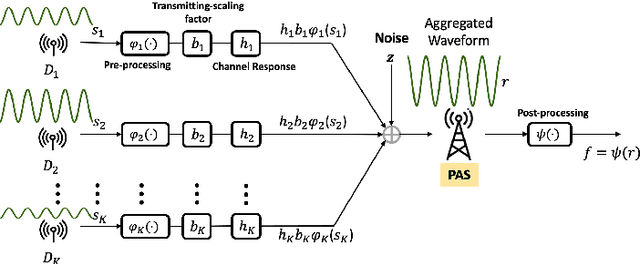
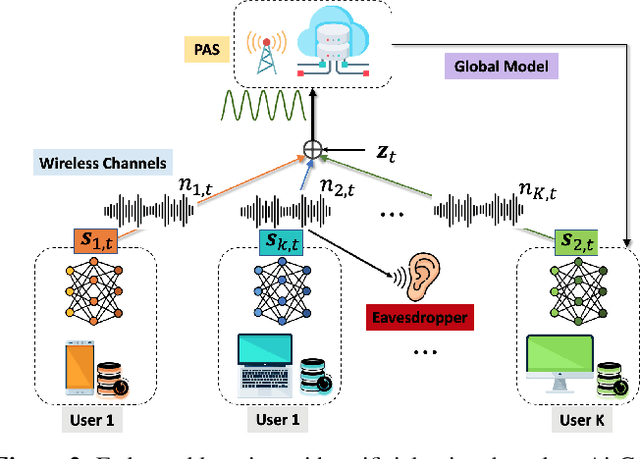
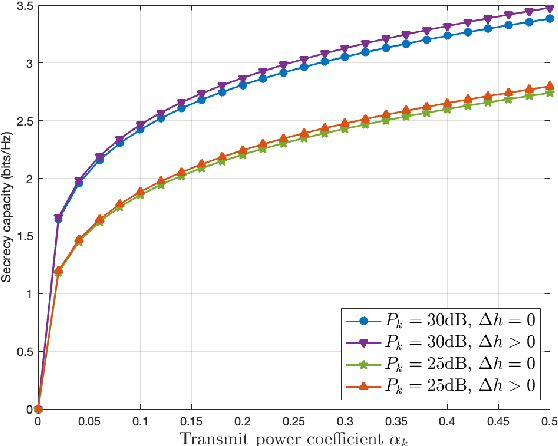
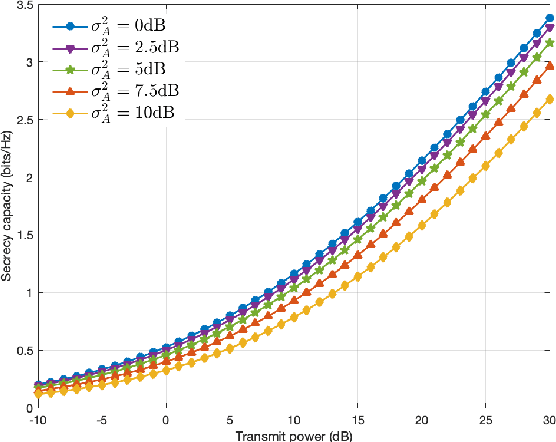
Federated learning (FL) has emerged as a promising learning paradigm in which only local model parameters (gradients) are shared. Private user data never leaves the local devices thus preserving data privacy. However, recent research has shown that even when local data is never shared by a user, exchanging model parameters without protection can also leak private information. Moreover, in wireless systems, the frequent transmission of model parameters can cause tremendous bandwidth consumption and network congestion when the model is large. To address this problem, we propose a new FL framework with efficient over-the-air parameter aggregation and strong privacy protection of both user data and models. We achieve this by introducing pairwise cancellable random artificial noises (PCR-ANs) on end devices. As compared to existing over-the-air computation (AirComp) based FL schemes, our design provides stronger privacy protection. We analytically show the secrecy capacity and the convergence rate of the proposed wireless FL aggregation algorithm.
Shakes on a Plane: Unsupervised Depth Estimation from Unstabilized Photography
Dec 22, 2022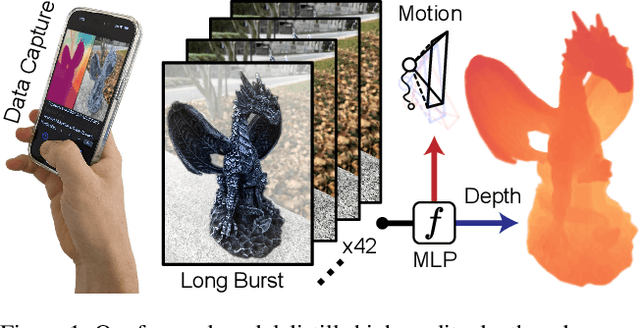
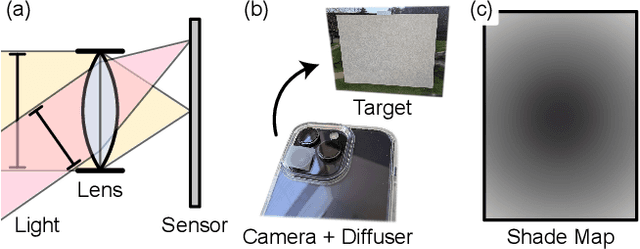
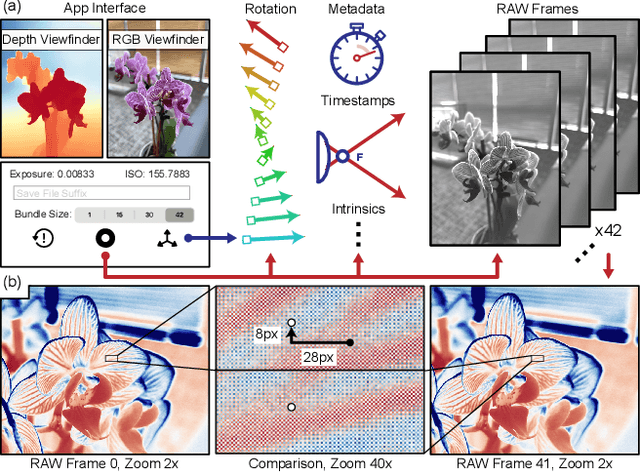
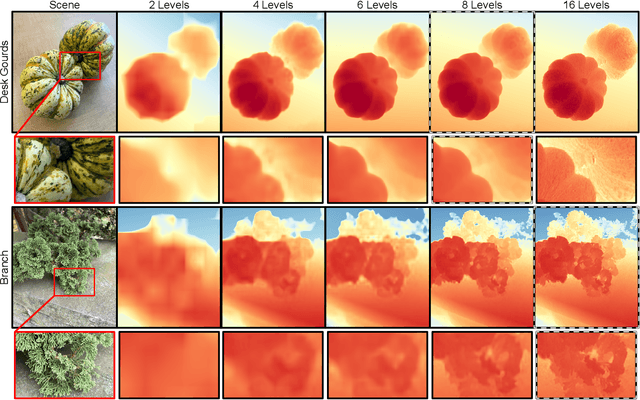
Modern mobile burst photography pipelines capture and merge a short sequence of frames to recover an enhanced image, but often disregard the 3D nature of the scene they capture, treating pixel motion between images as a 2D aggregation problem. We show that in a "long-burst", forty-two 12-megapixel RAW frames captured in a two-second sequence, there is enough parallax information from natural hand tremor alone to recover high-quality scene depth. To this end, we devise a test-time optimization approach that fits a neural RGB-D representation to long-burst data and simultaneously estimates scene depth and camera motion. Our plane plus depth model is trained end-to-end, and performs coarse-to-fine refinement by controlling which multi-resolution volume features the network has access to at what time during training. We validate the method experimentally, and demonstrate geometrically accurate depth reconstructions with no additional hardware or separate data pre-processing and pose-estimation steps.
A deep local attention network for pre-operative lymph node metastasis prediction in pancreatic cancer via multiphase CT imaging
Jan 04, 2023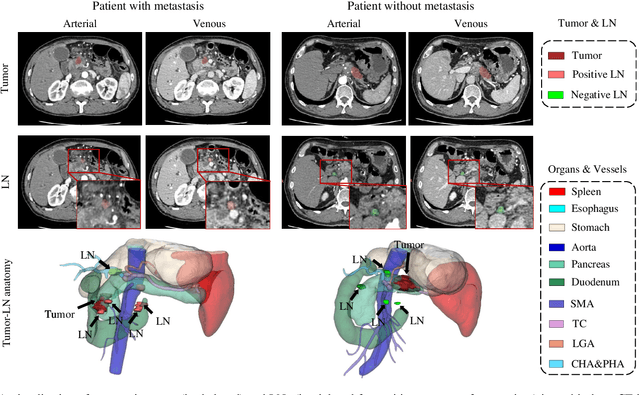
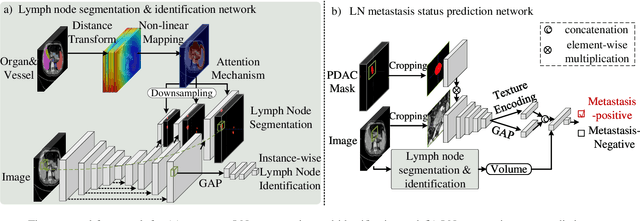
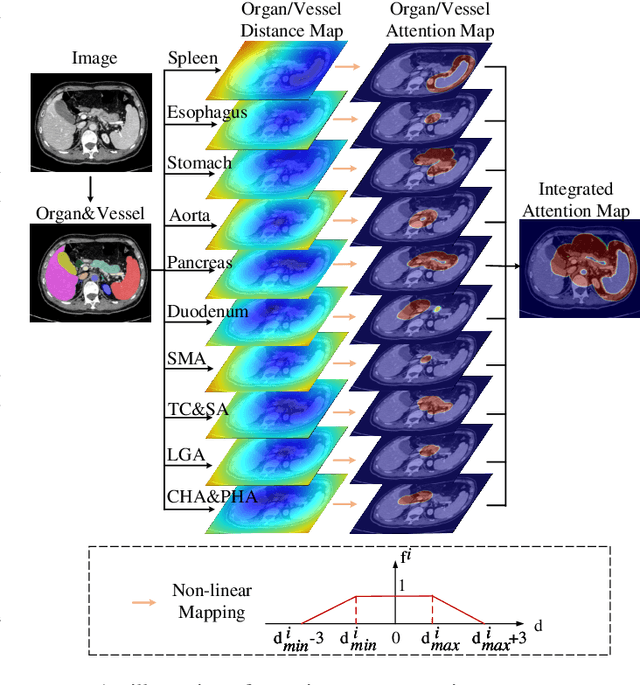
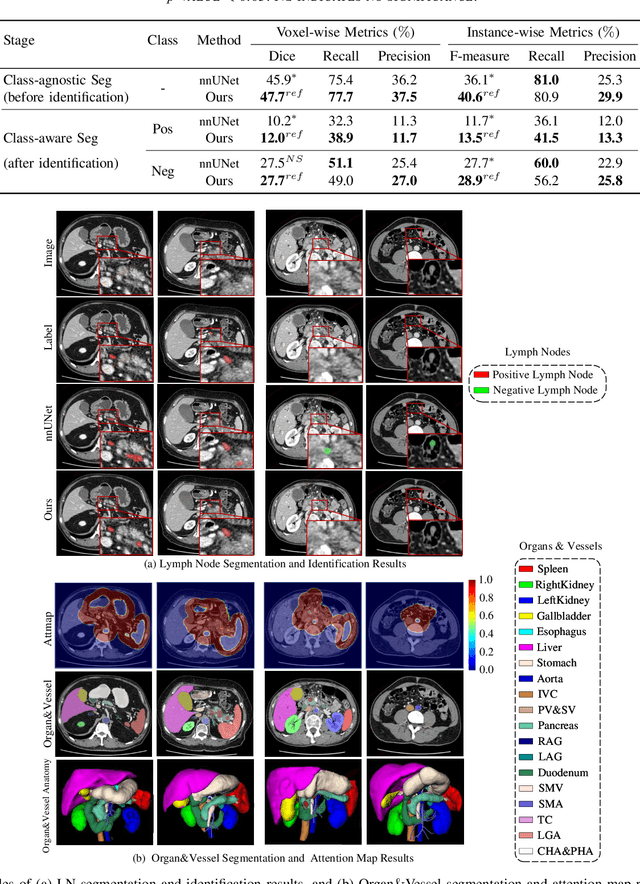
Lymph node (LN) metastasis status is one of the most critical prognostic and cancer staging factors for patients with resectable pancreatic ductal adenocarcinoma (PDAC), or in general, for any types of solid malignant tumors. Preoperative prediction of LN metastasis from non-invasive CT imaging is highly desired, as it might be straightforwardly used to guide the following neoadjuvant treatment decision and surgical planning. Most studies only capture the tumor characteristics in CT imaging to implicitly infer LN metastasis and very few work exploit direct LN's CT imaging information. To the best of our knowledge, this is the first work to propose a fully-automated LN segmentation and identification network to directly facilitate the LN metastasis status prediction task. Nevertheless LN segmentation/detection is very challenging since LN can be easily confused with other hard negative anatomic structures (e.g., vessels) from radiological images. We explore the anatomical spatial context priors of pancreatic LN locations by generating a guiding attention map from related organs and vessels to assist segmentation and infer LN status. As such, LN segmentation is impelled to focus on regions that are anatomically adjacent or plausible with respect to the specific organs and vessels. The metastasized LN identification network is trained to classify the segmented LN instances into positives or negatives by reusing the segmentation network as a pre-trained backbone and padding a new classification head. More importantly, we develop a LN metastasis status prediction network that combines the patient-wise aggregation results of LN segmentation/identification and deep imaging features extracted from the tumor region. Extensive quantitative nested five-fold cross-validation is conducted on a discovery dataset of 749 patients with PDAC.
Forecasting User Interests Through Topic Tag Predictions in Online Health Communities
Nov 05, 2022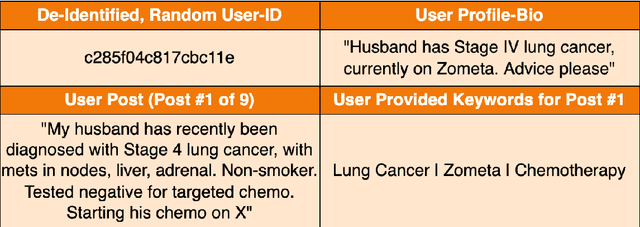
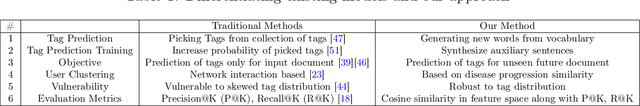
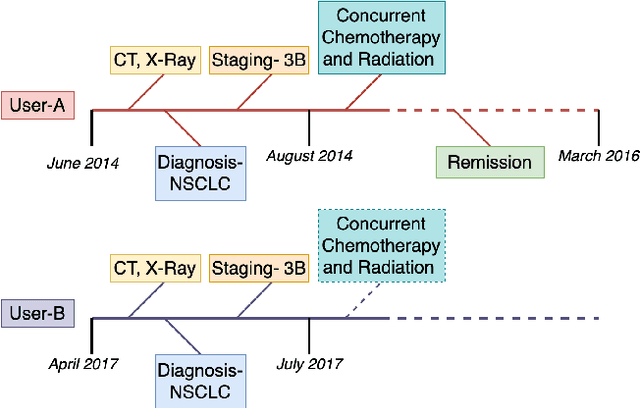
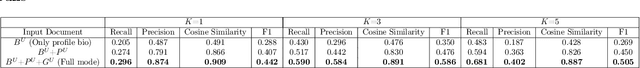
The increasing reliance on online communities for healthcare information by patients and caregivers has led to the increase in the spread of misinformation, or subjective, anecdotal and inaccurate or non-specific recommendations, which, if acted on, could cause serious harm to the patients. Hence, there is an urgent need to connect users with accurate and tailored health information in a timely manner to prevent such harm. This paper proposes an innovative approach to suggesting reliable information to participants in online communities as they move through different stages in their disease or treatment. We hypothesize that patients with similar histories of disease progression or course of treatment would have similar information needs at comparable stages. Specifically, we pose the problem of predicting topic tags or keywords that describe the future information needs of users based on their profiles, traces of their online interactions within the community (past posts, replies) and the profiles and traces of online interactions of other users with similar profiles and similar traces of past interaction with the target users. The result is a variant of the collaborative information filtering or recommendation system tailored to the needs of users of online health communities. We report results of our experiments on an expert curated data set which demonstrate the superiority of the proposed approach over the state of the art baselines with respect to accurate and timely prediction of topic tags (and hence information sources of interest).
SARAS-Net: Scale and Relation Aware Siamese Network for Change Detection
Dec 02, 2022
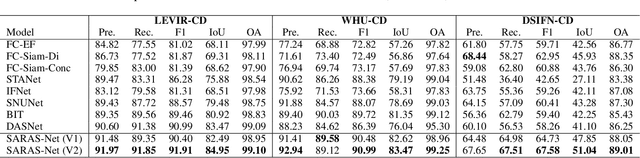
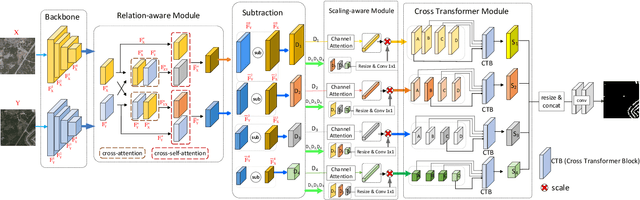
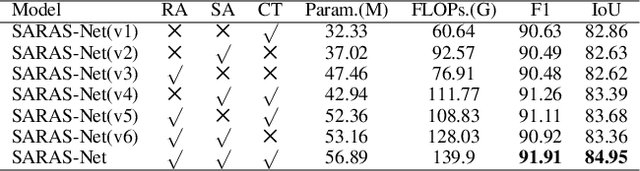
Change detection (CD) aims to find the difference between two images at different times and outputs a change map to represent whether the region has changed or not. To achieve a better result in generating the change map, many State-of-The-Art (SoTA) methods design a deep learning model that has a powerful discriminative ability. However, these methods still get lower performance because they ignore spatial information and scaling changes between objects, giving rise to blurry or wrong boundaries. In addition to these, they also neglect the interactive information of two different images. To alleviate these problems, we propose our network, the Scale and Relation-Aware Siamese Network (SARAS-Net) to deal with this issue. In this paper, three modules are proposed that include relation-aware, scale-aware, and cross-transformer to tackle the problem of scene change detection more effectively. To verify our model, we tested three public datasets, including LEVIR-CD, WHU-CD, and DSFIN, and obtained SoTA accuracy. Our code is available at https://github.com/f64051041/SARAS-Net.
Role of Audio in Audio-Visual Video Summarization
Dec 02, 2022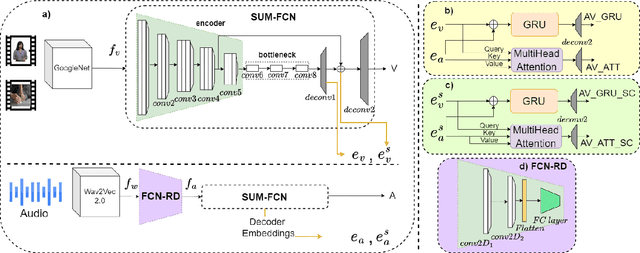
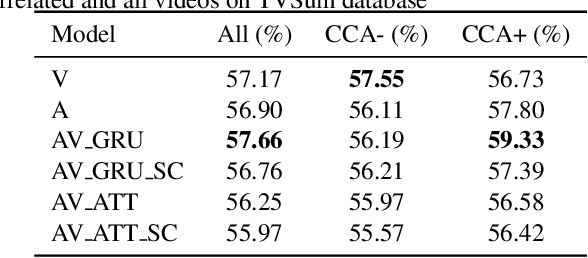
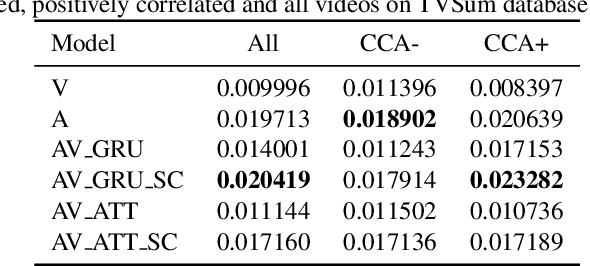
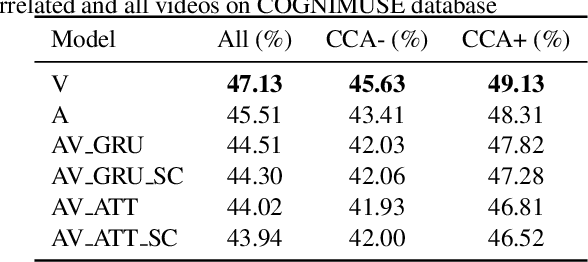
Video summarization attracts attention for efficient video representation, retrieval, and browsing to ease volume and traffic surge problems. Although video summarization mostly uses the visual channel for compaction, the benefits of audio-visual modeling appeared in recent literature. The information coming from the audio channel can be a result of audio-visual correlation in the video content. In this study, we propose a new audio-visual video summarization framework integrating four ways of audio-visual information fusion with GRU-based and attention-based networks. Furthermore, we investigate a new explainability methodology using audio-visual canonical correlation analysis (CCA) to better understand and explain the role of audio in the video summarization task. Experimental evaluations on the TVSum dataset attain F1 score and Kendall-tau score improvements for the audio-visual video summarization. Furthermore, splitting video content on TVSum and COGNIMUSE datasets based on audio-visual CCA as positively and negatively correlated videos yields a strong performance improvement over the positively correlated videos for audio-only and audio-visual video summarization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge