"Information": models, code, and papers
A Survey on Conversational Search and Applications in Biomedicine
Nov 28, 2022
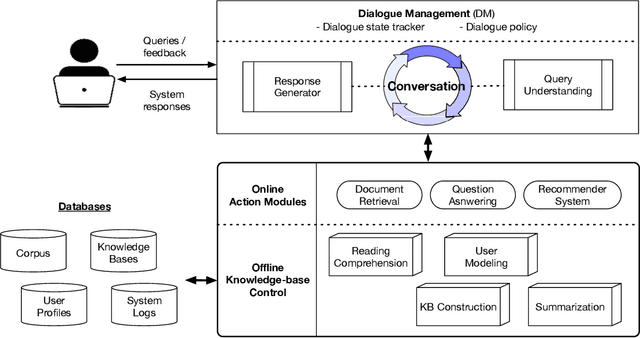
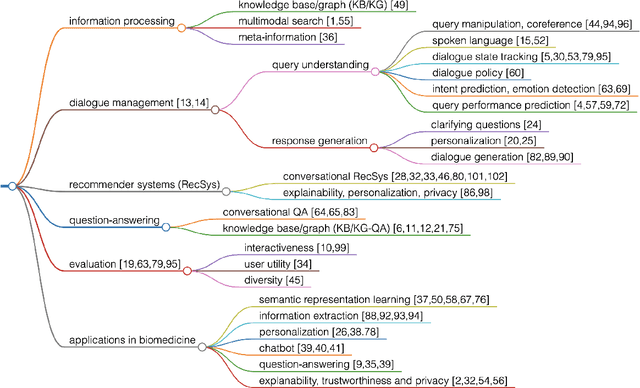
This paper aims to provide a radical rundown on Conversation Search (ConvSearch), an approach to enhance the information retrieval method where users engage in a dialogue for the information-seeking tasks. In this survey, we predominantly focused on the human interactive characteristics of the ConvSearch systems, highlighting the operations of the action modules, likely the Retrieval system, Question-Answering, and Recommender system. We labeled various ConvSearch research problems in knowledge bases, natural language processing, and dialogue management systems along with the action modules. We further categorized the framework to ConvSearch and the application is directed toward biomedical and healthcare fields for the utilization of clinical social technology. Finally, we conclude by talking through the challenges and issues of ConvSearch, particularly in Bio-Medicine. Our main aim is to provide an integrated and unified vision of the ConvSearch components from different fields, which benefit the information-seeking process in healthcare systems.
An Adaptive Kernel Approach to Federated Learning of Heterogeneous Causal Effects
Jan 01, 2023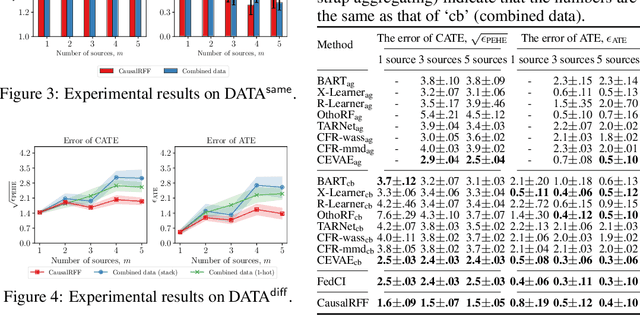
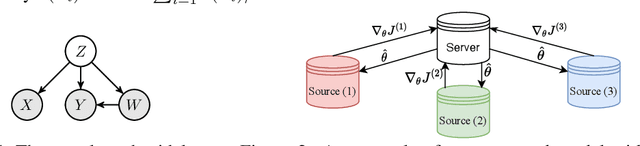
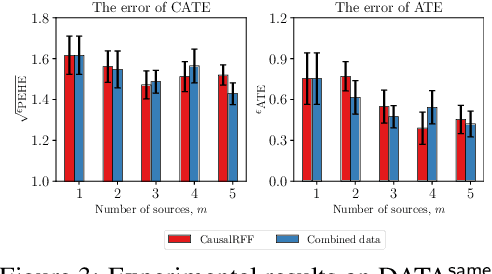
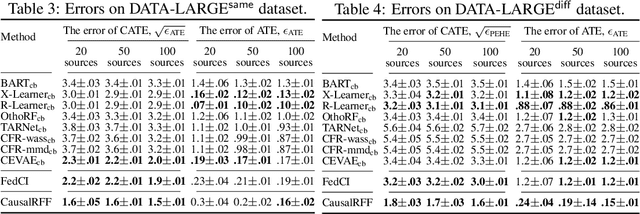
We propose a new causal inference framework to learn causal effects from multiple, decentralized data sources in a federated setting. We introduce an adaptive transfer algorithm that learns the similarities among the data sources by utilizing Random Fourier Features to disentangle the loss function into multiple components, each of which is associated with a data source. The data sources may have different distributions; the causal effects are independently and systematically incorporated. The proposed method estimates the similarities among the sources through transfer coefficients, and hence requiring no prior information about the similarity measures. The heterogeneous causal effects can be estimated with no sharing of the raw training data among the sources, thus minimizing the risk of privacy leak. We also provide minimax lower bounds to assess the quality of the parameters learned from the disparate sources. The proposed method is empirically shown to outperform the baselines on decentralized data sources with dissimilar distributions.
L-SeqSleepNet: Whole-cycle Long Sequence Modelling for Automatic Sleep Staging
Jan 09, 2023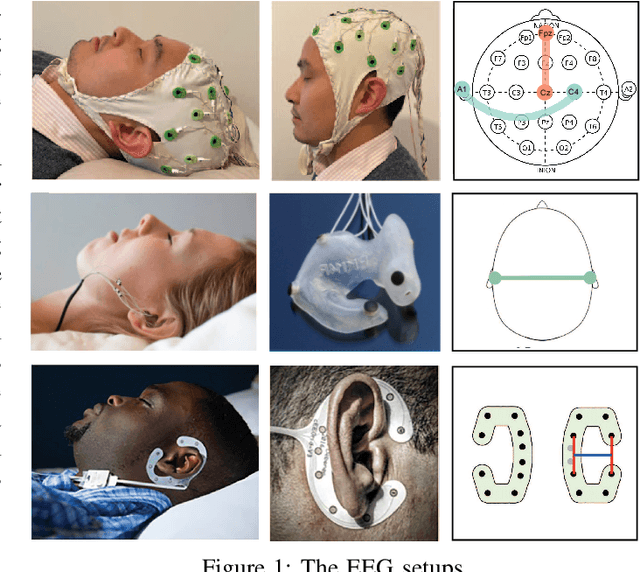
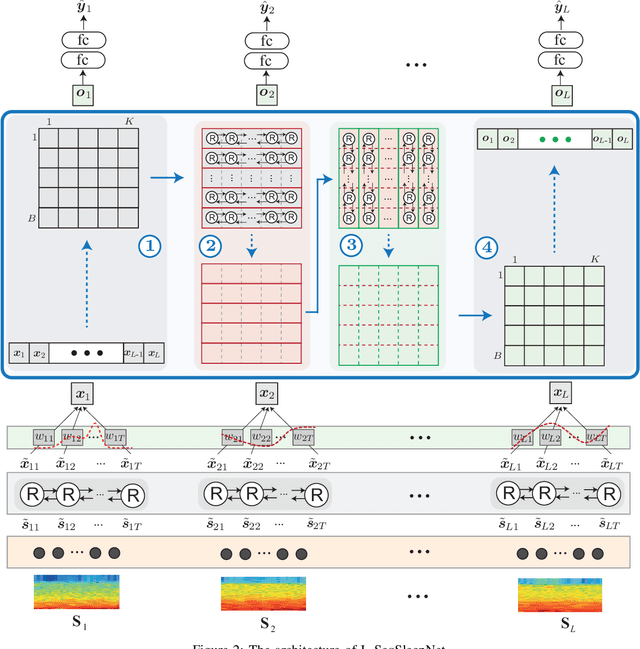
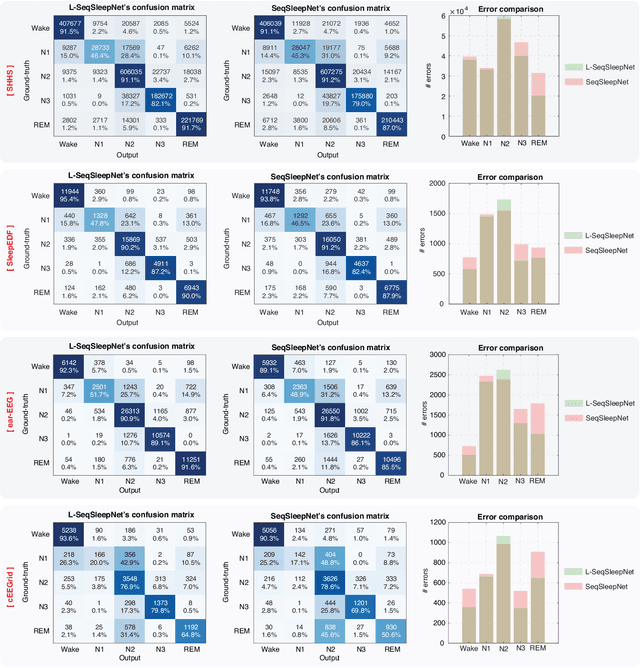
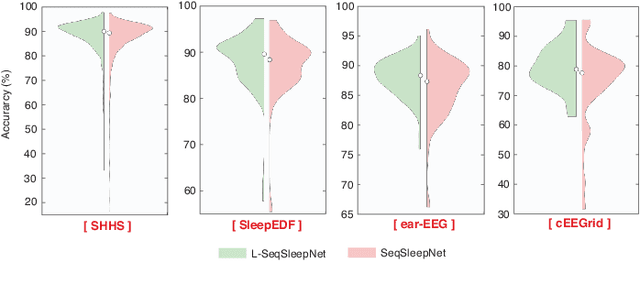
Human sleep is cyclical with a period of approximately 90 minutes, implying long temporal dependency in the sleep data. Yet, exploring this long-term dependency when developing sleep staging models has remained untouched. In this work, we show that while encoding the logic of a whole sleep cycle is crucial to improve sleep staging performance, the sequential modelling approach in existing state-of-the-art deep learning models are inefficient for that purpose. We then introduce a method for efficient long sequence modelling and propose a new deep learning model, L-SeqSleepNet, incorporating this method to take into account whole-cycle sleep information for sleep staging. Evaluating L-SeqSleepNet on a set of four distinct databases of various sizes, we demonstrate state-of-the-art performance obtained by the model over three different EEG setups, including scalp EEG in conventional Polysomnography (PSG), in-ear EEG, and around-the-ear EEG (cEEGrid), even with a single-EEG channel input. Our analyses also show that L-SeqSleepNet is able to remedy the effect of N2 sleep (the major class in terms of classification) to bring down errors in other sleep stages and that the network largely reduces exceptionally high errors seen in many subjects. Finally, the computation time only grows at a sub-linear rate when the sequence length increases.
MaNi: Maximizing Mutual Information for Nuclei Cross-Domain Unsupervised Segmentation
Jun 29, 2022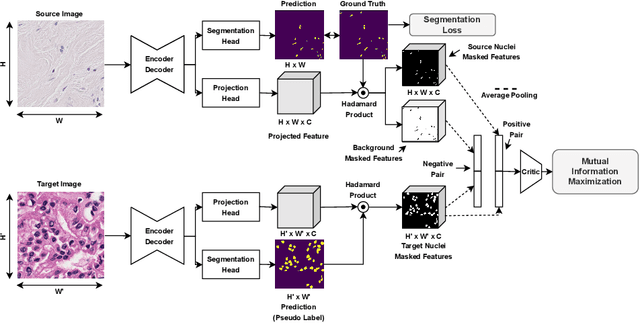
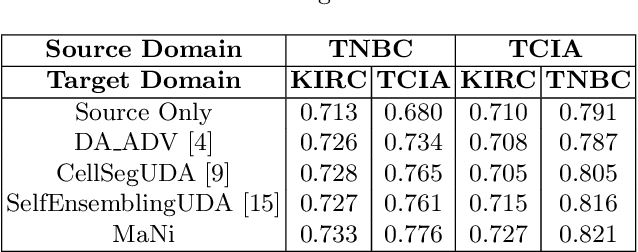
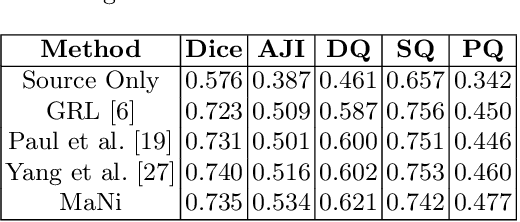
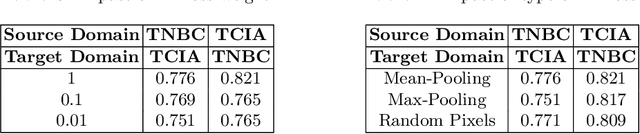
In this work, we propose a mutual information (MI) based unsupervised domain adaptation (UDA) method for the cross-domain nuclei segmentation. Nuclei vary substantially in structure and appearances across different cancer types, leading to a drop in performance of deep learning models when trained on one cancer type and tested on another. This domain shift becomes even more critical as accurate segmentation and quantification of nuclei is an essential histopathology task for the diagnosis/ prognosis of patients and annotating nuclei at the pixel level for new cancer types demands extensive effort by medical experts. To address this problem, we maximize the MI between labeled source cancer type data and unlabeled target cancer type data for transferring nuclei segmentation knowledge across domains. We use the Jensen-Shanon divergence bound, requiring only one negative pair per positive pair for MI maximization. We evaluate our set-up for multiple modeling frameworks and on different datasets comprising of over 20 cancer-type domain shifts and demonstrate competitive performance. All the recently proposed approaches consist of multiple components for improving the domain adaptation, whereas our proposed module is light and can be easily incorporated into other methods (Implementation: https://github.com/YashSharma/MaNi ).
DetIE: Multilingual Open Information Extraction Inspired by Object Detection
Jun 24, 2022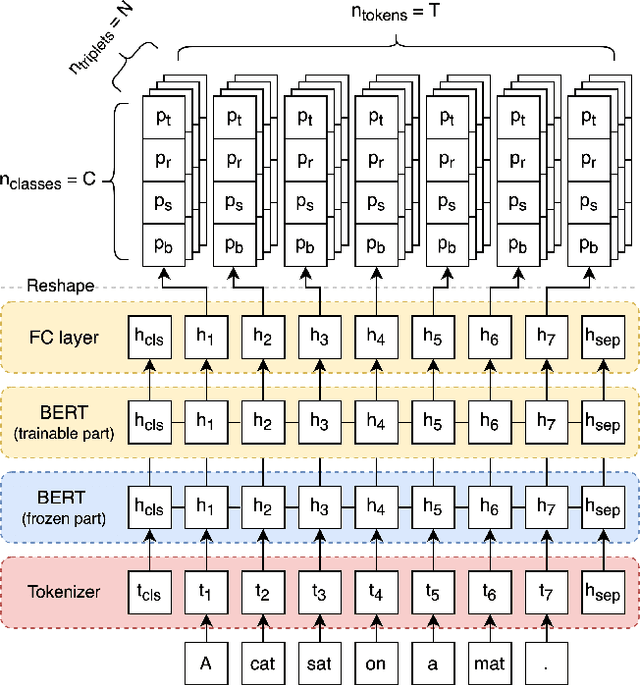
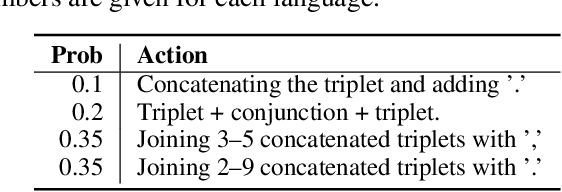
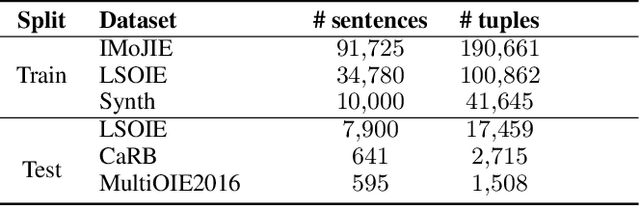
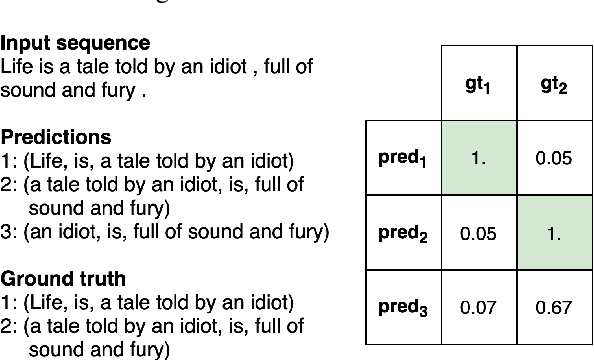

State of the art neural methods for open information extraction (OpenIE) usually extract triplets (or tuples) iteratively in an autoregressive or predicate-based manner in order not to produce duplicates. In this work, we propose a different approach to the problem that can be equally or more successful. Namely, we present a novel single-pass method for OpenIE inspired by object detection algorithms from computer vision. We use an order-agnostic loss based on bipartite matching that forces unique predictions and a Transformer-based encoder-only architecture for sequence labeling. The proposed approach is faster and shows superior or similar performance in comparison with state of the art models on standard benchmarks in terms of both quality metrics and inference time. Our model sets the new state of the art performance of 67.7% F1 on CaRB evaluated as OIE2016 while being 3.35x faster at inference than previous state of the art. We also evaluate the multilingual version of our model in the zero-shot setting for two languages and introduce a strategy for generating synthetic multilingual data to fine-tune the model for each specific language. In this setting, we show performance improvement 15% on multilingual Re-OIE2016, reaching 75% F1 for both Portuguese and Spanish languages. Code and models are available at https://github.com/sberbank-ai/DetIE.
GreenDB -- A Dataset and Benchmark for Extraction of Sustainability Information of Consumer Goods
Jul 21, 2022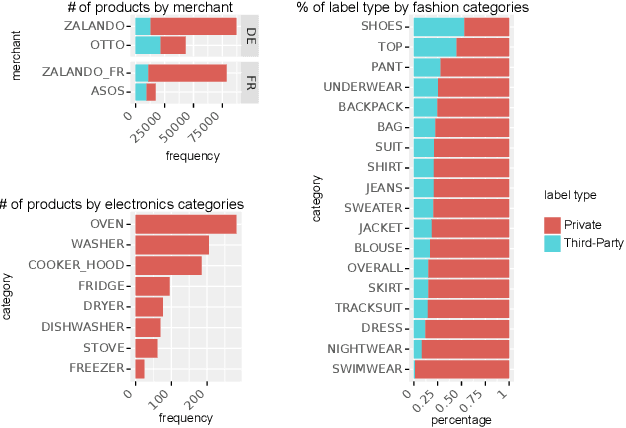
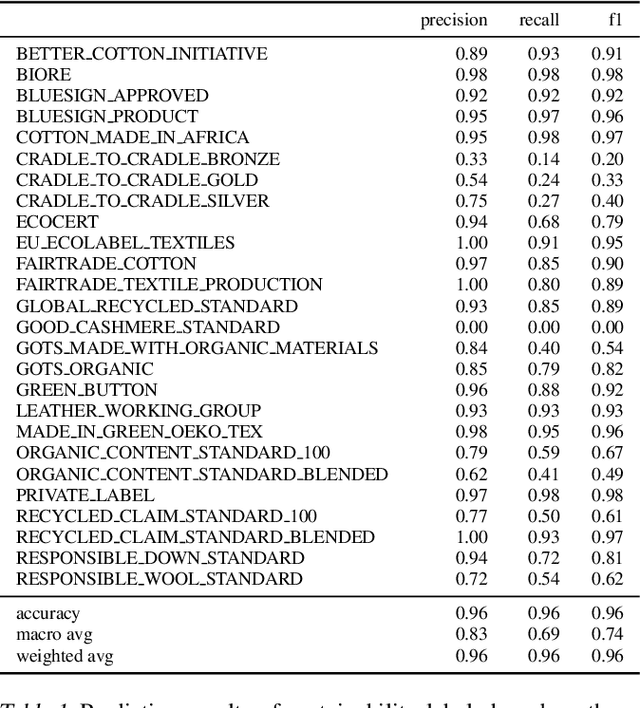
The production, shipping, usage, and disposal of consumer goods have a substantial impact on greenhouse gas emissions and the depletion of resources. Machine Learning (ML) can help to foster sustainable consumption patterns by accounting for sustainability aspects in product search or recommendations of modern retail platforms. However, the lack of large high quality publicly available product data with trustworthy sustainability information impedes the development of ML technology that can help to reach our sustainability goals. Here we present GreenDB, a database that collects products from European online shops on a weekly basis. As proxy for the products' sustainability, it relies on sustainability labels, which are evaluated by experts. The GreenDB schema extends the well-known schema.org Product definition and can be readily integrated into existing product catalogs. We present initial results demonstrating that ML models trained with our data can reliably (F1 score 96%) predict the sustainability label of products. These contributions can help to complement existing e-commerce experiences and ultimately encourage users to more sustainable consumption patterns.
Mutual Guidance and Residual Integration for Image Enhancement
Nov 25, 2022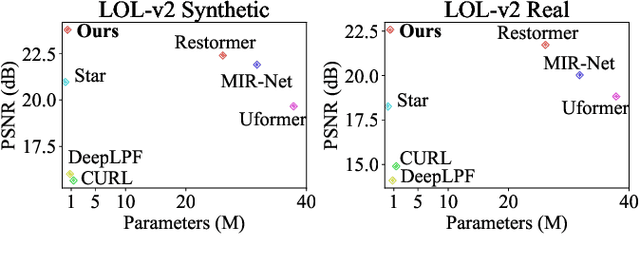
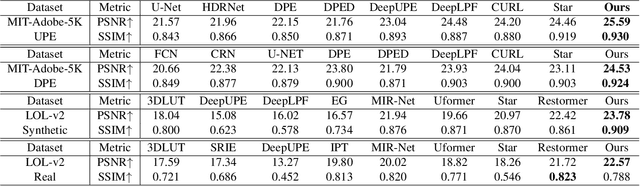

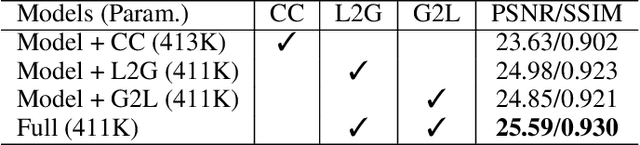
Previous studies show the necessity of global and local adjustment for image enhancement. However, existing convolutional neural networks (CNNs) and transformer-based models face great challenges in balancing the computational efficiency and effectiveness of global-local information usage. Especially, existing methods typically adopt the global-to-local fusion mode, ignoring the importance of bidirectional interactions. To address those issues, we propose a novel mutual guidance network (MGN) to perform effective bidirectional global-local information exchange while keeping a compact architecture. In our design, we adopt a two-branch framework where one branch focuses more on modeling global relations while the other is committed to processing local information. Then, we develop an efficient attention-based mutual guidance approach throughout our framework for bidirectional global-local interactions. As a result, both the global and local branches can enjoy the merits of mutual information aggregation. Besides, to further refine the results produced by our MGN, we propose a novel residual integration scheme following the divide-and-conquer philosophy. The extensive experiments demonstrate the effectiveness of our proposed method, which achieves state-of-the-art performance on several public image enhancement benchmarks.
Self-Supervised Correction Learning for Semi-Supervised Biomedical Image Segmentation
Jan 12, 2023Biomedical image segmentation plays a significant role in computer-aided diagnosis. However, existing CNN based methods rely heavily on massive manual annotations, which are very expensive and require huge human resources. In this work, we adopt a coarse-to-fine strategy and propose a self-supervised correction learning paradigm for semi-supervised biomedical image segmentation. Specifically, we design a dual-task network, including a shared encoder and two independent decoders for segmentation and lesion region inpainting, respectively. In the first phase, only the segmentation branch is used to obtain a relatively rough segmentation result. In the second step, we mask the detected lesion regions on the original image based on the initial segmentation map, and send it together with the original image into the network again to simultaneously perform inpainting and segmentation separately. For labeled data, this process is supervised by the segmentation annotations, and for unlabeled data, it is guided by the inpainting loss of masked lesion regions. Since the two tasks rely on similar feature information, the unlabeled data effectively enhances the representation of the network to the lesion regions and further improves the segmentation performance. Moreover, a gated feature fusion (GFF) module is designed to incorporate the complementary features from the two tasks. Experiments on three medical image segmentation datasets for different tasks including polyp, skin lesion and fundus optic disc segmentation well demonstrate the outstanding performance of our method compared with other semi-supervised approaches. The code is available at https://github.com/ReaFly/SemiMedSeg.
CovidRhythm: A Deep Learning Model for Passive Prediction of Covid-19 using Biobehavioral Rhythms Derived from Wearable Physiological Data
Jan 12, 2023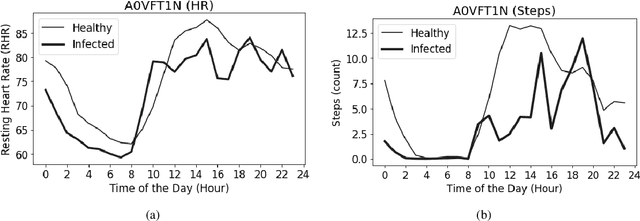
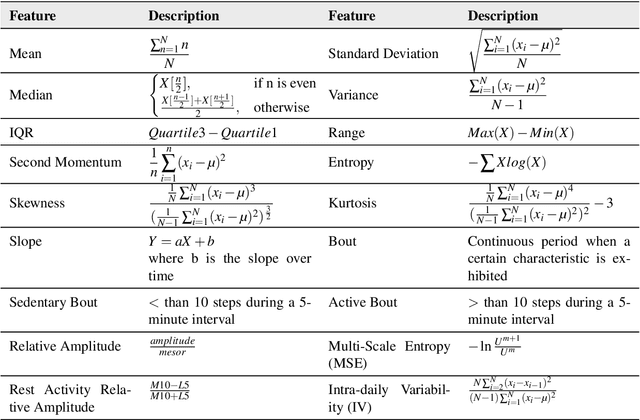
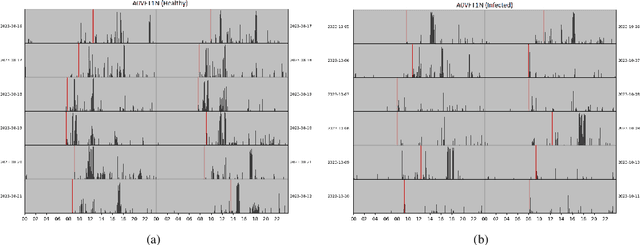
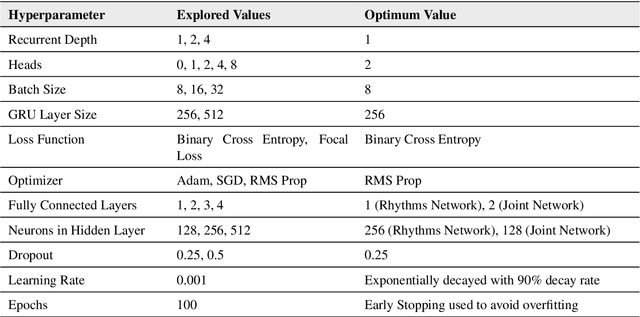
To investigate whether a deep learning model can detect Covid-19 from disruptions in the human body's physiological (heart rate) and rest-activity rhythms (rhythmic dysregulation) caused by the SARS-CoV-2 virus. We propose CovidRhythm, a novel Gated Recurrent Unit (GRU) Network with Multi-Head Self-Attention (MHSA) that combines sensor and rhythmic features extracted from heart rate and activity (steps) data gathered passively using consumer-grade smart wearable to predict Covid-19. A total of 39 features were extracted (standard deviation, mean, min/max/avg length of sedentary and active bouts) from wearable sensor data. Biobehavioral rhythms were modeled using nine parameters (mesor, amplitude, acrophase, and intra-daily variability). These features were then input to CovidRhythm for predicting Covid-19 in the incubation phase (one day before biological symptoms manifest). A combination of sensor and biobehavioral rhythm features achieved the highest AUC-ROC of 0.79 [Sensitivity = 0.69, Specificity=0.89, F$_{0.1}$ = 0.76], outperforming prior approaches in discriminating Covid-positive patients from healthy controls using 24 hours of historical wearable physiological. Rhythmic features were the most predictive of Covid-19 infection when utilized either alone or in conjunction with sensor features. Sensor features predicted healthy subjects best. Circadian rest-activity rhythms that combine 24h activity and sleep information were the most disrupted. CovidRhythm demonstrates that biobehavioral rhythms derived from consumer-grade wearable data can facilitate timely Covid-19 detection. To the best of our knowledge, our work is the first to detect Covid-19 using deep learning and biobehavioral rhythms features derived from consumer-grade wearable data.
Choosing the Number of Topics in LDA Models -- A Monte Carlo Comparison of Selection Criteria
Dec 28, 2022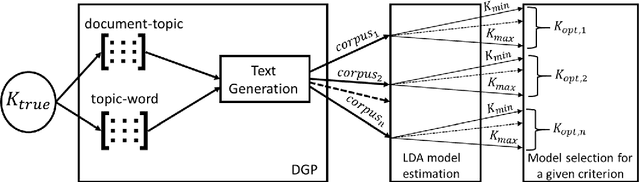
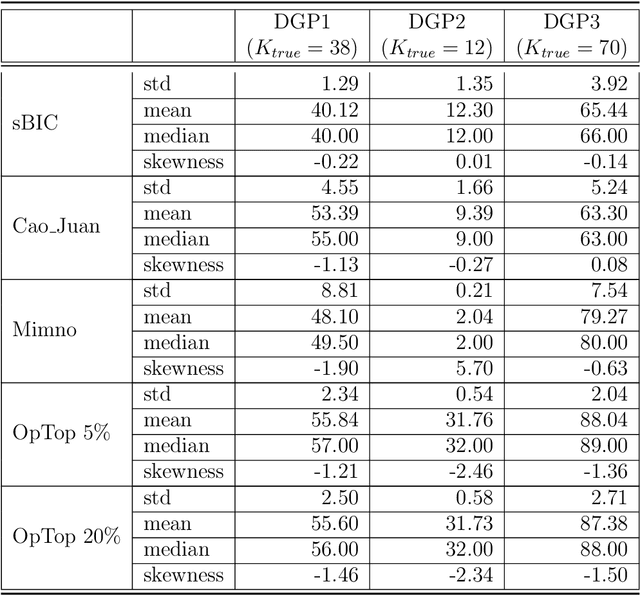
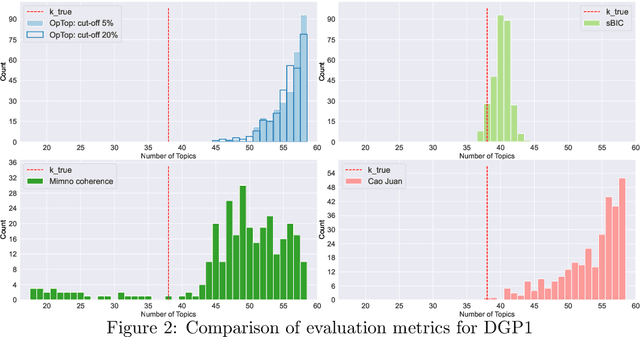
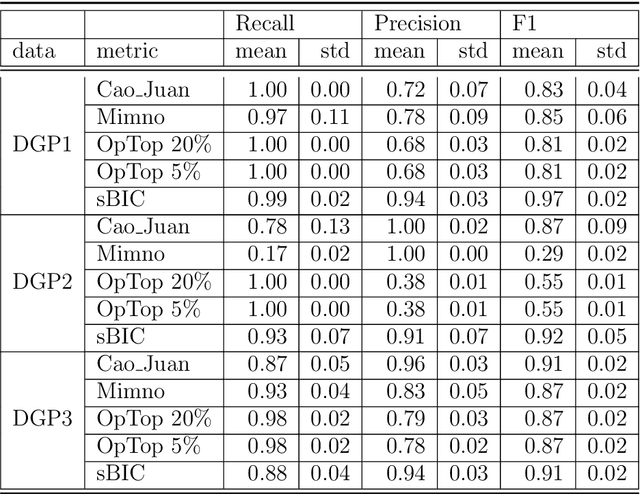
Selecting the number of topics in LDA models is considered to be a difficult task, for which alternative approaches have been proposed. The performance of the recently developed singular Bayesian information criterion (sBIC) is evaluated and compared to the performance of alternative model selection criteria. The sBIC is a generalization of the standard BIC that can be implemented to singular statistical models. The comparison is based on Monte Carlo simulations and carried out for several alternative settings, varying with respect to the number of topics, the number of documents and the size of documents in the corpora. Performance is measured using different criteria which take into account the correct number of topics, but also whether the relevant topics from the DGPs are identified. Practical recommendations for LDA model selection in applications are derived.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge