"Information": models, code, and papers
DEPAC: a Corpus for Depression and Anxiety Detection from Speech
Jun 20, 2023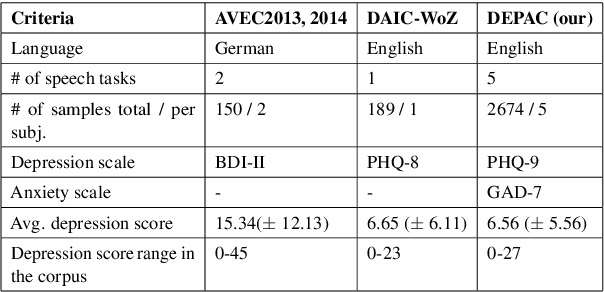
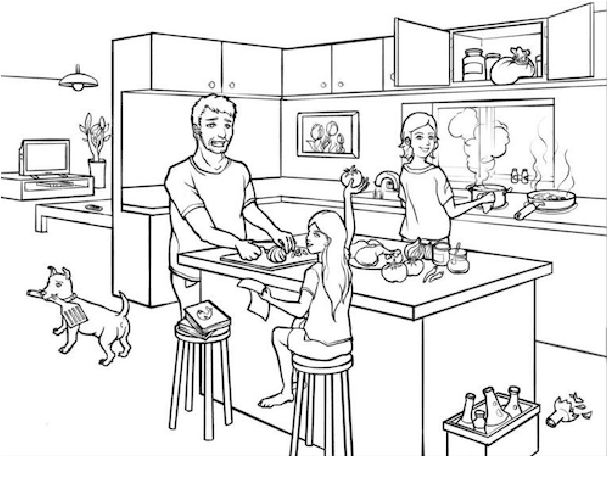

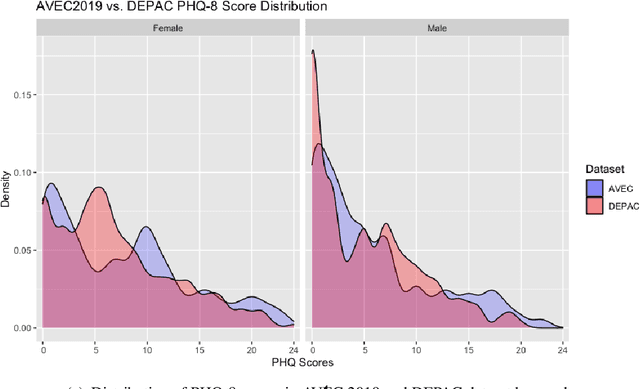
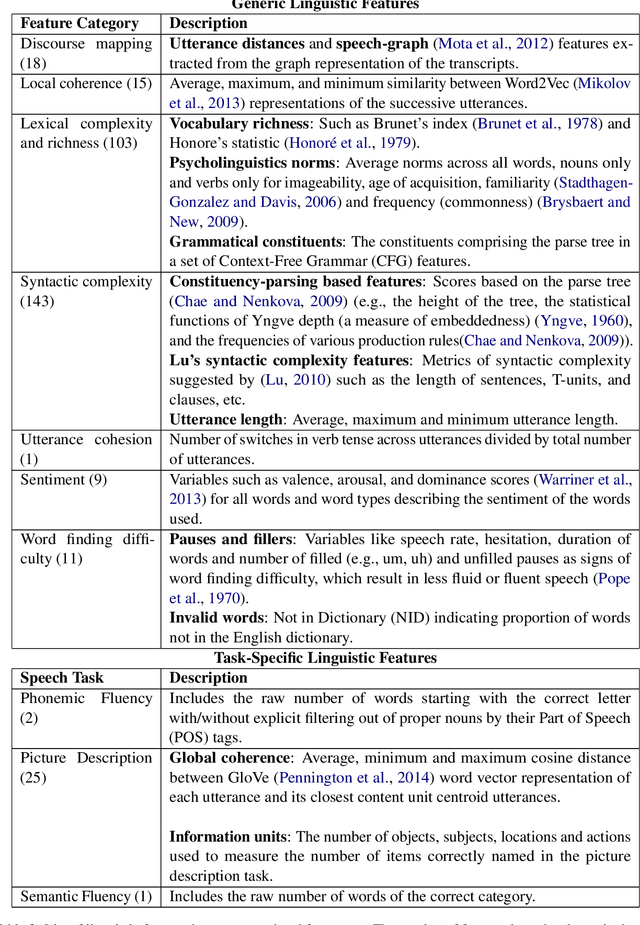
Mental distress like depression and anxiety contribute to the largest proportion of the global burden of diseases. Automated diagnosis systems of such disorders, empowered by recent innovations in Artificial Intelligence, can pave the way to reduce the sufferings of the affected individuals. Development of such systems requires information-rich and balanced corpora. In this work, we introduce a novel mental distress analysis audio dataset DEPAC, labeled based on established thresholds on depression and anxiety standard screening tools. This large dataset comprises multiple speech tasks per individual, as well as relevant demographic information. Alongside, we present a feature set consisting of hand-curated acoustic and linguistic features, which were found effective in identifying signs of mental illnesses in human speech. Finally, we justify the quality and effectiveness of our proposed audio corpus and feature set in predicting depression severity by comparing the performance of baseline machine learning models built on this dataset with baseline models trained on other well-known depression corpora.
LVM-Med: Learning Large-Scale Self-Supervised Vision Models for Medical Imaging via Second-order Graph Matching
Jul 09, 2023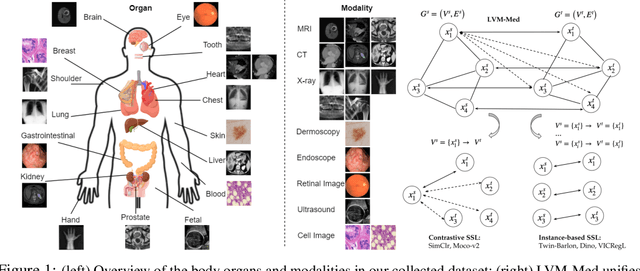
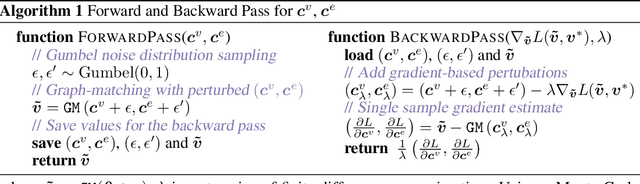

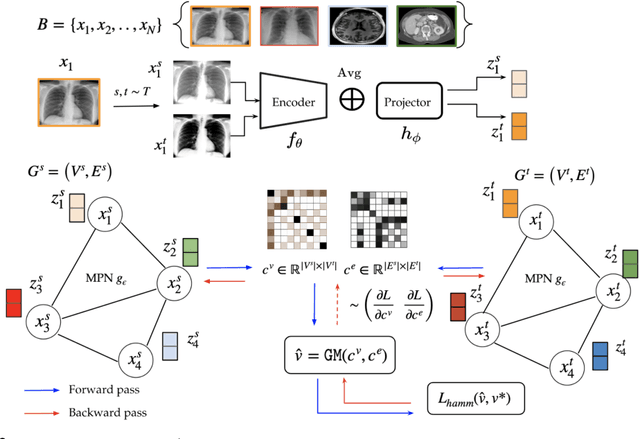
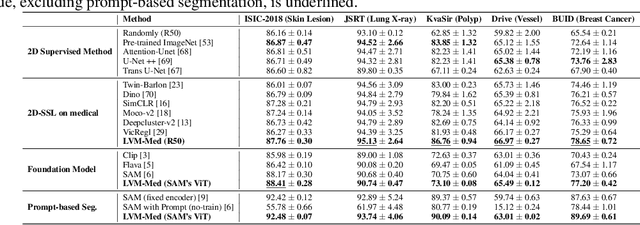
Obtaining large pre-trained models that can be fine-tuned to new tasks with limited annotated samples has remained an open challenge for medical imaging data. While pre-trained deep networks on ImageNet and vision-language foundation models trained on web-scale data are prevailing approaches, their effectiveness on medical tasks is limited due to the significant domain shift between natural and medical images. To bridge this gap, we introduce LVM-Med, the first family of deep networks trained on large-scale medical datasets. We have collected approximately 1.3 million medical images from 55 publicly available datasets, covering a large number of organs and modalities such as CT, MRI, X-ray, and Ultrasound. We benchmark several state-of-the-art self-supervised algorithms on this dataset and propose a novel self-supervised contrastive learning algorithm using a graph-matching formulation. The proposed approach makes three contributions: (i) it integrates prior pair-wise image similarity metrics based on local and global information; (ii) it captures the structural constraints of feature embeddings through a loss function constructed via a combinatorial graph-matching objective; and (iii) it can be trained efficiently end-to-end using modern gradient-estimation techniques for black-box solvers. We thoroughly evaluate the proposed LVM-Med on 15 downstream medical tasks ranging from segmentation and classification to object detection, and both for the in and out-of-distribution settings. LVM-Med empirically outperforms a number of state-of-the-art supervised, self-supervised, and foundation models. For challenging tasks such as Brain Tumor Classification or Diabetic Retinopathy Grading, LVM-Med improves previous vision-language models trained on 1 billion masks by 6-7% while using only a ResNet-50.
Learning World Models with Identifiable Factorization
Jun 11, 2023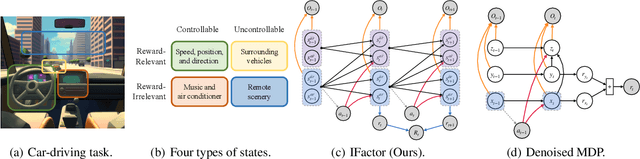
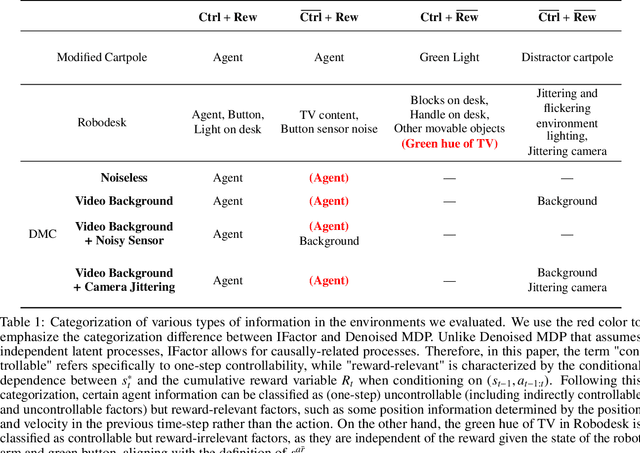
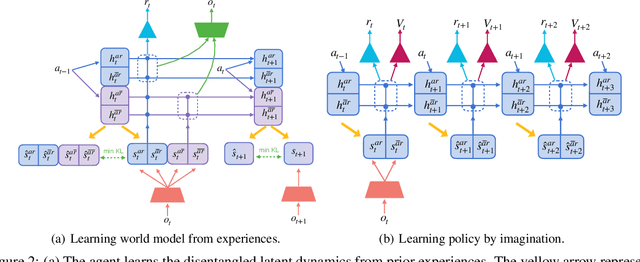
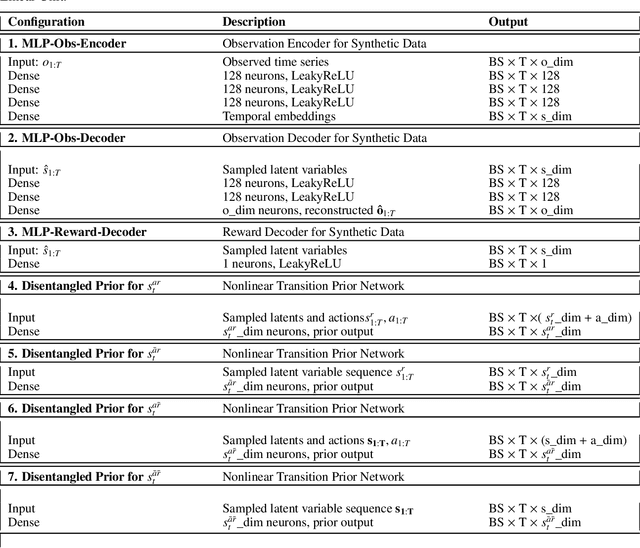
Extracting a stable and compact representation of the environment is crucial for efficient reinforcement learning in high-dimensional, noisy, and non-stationary environments. Different categories of information coexist in such environments -- how to effectively extract and disentangle these information remains a challenging problem. In this paper, we propose IFactor, a general framework to model four distinct categories of latent state variables that capture various aspects of information within the RL system, based on their interactions with actions and rewards. Our analysis establishes block-wise identifiability of these latent variables, which not only provides a stable and compact representation but also discloses that all reward-relevant factors are significant for policy learning. We further present a practical approach to learning the world model with identifiable blocks, ensuring the removal of redundants but retaining minimal and sufficient information for policy optimization. Experiments in synthetic worlds demonstrate that our method accurately identifies the ground-truth latent variables, substantiating our theoretical findings. Moreover, experiments in variants of the DeepMind Control Suite and RoboDesk showcase the superior performance of our approach over baselines.
Physics-informed invertible neural network for the Koopman operator learning
Jun 30, 2023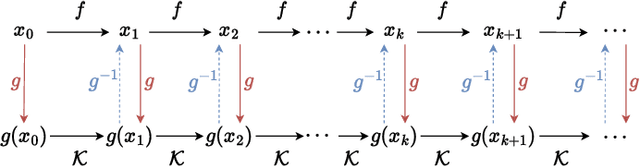

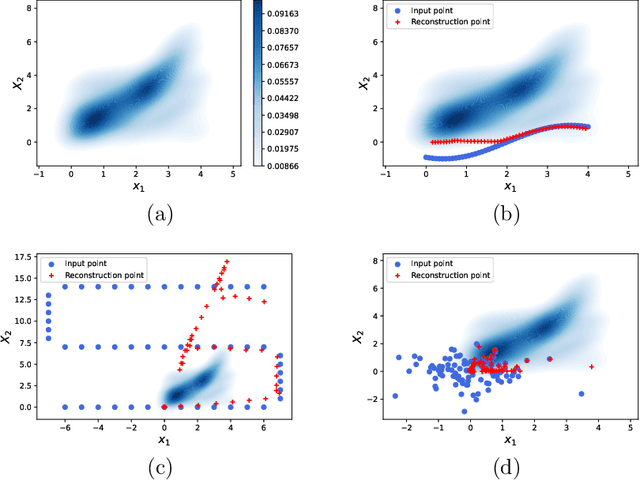
In Koopman operator theory, a finite-dimensional nonlinear system is transformed into an infinite but linear system using a set of observable functions. However, manually selecting observable functions that span the invariant subspace of the Koopman operator based on prior knowledge is inefficient and challenging, particularly when little or no information is available about the underlying systems. Furthermore, current methodologies tend to disregard the importance of the invertibility of observable functions, which leads to inaccurate results. To address these challenges, we propose the so-called FlowDMD, a Flow-based Dynamic Mode Decomposition that utilizes the Coupling Flow Invertible Neural Network (CF-INN) framework. FlowDMD leverages the intrinsically invertible characteristics of the CF-INN to learn the invariant subspaces of the Koopman operator and accurately reconstruct state variables. Numerical experiments demonstrate the superior performance of our algorithm compared to state-of-the-art methodologies.
A-STAR: Test-time Attention Segregation and Retention for Text-to-image Synthesis
Jun 26, 2023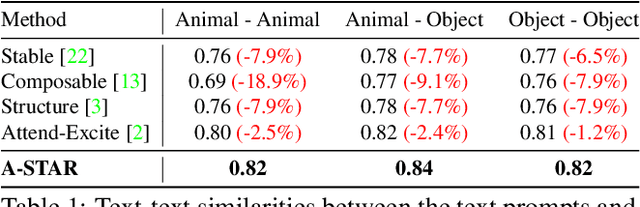
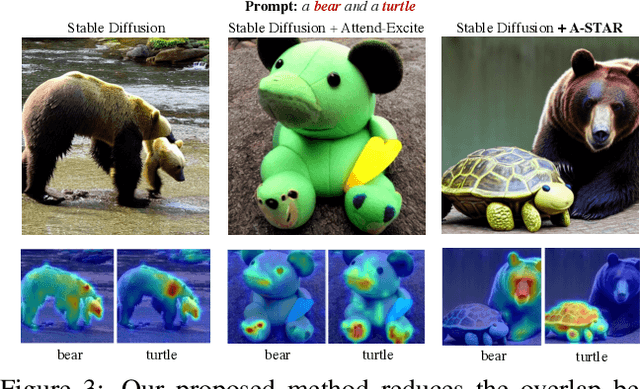
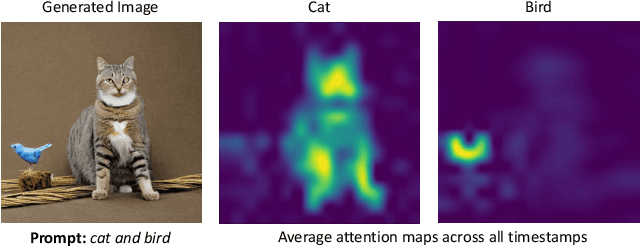
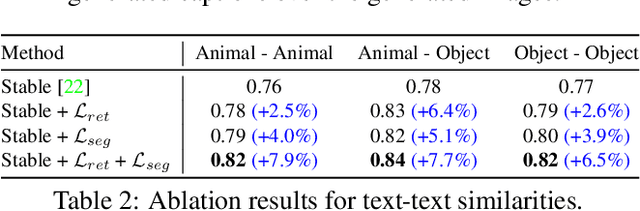

While recent developments in text-to-image generative models have led to a suite of high-performing methods capable of producing creative imagery from free-form text, there are several limitations. By analyzing the cross-attention representations of these models, we notice two key issues. First, for text prompts that contain multiple concepts, there is a significant amount of pixel-space overlap (i.e., same spatial regions) among pairs of different concepts. This eventually leads to the model being unable to distinguish between the two concepts and one of them being ignored in the final generation. Next, while these models attempt to capture all such concepts during the beginning of denoising (e.g., first few steps) as evidenced by cross-attention maps, this knowledge is not retained by the end of denoising (e.g., last few steps). Such loss of knowledge eventually leads to inaccurate generation outputs. To address these issues, our key innovations include two test-time attention-based loss functions that substantially improve the performance of pretrained baseline text-to-image diffusion models. First, our attention segregation loss reduces the cross-attention overlap between attention maps of different concepts in the text prompt, thereby reducing the confusion/conflict among various concepts and the eventual capture of all concepts in the generated output. Next, our attention retention loss explicitly forces text-to-image diffusion models to retain cross-attention information for all concepts across all denoising time steps, thereby leading to reduced information loss and the preservation of all concepts in the generated output.
Communication-Efficient Federated Learning through Importance Sampling
Jun 22, 2023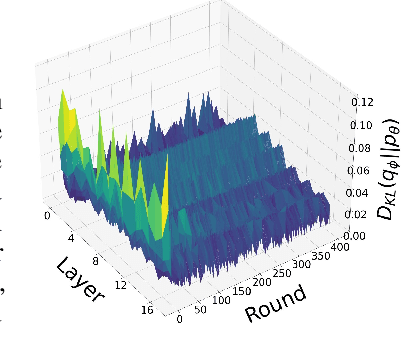
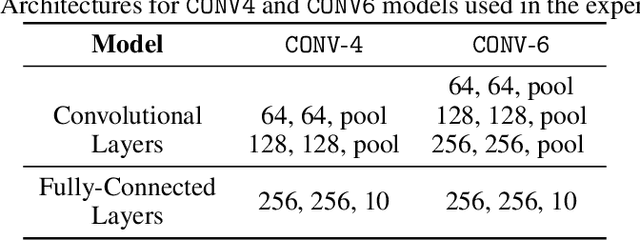
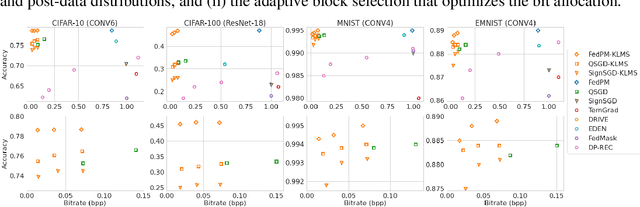
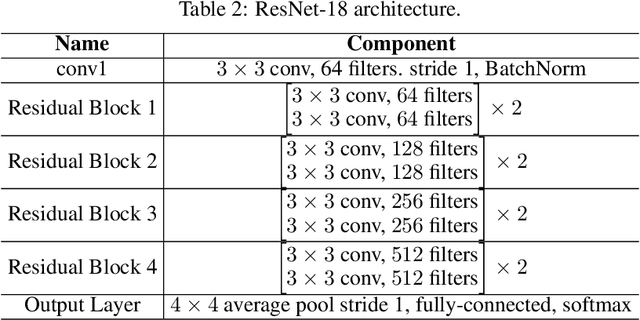
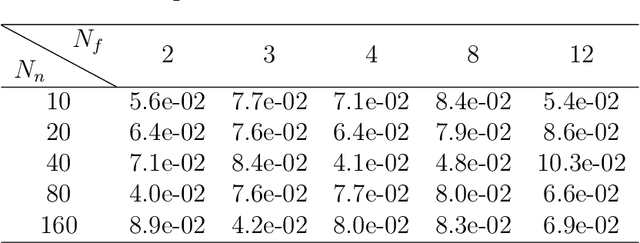
The high communication cost of sending model updates from the clients to the server is a significant bottleneck for scalable federated learning (FL). Among existing approaches, state-of-the-art bitrate-accuracy tradeoffs have been achieved using stochastic compression methods -- in which the client $n$ sends a sample from a client-only probability distribution $q_{\phi^{(n)}}$, and the server estimates the mean of the clients' distributions using these samples. However, such methods do not take full advantage of the FL setup where the server, throughout the training process, has side information in the form of a pre-data distribution $p_{\theta}$ that is close to the client's distribution $q_{\phi^{(n)}}$ in Kullback-Leibler (KL) divergence. In this work, we exploit this closeness between the clients' distributions $q_{\phi^{(n)}}$'s and the side information $p_{\theta}$ at the server, and propose a framework that requires approximately $D_{KL}(q_{\phi^{(n)}}|| p_{\theta})$ bits of communication. We show that our method can be integrated into many existing stochastic compression frameworks such as FedPM, Federated SGLD, and QSGD to attain the same (and often higher) test accuracy with up to $50$ times reduction in the bitrate.
PromptIR: Prompting for All-in-One Blind Image Restoration
Jun 22, 2023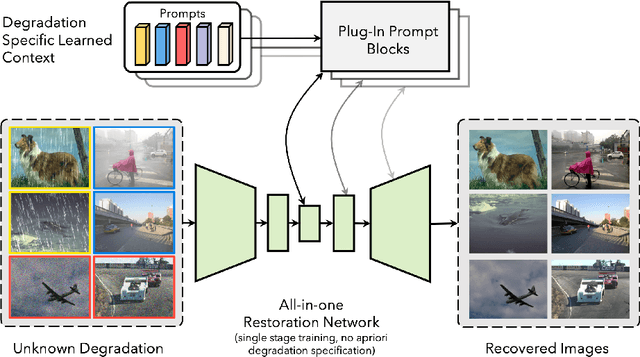
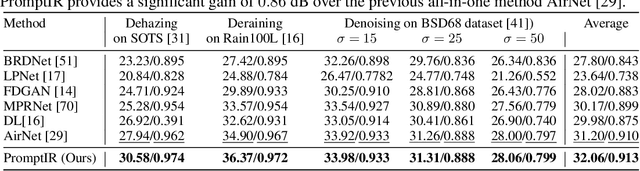
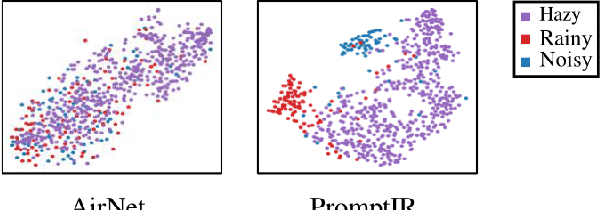

Image restoration involves recovering a high-quality clean image from its degraded version. Deep learning-based methods have significantly improved image restoration performance, however, they have limited generalization ability to different degradation types and levels. This restricts their real-world application since it requires training individual models for each specific degradation and knowing the input degradation type to apply the relevant model. We present a prompt-based learning approach, PromptIR, for All-In-One image restoration that can effectively restore images from various types and levels of degradation. In particular, our method uses prompts to encode degradation-specific information, which is then used to dynamically guide the restoration network. This allows our method to generalize to different degradation types and levels, while still achieving state-of-the-art results on image denoising, deraining, and dehazing. Overall, PromptIR offers a generic and efficient plugin module with few lightweight prompts that can be used to restore images of various types and levels of degradation with no prior information on the corruptions present in the image. Our code and pretrained models are available here: https://github.com/va1shn9v/PromptIR
Super-Resolution of BVOC Emission Maps Via Domain Adaptation
Jun 22, 2023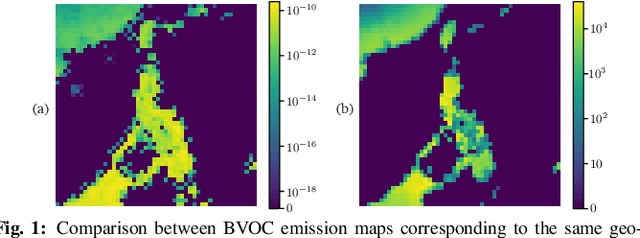
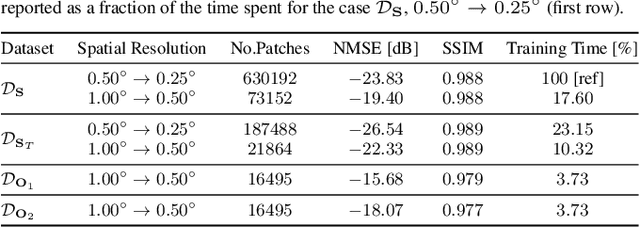
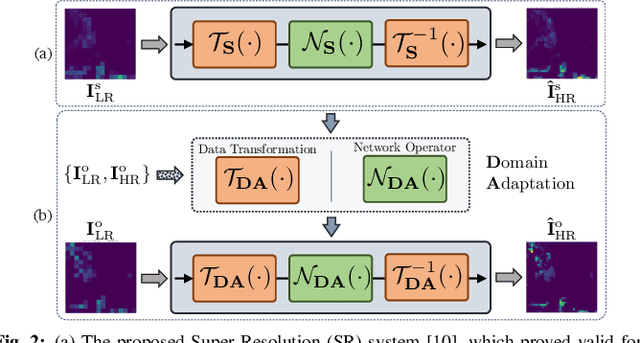
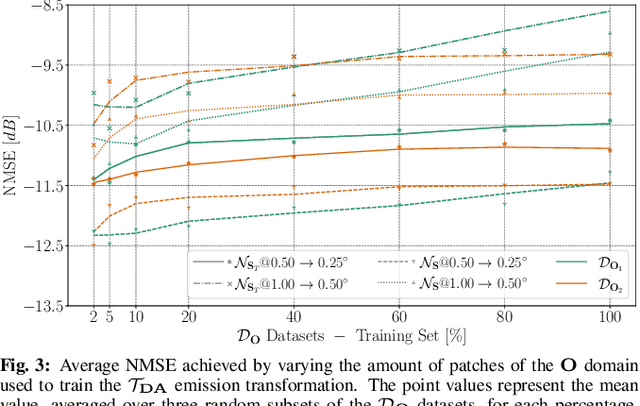
Enhancing the resolution of Biogenic Volatile Organic Compound (BVOC) emission maps is a critical task in remote sensing. Recently, some Super-Resolution (SR) methods based on Deep Learning (DL) have been proposed, leveraging data from numerical simulations for their training process. However, when dealing with data derived from satellite observations, the reconstruction is particularly challenging due to the scarcity of measurements to train SR algorithms with. In our work, we aim at super-resolving low resolution emission maps derived from satellite observations by leveraging the information of emission maps obtained through numerical simulations. To do this, we combine a SR method based on DL with Domain Adaptation (DA) techniques, harmonizing the different aggregation strategies and spatial information used in simulated and observed domains to ensure compatibility. We investigate the effectiveness of DA strategies at different stages by systematically varying the number of simulated and observed emissions used, exploring the implications of data scarcity on the adaptation strategies. To the best of our knowledge, there are no prior investigations of DA in satellite-derived BVOC maps enhancement. Our work represents a first step toward the development of robust strategies for the reconstruction of observed BVOC emissions.
MIRACLE: Multi-task Learning based Interpretable Regulation of Autoimmune Diseases through Common Latent Epigenetics
Jun 24, 2023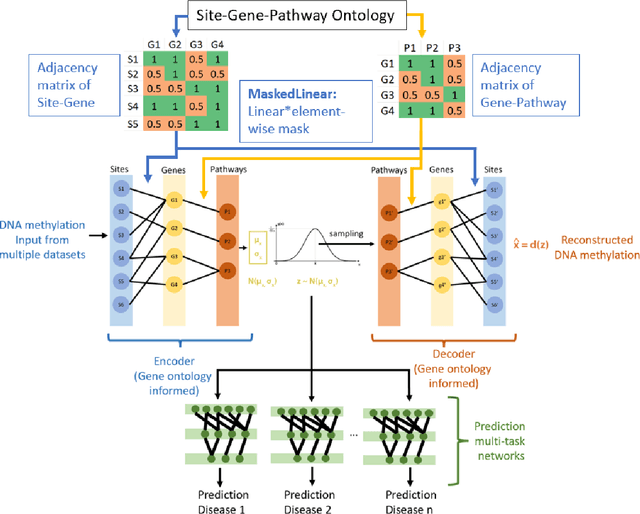
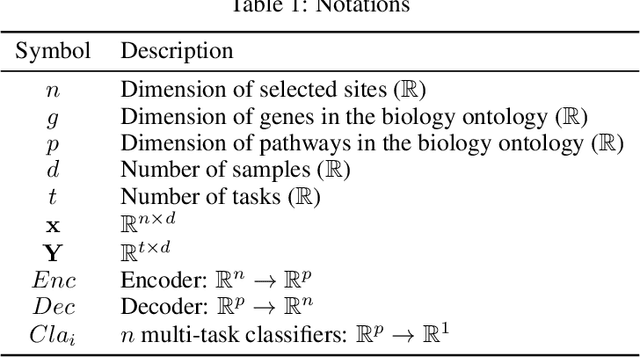

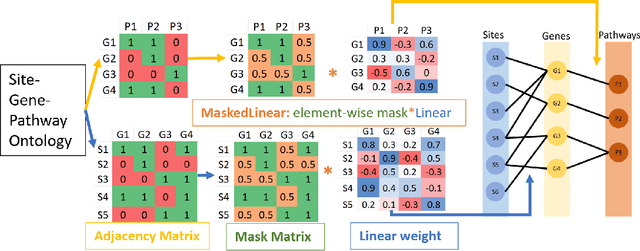
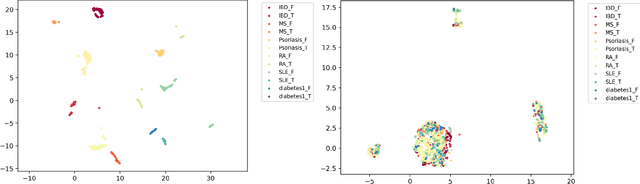
DNA methylation is a crucial regulator of gene transcription and has been linked to various diseases, including autoimmune diseases and cancers. However, diagnostics based on DNA methylation face challenges due to large feature sets and small sample sizes, resulting in overfitting and suboptimal performance. To address these issues, we propose MIRACLE, a novel interpretable neural network that leverages autoencoder-based multi-task learning to integrate multiple datasets and jointly identify common patterns in DNA methylation. MIRACLE's architecture reflects the relationships between methylation sites, genes, and pathways, ensuring biological interpretability and meaningfulness. The network comprises an encoder and a decoder, with a bottleneck layer representing pathway information as the basic unit of heredity. Customized defined MaskedLinear Layer is constrained by site-gene-pathway graph adjacency matrix information, which provides explainability and expresses the site-gene-pathway hierarchical structure explicitly. And from the embedding, there are different multi-task classifiers to predict diseases. Tested on six datasets, including rheumatoid arthritis, systemic lupus erythematosus, multiple sclerosis, inflammatory bowel disease, psoriasis, and type 1 diabetes, MIRACLE demonstrates robust performance in identifying common functions of DNA methylation across different phenotypes, with higher accuracy in prediction dieseases than baseline methods. By incorporating biological prior knowledge, MIRACLE offers a meaningful and interpretable framework for DNA methylation data analysis in the context of autoimmune diseases.
TACOformer:Token-channel compounded Cross Attention for Multimodal Emotion Recognition
Jun 23, 2023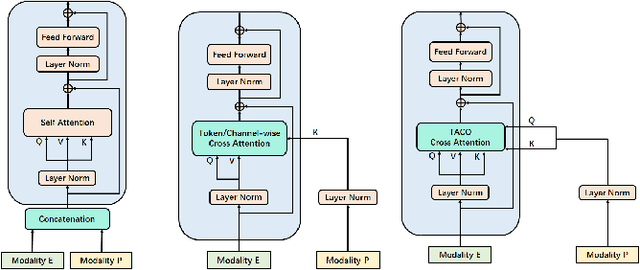
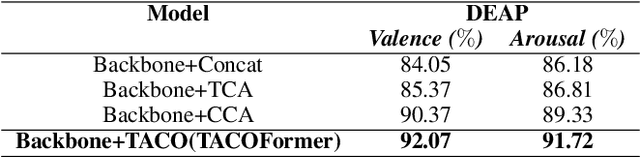
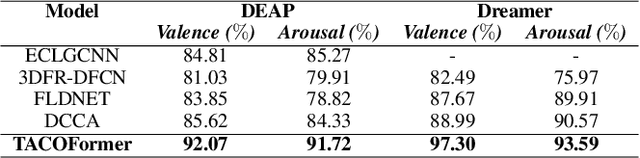
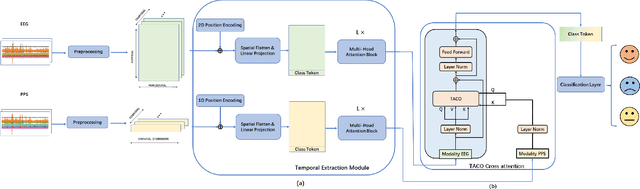

Recently, emotion recognition based on physiological signals has emerged as a field with intensive research. The utilization of multi-modal, multi-channel physiological signals has significantly improved the performance of emotion recognition systems, due to their complementarity. However, effectively integrating emotion-related semantic information from different modalities and capturing inter-modal dependencies remains a challenging issue. Many existing multimodal fusion methods ignore either token-to-token or channel-to-channel correlations of multichannel signals from different modalities, which limits the classification capability of the models to some extent. In this paper, we propose a comprehensive perspective of multimodal fusion that integrates channel-level and token-level cross-modal interactions. Specifically, we introduce a unified cross attention module called Token-chAnnel COmpound (TACO) Cross Attention to perform multimodal fusion, which simultaneously models channel-level and token-level dependencies between modalities. Additionally, we propose a 2D position encoding method to preserve information about the spatial distribution of EEG signal channels, then we use two transformer encoders ahead of the fusion module to capture long-term temporal dependencies from the EEG signal and the peripheral physiological signal, respectively. Subject-independent experiments on emotional dataset DEAP and Dreamer demonstrate that the proposed model achieves state-of-the-art performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge