"Information": models, code, and papers
Age of Gossip on a Grid
Jul 17, 2023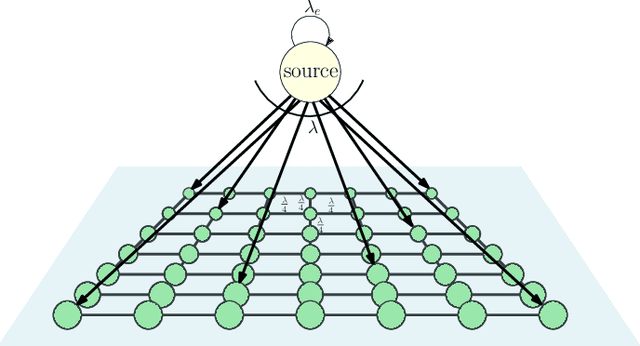
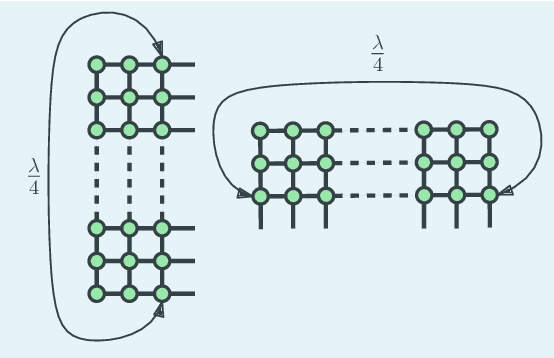
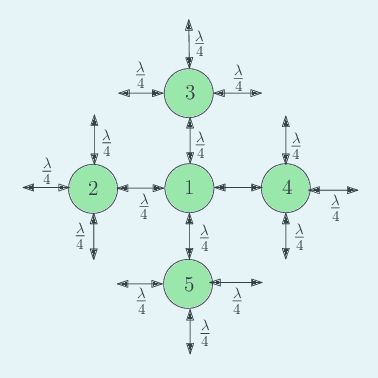
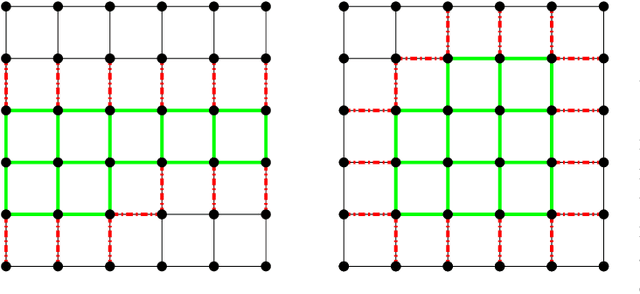
We consider a gossip network consisting of a source generating updates and $n$ nodes connected in a two-dimensional square grid. The source keeps updates of a process, that might be generated or observed, and shares them with the grid network. The nodes in the grid network communicate with their neighbors and disseminate these version updates using a push-style gossip strategy. We use the version age metric to quantify the timeliness of information at the nodes. We find an upper bound for the average version age for a set of nodes in a general network. Using this, we show that the average version age at a node scales as $O(n^{\frac{1}{3}})$ in a grid network. Prior to our work, it has been known that when $n$ nodes are connected on a ring the version age scales as $O(n^{\frac{1}{2}})$, and when they are connected on a fully-connected graph the version age scales as $O(\log n)$. Ours is the first work to show an age scaling result for a connectivity structure other than the ring and fully-connected networks that represent two extremes of network connectivity. Our work shows that higher connectivity on a grid compared to a ring lowers the age experience of each node from $O(n^{\frac{1}{2}})$ to $O(n^{\frac{1}{3}})$.
CoAD: Automatic Diagnosis through Symptom and Disease Collaborative Generation
Jul 17, 2023
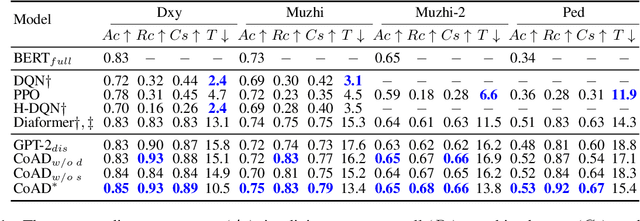
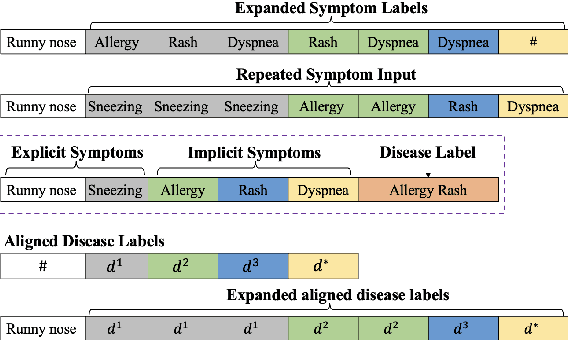
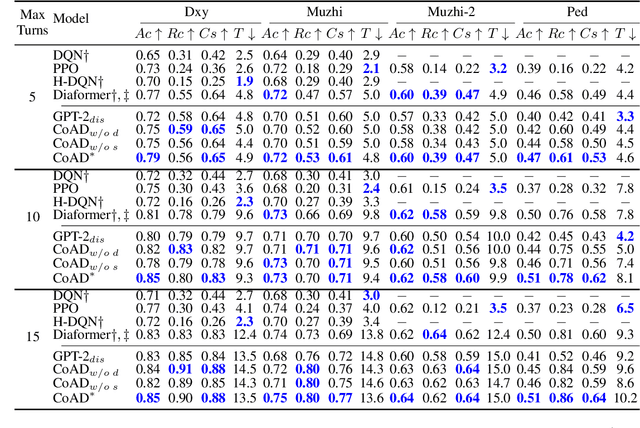
Automatic diagnosis (AD), a critical application of AI in healthcare, employs machine learning techniques to assist doctors in gathering patient symptom information for precise disease diagnosis. The Transformer-based method utilizes an input symptom sequence, predicts itself through auto-regression, and employs the hidden state of the final symptom to determine the disease. Despite its simplicity and superior performance demonstrated, a decline in disease diagnosis accuracy is observed caused by 1) a mismatch between symptoms observed during training and generation, and 2) the effect of different symptom orders on disease prediction. To address the above obstacles, we introduce the CoAD, a novel disease and symptom collaborative generation framework, which incorporates several key innovations to improve AD: 1) aligning sentence-level disease labels with multiple possible symptom inquiry steps to bridge the gap between training and generation; 2) expanding symptom labels for each sub-sequence of symptoms to enhance annotation and eliminate the effect of symptom order; 3) developing a repeated symptom input schema to effectively and efficiently learn the expanded disease and symptom labels. We evaluate the CoAD framework using four datasets, including three public and one private, and demonstrate that it achieves an average 2.3% improvement over previous state-of-the-art results in automatic disease diagnosis. For reproducibility, we release the code and data at https://github.com/KwanWaiChung/coad.
Cross-Language Speech Emotion Recognition Using Multimodal Dual Attention Transformers
Jul 14, 2023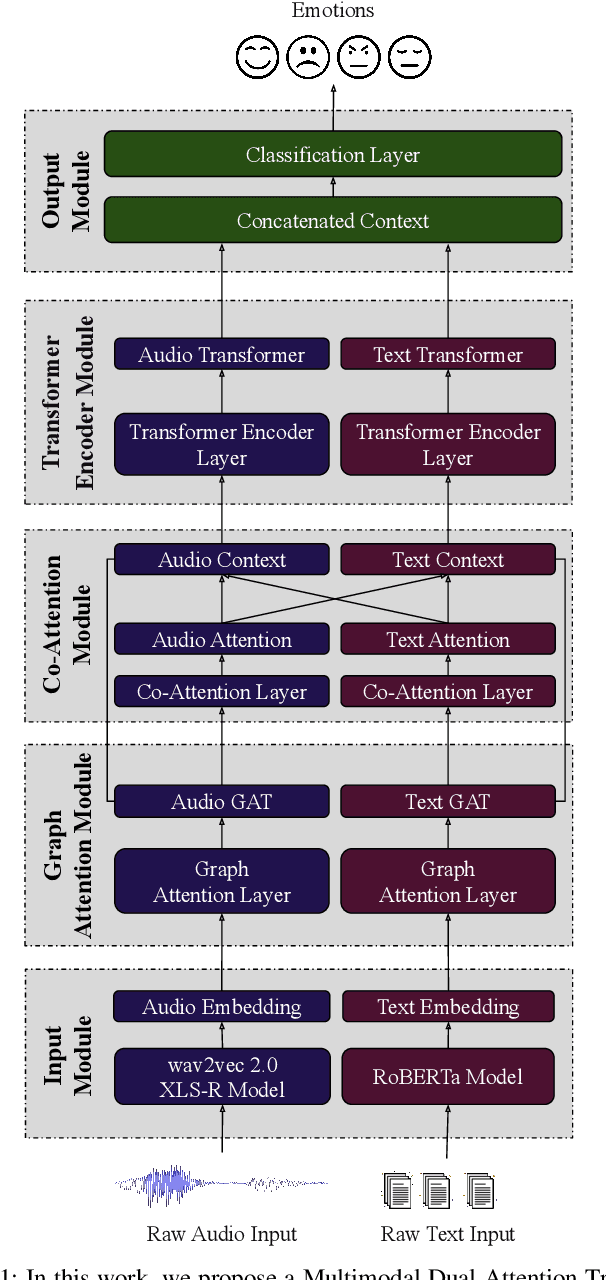
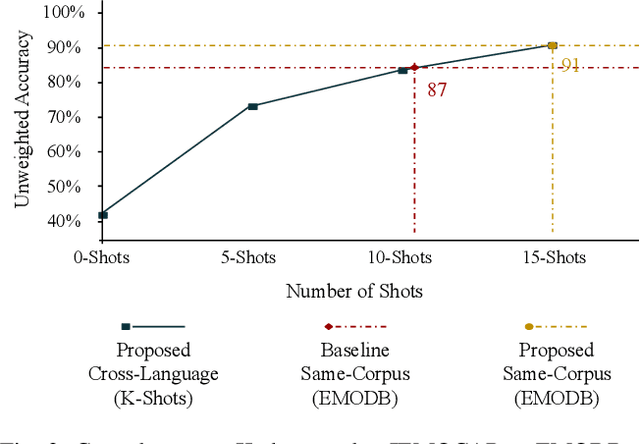
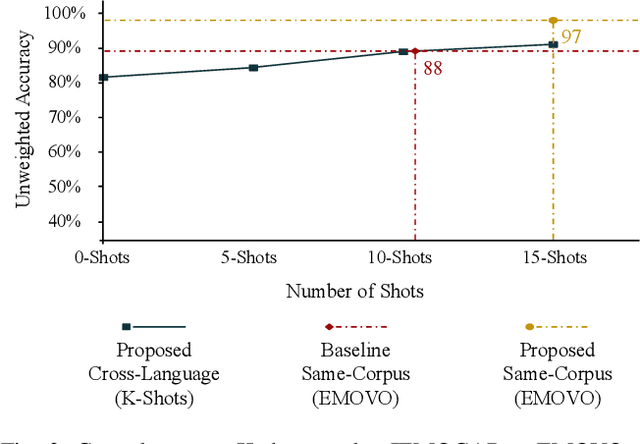
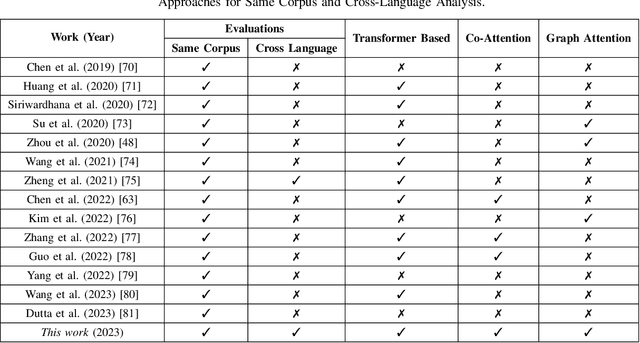
Despite the recent progress in speech emotion recognition (SER), state-of-the-art systems are unable to achieve improved performance in cross-language settings. In this paper, we propose a Multimodal Dual Attention Transformer (MDAT) model to improve cross-language SER. Our model utilises pre-trained models for multimodal feature extraction and is equipped with a dual attention mechanism including graph attention and co-attention to capture complex dependencies across different modalities and achieve improved cross-language SER results using minimal target language data. In addition, our model also exploits a transformer encoder layer for high-level feature representation to improve emotion classification accuracy. In this way, MDAT performs refinement of feature representation at various stages and provides emotional salient features to the classification layer. This novel approach also ensures the preservation of modality-specific emotional information while enhancing cross-modality and cross-language interactions. We assess our model's performance on four publicly available SER datasets and establish its superior effectiveness compared to recent approaches and baseline models.
SLSSNN: High energy efficiency spike-train level spiking neural networks with spatio-temporal conversion
Jul 14, 2023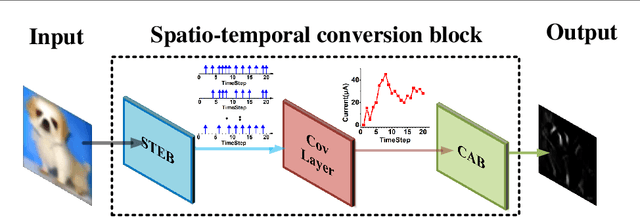
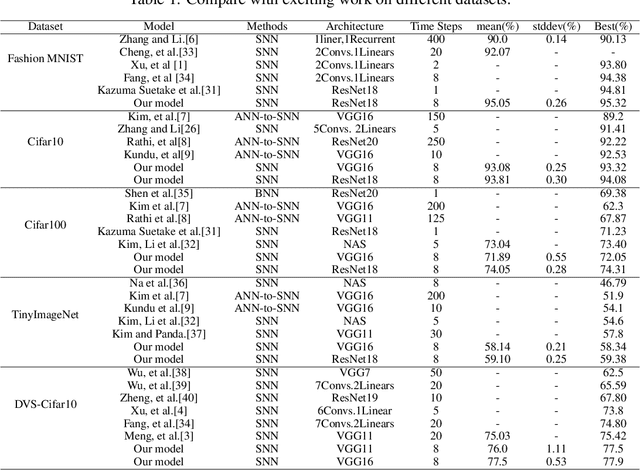
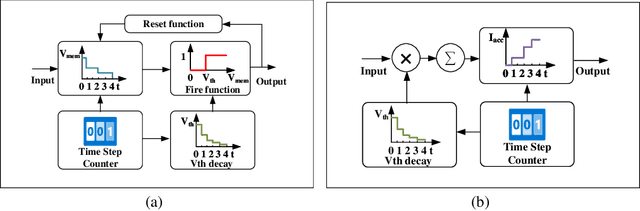
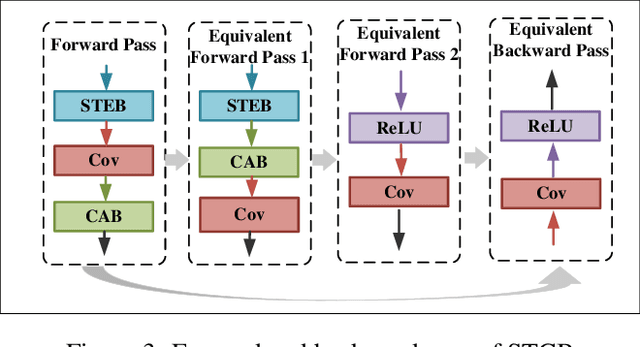
Brain-inspired spiking neuron networks (SNNs) have attracted widespread research interest due to their low power features, high biological plausibility, and strong spatiotemporal information processing capability. Although adopting a surrogate gradient (SG) makes the non-differentiability SNN trainable, achieving comparable accuracy for ANNs and keeping low-power features simultaneously is still tricky. In this paper, we proposed an energy-efficient spike-train level spiking neural network (SLSSNN) with low computational cost and high accuracy. In the SLSSNN, spatio-temporal conversion blocks (STCBs) are applied to replace the convolutional and ReLU layers to keep the low power features of SNNs and improve accuracy. However, SLSSNN cannot adopt backpropagation algorithms directly due to the non-differentiability nature of spike trains. We proposed a suitable learning rule for SLSSNNs by deducing the equivalent gradient of STCB. We evaluate the proposed SLSSNN on static and neuromorphic datasets, including Fashion-Mnist, Cifar10, Cifar100, TinyImageNet, and DVS-Cifar10. The experiment results show that our proposed SLSSNN outperforms the state-of-the-art accuracy on nearly all datasets, using fewer time steps and being highly energy-efficient.
Value-based Fast and Slow AI Nudging
Jul 14, 2023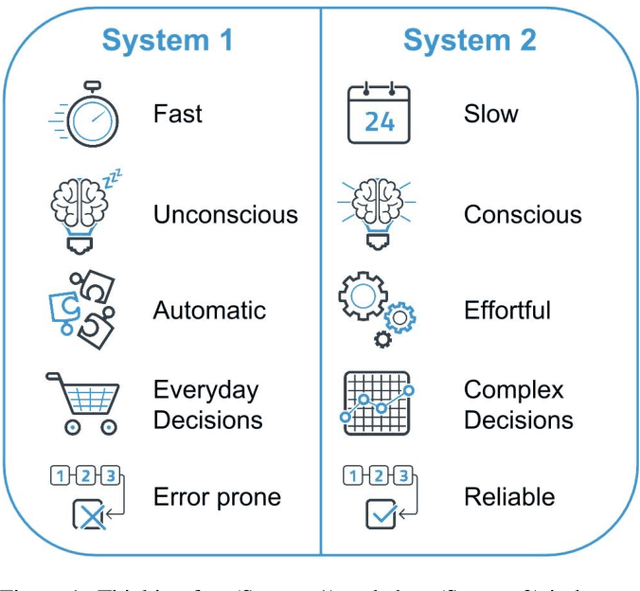
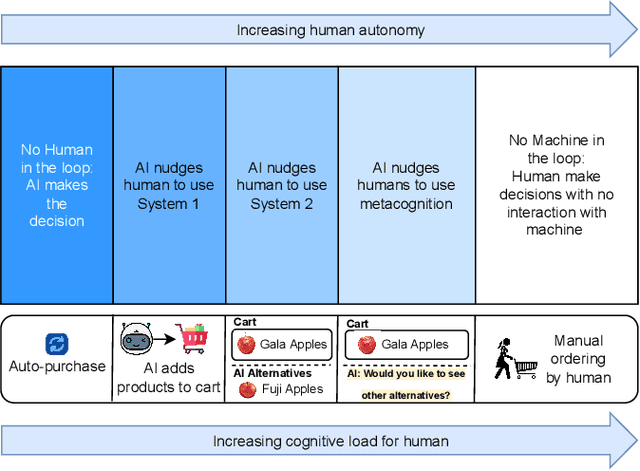
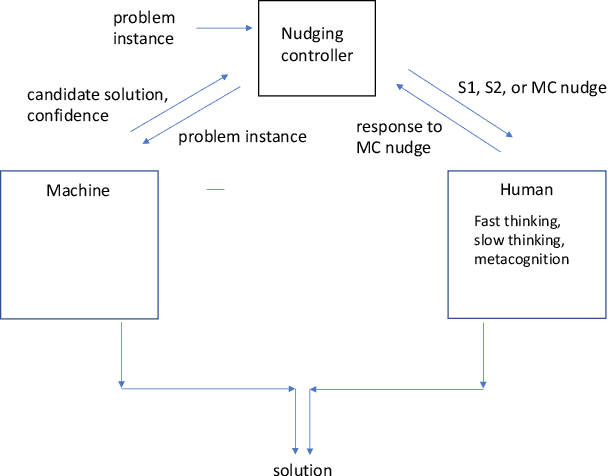
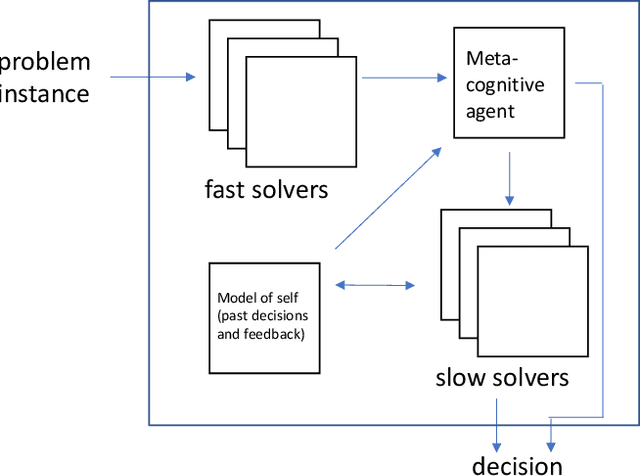
Nudging is a behavioral strategy aimed at influencing people's thoughts and actions. Nudging techniques can be found in many situations in our daily lives, and these nudging techniques can targeted at human fast and unconscious thinking, e.g., by using images to generate fear or the more careful and effortful slow thinking, e.g., by releasing information that makes us reflect on our choices. In this paper, we propose and discuss a value-based AI-human collaborative framework where AI systems nudge humans by proposing decision recommendations. Three different nudging modalities, based on when recommendations are presented to the human, are intended to stimulate human fast thinking, slow thinking, or meta-cognition. Values that are relevant to a specific decision scenario are used to decide when and how to use each of these nudging modalities. Examples of values are decision quality, speed, human upskilling and learning, human agency, and privacy. Several values can be present at the same time, and their priorities can vary over time. The framework treats values as parameters to be instantiated in a specific decision environment.
milliFlow: Scene Flow Estimation on mmWave Radar Point Cloud for Human Motion Sensing
Jul 03, 2023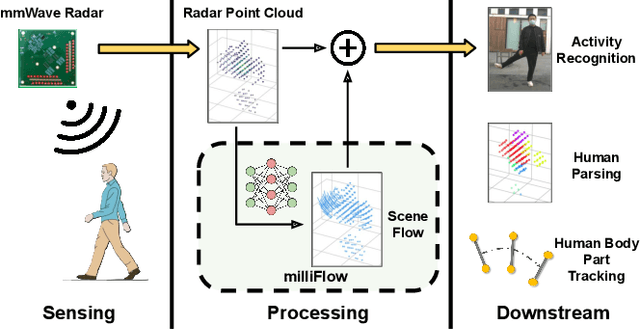
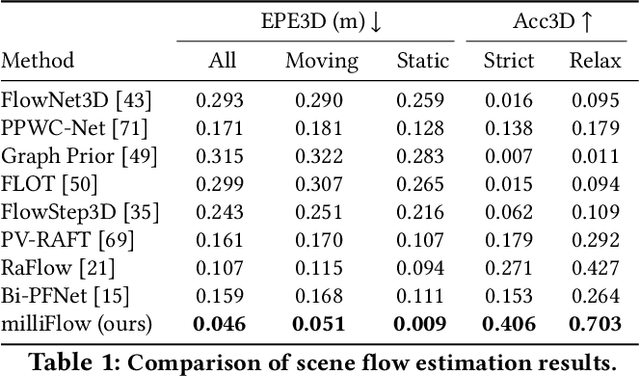
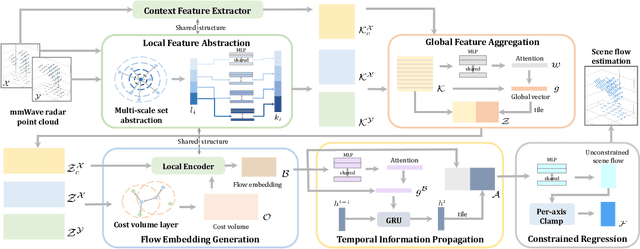
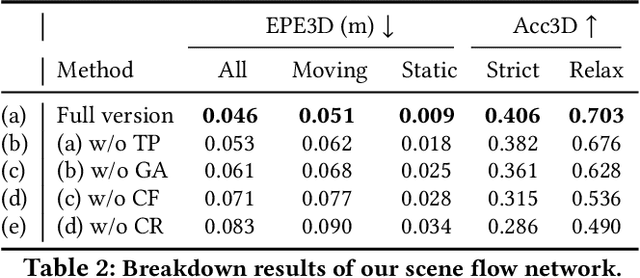
Approaching the era of ubiquitous computing, human motion sensing plays a crucial role in smart systems for decision making, user interaction, and personalized services. Extensive research has been conducted on human tracking, pose estimation, gesture recognition, and activity recognition, which are predominantly based on cameras in traditional methods. However, the intrusive nature of cameras limits their use in smart home applications. To address this, mmWave radars have gained popularity due to their privacy-friendly features. In this work, we propose \textit{milliFlow}, a novel deep learning method for scene flow estimation as a complementary motion information for mmWave point cloud, serving as an intermediate level of features and directly benefiting downstream human motion sensing tasks. Experimental results demonstrate the superior performance of our method with an average 3D endpoint error of 4.6cm, significantly surpassing the competing approaches. Furthermore, by incorporating scene flow information, we achieve remarkable improvements in human activity recognition, human parsing, and human body part tracking. To foster further research in this area, we provide our codebase and dataset for open access.
Enhancing Mapless Trajectory Prediction through Knowledge Distillation
Jun 25, 2023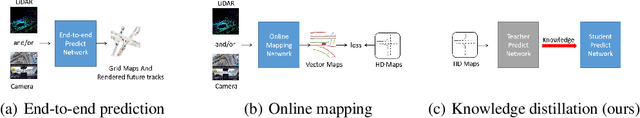
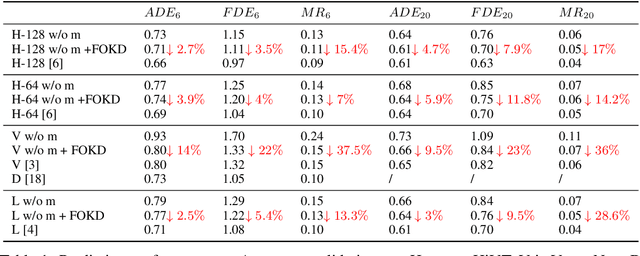
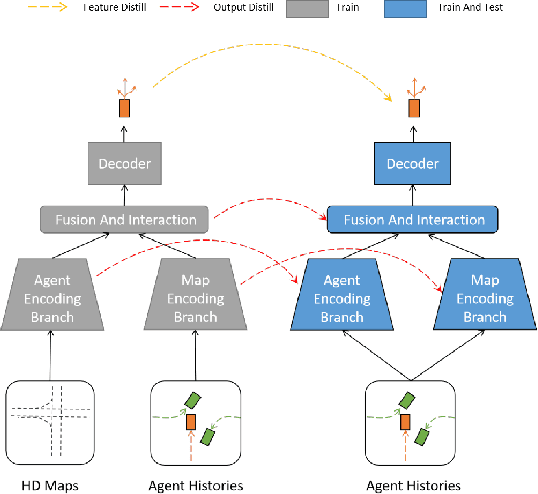
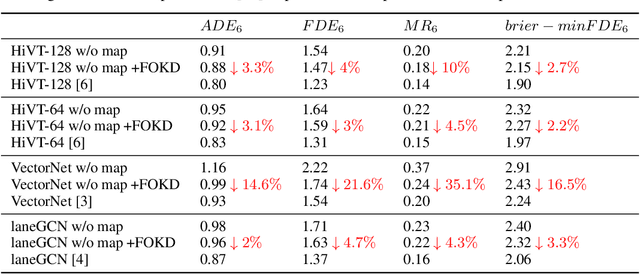
Scene information plays a crucial role in trajectory forecasting systems for autonomous driving by providing semantic clues and constraints on potential future paths of traffic agents. Prevalent trajectory prediction techniques often take high-definition maps (HD maps) as part of the inputs to provide scene knowledge. Although HD maps offer accurate road information, they may suffer from the high cost of annotation or restrictions of law that limits their widespread use. Therefore, those methods are still expected to generate reliable prediction results in mapless scenarios. In this paper, we tackle the problem of improving the consistency of multi-modal prediction trajectories and the real road topology when map information is unavailable during the test phase. Specifically, we achieve this by training a map-based prediction teacher network on the annotated samples and transferring the knowledge to a student mapless prediction network using a two-fold knowledge distillation framework. Our solution is generalizable for common trajectory prediction networks and does not bring extra computation burden. Experimental results show that our method stably improves prediction performance in mapless mode on many widely used state-of-the-art trajectory prediction baselines, compensating for the gaps caused by the absence of HD maps. Qualitative visualization results demonstrate that our approach helps infer unseen map information.
SHAMSUL: Simultaneous Heatmap-Analysis to investigate Medical Significance Utilizing Local interpretability methods
Jul 16, 2023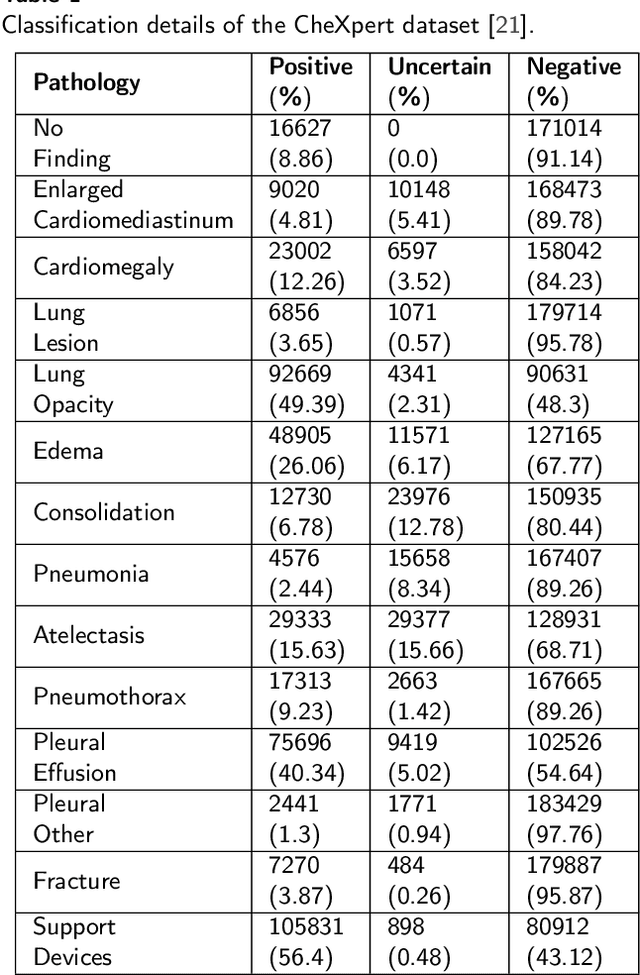
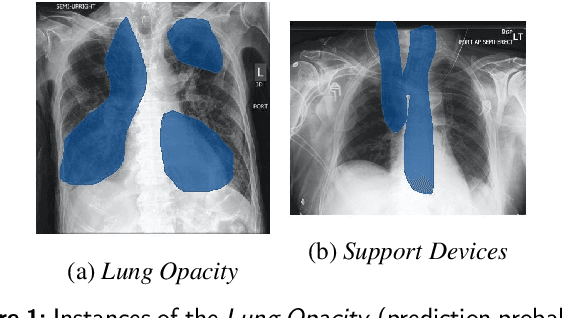
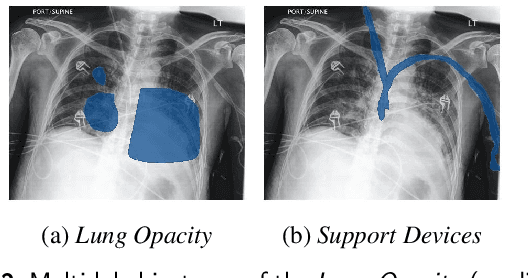
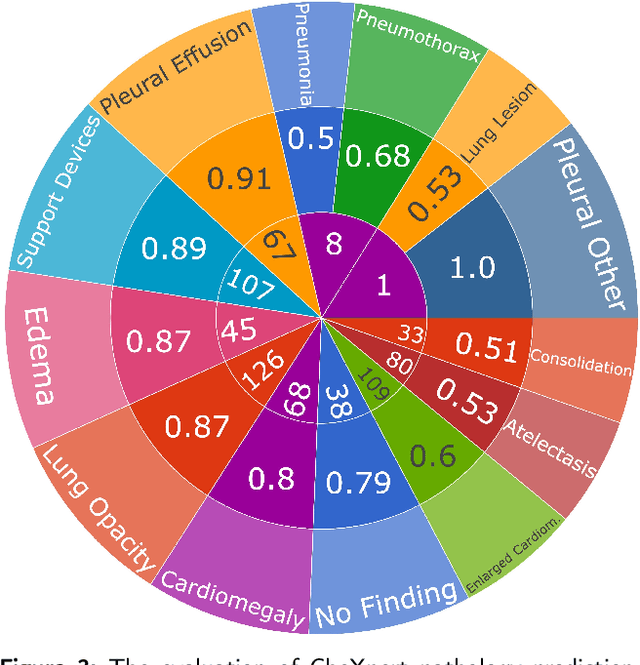
The interpretability of deep neural networks has become a subject of great interest within the medical and healthcare domain. This attention stems from concerns regarding transparency, legal and ethical considerations, and the medical significance of predictions generated by these deep neural networks in clinical decision support systems. To address this matter, our study delves into the application of four well-established interpretability methods: Local Interpretable Model-agnostic Explanations (LIME), Shapley Additive exPlanations (SHAP), Gradient-weighted Class Activation Mapping (Grad-CAM), and Layer-wise Relevance Propagation (LRP). Leveraging the approach of transfer learning with a multi-label-multi-class chest radiography dataset, we aim to interpret predictions pertaining to specific pathology classes. Our analysis encompasses both single-label and multi-label predictions, providing a comprehensive and unbiased assessment through quantitative and qualitative investigations, which are compared against human expert annotation. Notably, Grad-CAM demonstrates the most favorable performance in quantitative evaluation, while the LIME heatmap segmentation visualization exhibits the highest level of medical significance. Our research highlights the strengths and limitations of these interpretability methods and suggests that a multimodal-based approach, incorporating diverse sources of information beyond chest radiography images, could offer additional insights for enhancing interpretability in the medical domain.
Untrained neural network embedded Fourier phase retrieval from few measurements
Jul 16, 2023Fourier phase retrieval (FPR) is a challenging task widely used in various applications. It involves recovering an unknown signal from its Fourier phaseless measurements. FPR with few measurements is important for reducing time and hardware costs, but it suffers from serious ill-posedness. Recently, untrained neural networks have offered new approaches by introducing learned priors to alleviate the ill-posedness without requiring any external data. However, they may not be ideal for reconstructing fine details in images and can be computationally expensive. This paper proposes an untrained neural network (NN) embedded algorithm based on the alternating direction method of multipliers (ADMM) framework to solve FPR with few measurements. Specifically, we use a generative network to represent the image to be recovered, which confines the image to the space defined by the network structure. To improve the ability to represent high-frequency information, total variation (TV) regularization is imposed to facilitate the recovery of local structures in the image. Furthermore, to reduce the computational cost mainly caused by the parameter updates of the untrained NN, we develop an accelerated algorithm that adaptively trades off between explicit and implicit regularization. Experimental results indicate that the proposed algorithm outperforms existing untrained NN-based algorithms with fewer computational resources and even performs competitively against trained NN-based algorithms.
Generative Language Models on Nucleotide Sequences of Human Genes
Jul 20, 2023Language models, primarily transformer-based ones, obtained colossal success in NLP. To be more precise, studies like BERT in NLU and works such as GPT-3 for NLG are very crucial. DNA sequences are very close to natural language in terms of structure, so if the DNA-related bioinformatics domain is concerned, discriminative models, like DNABert, exist. Yet, the generative side of the coin is mainly unexplored to the best of our knowledge. Consequently, we focused on developing an autoregressive generative language model like GPT-3 for DNA sequences. Because working with whole DNA sequences is challenging without substantial computational resources, we decided to carry out our study on a smaller scale, focusing on nucleotide sequences of human genes, unique parts in DNA with specific functionalities, instead of the whole DNA. This decision did not change the problem structure a lot due to the fact that both DNA and genes can be seen as 1D sequences consisting of four different nucleotides without losing much information and making too much simplification. First of all, we systematically examined an almost entirely unexplored problem and observed that RNNs performed the best while simple techniques like N-grams were also promising. Another beneficial point was learning how to work with generative models on languages we do not understand, unlike natural language. How essential using real-life tasks beyond the classical metrics such as perplexity is observed. Furthermore, checking whether the data-hungry nature of these models can be changed through selecting a language with minimal vocabulary size, four owing to four different types of nucleotides, is examined. The reason for reviewing this was that choosing such a language might make the problem easier. However, what we observed in this study was it did not provide that much of a change in the amount of data needed.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge