"Information": models, code, and papers
A Generalized Bandsplit Neural Network for Cinematic Audio Source Separation
Sep 07, 2023Cinematic audio source separation is a relatively new subtask of audio source separation, with the aim of extracting the dialogue stem, the music stem, and the effects stem from their mixture. In this work, we developed a model generalizing the Bandsplit RNN for any complete or overcomplete partitions of the frequency axis. Psycho-acoustically motivated frequency scales were used to inform the band definitions which are now defined with redundancy for more reliable feature extraction. A loss function motivated by the signal-to-noise ratio and the sparsity-promoting property of the 1-norm was proposed. We additionally exploit the information-sharing property of a common-encoder setup to reduce computational complexity during both training and inference, improve separation performance for hard-to-generalize classes of sounds, and allow flexibility during inference time with easily detachable decoders. Our best model sets the state of the art on the Divide and Remaster dataset with performance above the ideal ratio mask for the dialogue stem.
Feature Enhancer Segmentation Network (FES-Net) for Vessel Segmentation
Sep 07, 2023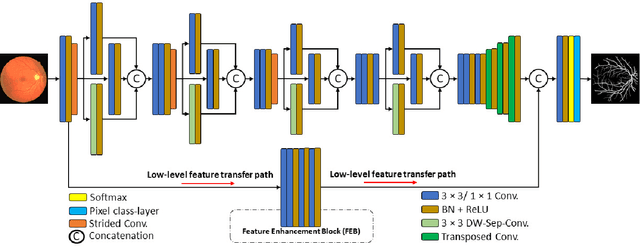
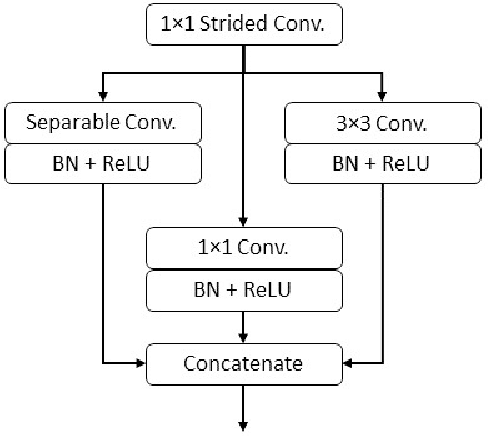
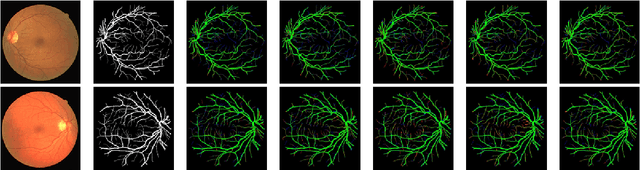
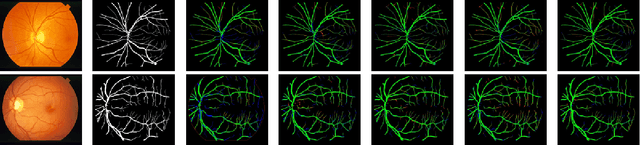
Diseases such as diabetic retinopathy and age-related macular degeneration pose a significant risk to vision, highlighting the importance of precise segmentation of retinal vessels for the tracking and diagnosis of progression. However, existing vessel segmentation methods that heavily rely on encoder-decoder structures struggle to capture contextual information about retinal vessel configurations, leading to challenges in reconciling semantic disparities between encoder and decoder features. To address this, we propose a novel feature enhancement segmentation network (FES-Net) that achieves accurate pixel-wise segmentation without requiring additional image enhancement steps. FES-Net directly processes the input image and utilizes four prompt convolutional blocks (PCBs) during downsampling, complemented by a shallow upsampling approach to generate a binary mask for each class. We evaluate the performance of FES-Net on four publicly available state-of-the-art datasets: DRIVE, STARE, CHASE, and HRF. The evaluation results clearly demonstrate the superior performance of FES-Net compared to other competitive approaches documented in the existing literature.
BluNF: Blueprint Neural Field
Sep 07, 2023Neural Radiance Fields (NeRFs) have revolutionized scene novel view synthesis, offering visually realistic, precise, and robust implicit reconstructions. While recent approaches enable NeRF editing, such as object removal, 3D shape modification, or material property manipulation, the manual annotation prior to such edits makes the process tedious. Additionally, traditional 2D interaction tools lack an accurate sense of 3D space, preventing precise manipulation and editing of scenes. In this paper, we introduce a novel approach, called Blueprint Neural Field (BluNF), to address these editing issues. BluNF provides a robust and user-friendly 2D blueprint, enabling intuitive scene editing. By leveraging implicit neural representation, BluNF constructs a blueprint of a scene using prior semantic and depth information. The generated blueprint allows effortless editing and manipulation of NeRF representations. We demonstrate BluNF's editability through an intuitive click-and-change mechanism, enabling 3D manipulations, such as masking, appearance modification, and object removal. Our approach significantly contributes to visual content creation, paving the way for further research in this area.
Aggregating Correlated Estimations with (Almost) no Training
Sep 05, 2023Many decision problems cannot be solved exactly and use several estimation algorithms that assign scores to the different available options. The estimation errors can have various correlations, from low (e.g. between two very different approaches) to high (e.g. when using a given algorithm with different hyperparameters). Most aggregation rules would suffer from this diversity of correlations. In this article, we propose different aggregation rules that take correlations into account, and we compare them to naive rules in various experiments based on synthetic data. Our results show that when sufficient information is known about the correlations between errors, a maximum likelihood aggregation should be preferred. Otherwise, typically with limited training data, we recommend a method that we call Embedded Voting (EV).
SdCT-GAN: Reconstructing CT from Biplanar X-Rays with Self-driven Generative Adversarial Networks
Sep 10, 2023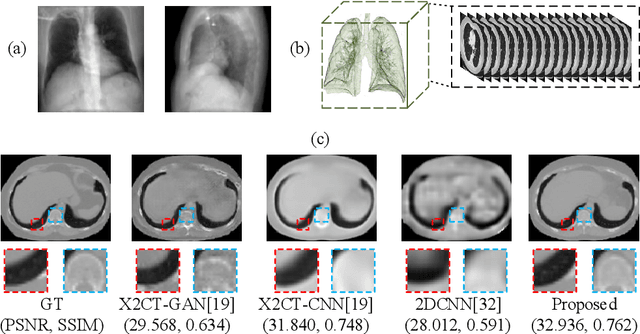
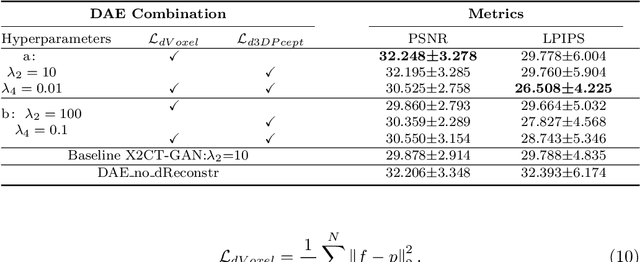
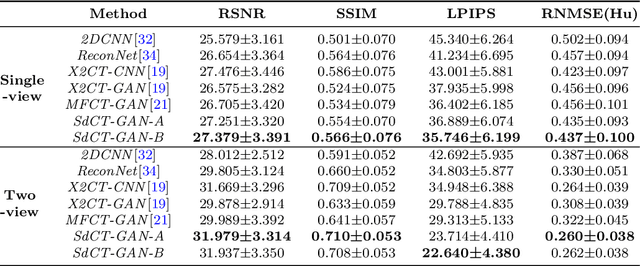
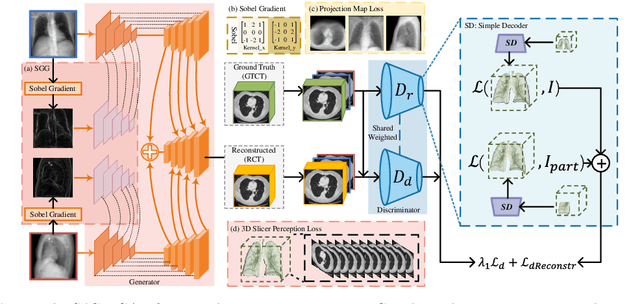

Computed Tomography (CT) is a medical imaging modality that can generate more informative 3D images than 2D X-rays. However, this advantage comes at the expense of more radiation exposure, higher costs, and longer acquisition time. Hence, the reconstruction of 3D CT images using a limited number of 2D X-rays has gained significant importance as an economical alternative. Nevertheless, existing methods primarily prioritize minimizing pixel/voxel-level intensity discrepancies, often neglecting the preservation of textural details in the synthesized images. This oversight directly impacts the quality of the reconstructed images and thus affects the clinical diagnosis. To address the deficits, this paper presents a new self-driven generative adversarial network model (SdCT-GAN), which is motivated to pay more attention to image details by introducing a novel auto-encoder structure in the discriminator. In addition, a Sobel Gradient Guider (SGG) idea is applied throughout the model, where the edge information from the 2D X-ray image at the input can be integrated. Moreover, LPIPS (Learned Perceptual Image Patch Similarity) evaluation metric is adopted that can quantitatively evaluate the fine contours and textures of reconstructed images better than the existing ones. Finally, the qualitative and quantitative results of the empirical studies justify the power of the proposed model compared to mainstream state-of-the-art baselines.
Information criteria for structured parameter selection in high dimensional tree and graph models
Jun 24, 2023Parameter selection in high-dimensional models is typically finetuned in a way that keeps the (relative) number of false positives under control. This is because otherwise the few true positives may be dominated by the many possible false positives. This happens, for instance, when the selection follows from a naive optimisation of an information criterion, such as AIC or Mallows's Cp. It can be argued that the overestimation of the selection comes from the optimisation process itself changing the statistics of the selected variables, in a way that the information criterion no longer reflects the true divergence between the selection and the data generating process. In lasso, the overestimation can also be linked to the shrinkage estimator, which makes the selection too tolerant of false positive selections. For these reasons, this paper works on refined information criteria, carefully balancing false positives and false negatives, for use with estimators without shrinkage. In particular, the paper develops corrected Mallows's Cp criteria for structured selection in trees and graphical models.
Geometry-aware Line Graph Transformer Pre-training for Molecular Property Prediction
Sep 01, 2023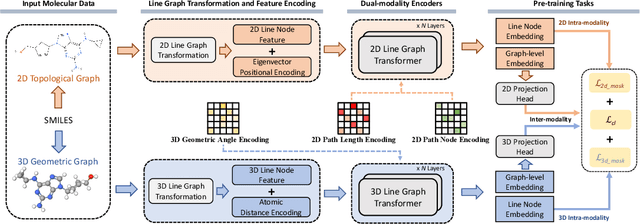
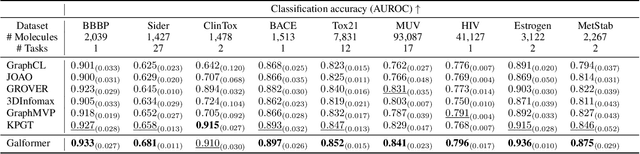
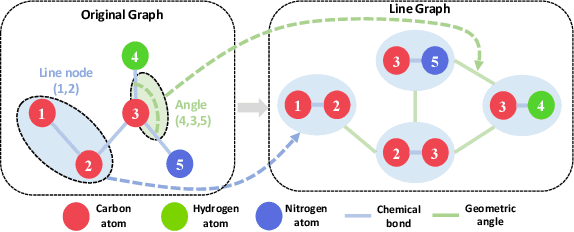
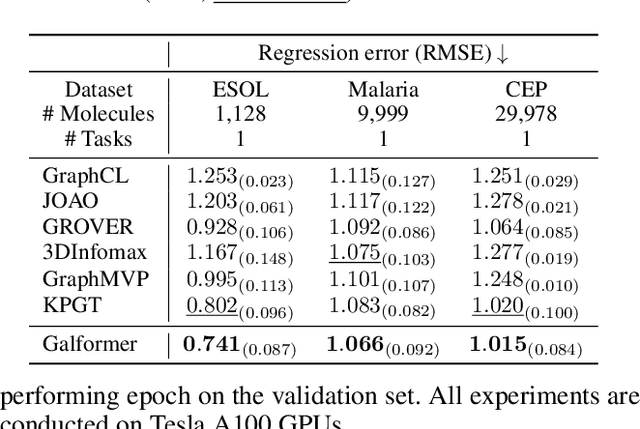
Molecular property prediction with deep learning has gained much attention over the past years. Owing to the scarcity of labeled molecules, there has been growing interest in self-supervised learning methods that learn generalizable molecular representations from unlabeled data. Molecules are typically treated as 2D topological graphs in modeling, but it has been discovered that their 3D geometry is of great importance in determining molecular functionalities. In this paper, we propose the Geometry-aware line graph transformer (Galformer) pre-training, a novel self-supervised learning framework that aims to enhance molecular representation learning with 2D and 3D modalities. Specifically, we first design a dual-modality line graph transformer backbone to encode the topological and geometric information of a molecule. The designed backbone incorporates effective structural encodings to capture graph structures from both modalities. Then we devise two complementary pre-training tasks at the inter and intra-modality levels. These tasks provide properly supervised information and extract discriminative 2D and 3D knowledge from unlabeled molecules. Finally, we evaluate Galformer against six state-of-the-art baselines on twelve property prediction benchmarks via downstream fine-tuning. Experimental results show that Galformer consistently outperforms all baselines on both classification and regression tasks, demonstrating its effectiveness.
A Circuit Complexity Formulation of Algorithmic Information Theory
Jun 25, 2023Inspired by Solomonoffs theory of inductive inference, we propose a prior based on circuit complexity. There are several advantages to this approach. First, it relies on a complexity measure that does not depend on the choice of UTM. There is one universal definition for Boolean circuits involving an universal operation such as nand with simple conversions to alternative definitions such as and, or, and not. Second, there is no analogue of the halting problem. The output value of a circuit can be calculated recursively by computer in time proportional to the number of gates, while a short program may run for a very long time. Our prior assumes that a Boolean function, or equivalently, Boolean string of fixed length, is generated by some Bayesian mixture of circuits. This model is appropriate for learning Boolean functions from partial information, a problem often encountered within machine learning as "binary classification." We argue that an inductive bias towards simple explanations as measured by circuit complexity is appropriate for this problem.
Data-Driven Information Extraction and Enrichment of Molecular Profiling Data for Cancer Cell Lines
Jul 03, 2023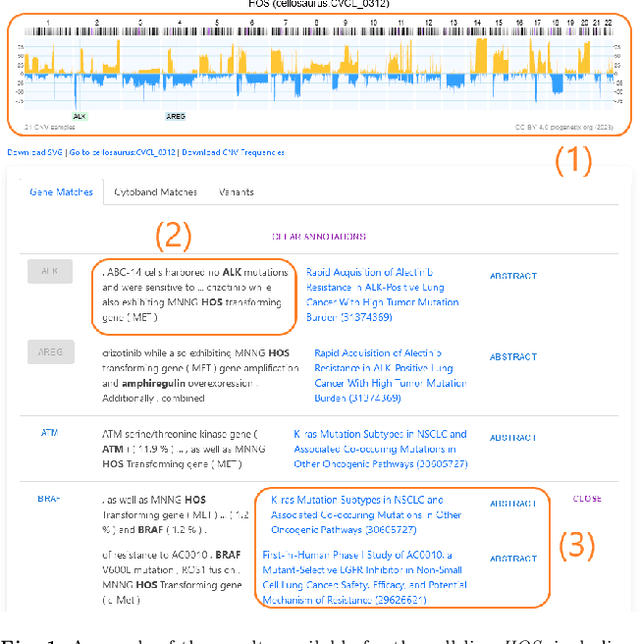
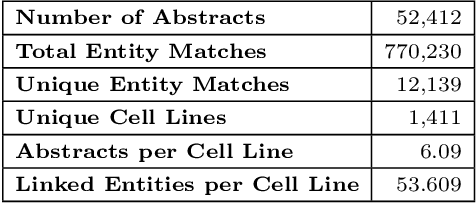

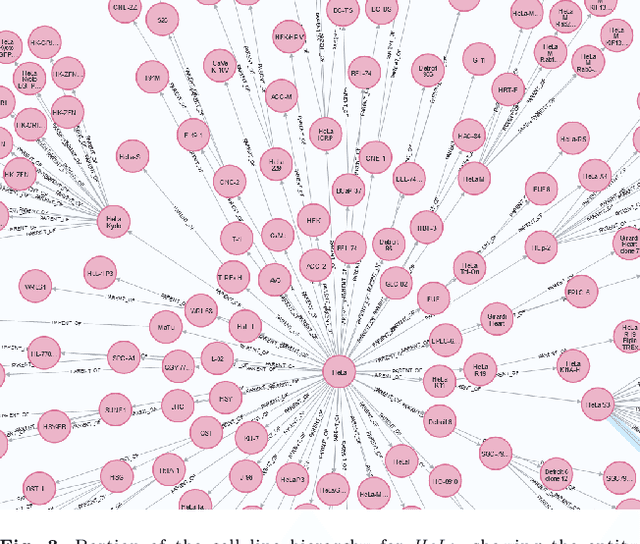
With the proliferation of research means and computational methodologies, published biomedical literature is growing exponentially in numbers and volume. As a consequence, in the fields of biological, medical and clinical research, domain experts have to sift through massive amounts of scientific text to find relevant information. However, this process is extremely tedious and slow to be performed by humans. Hence, novel computational information extraction and correlation mechanisms are required to boost meaningful knowledge extraction. In this work, we present the design, implementation and application of a novel data extraction and exploration system. This system extracts deep semantic relations between textual entities from scientific literature to enrich existing structured clinical data in the domain of cancer cell lines. We introduce a new public data exploration portal, which enables automatic linking of genomic copy number variants plots with ranked, related entities such as affected genes. Each relation is accompanied by literature-derived evidences, allowing for deep, yet rapid, literature search, using existing structured data as a springboard. Our system is publicly available on the web at https://cancercelllines.org
Knowledge-Enhanced Hierarchical Information Correlation Learning for Multi-Modal Rumor Detection
Jun 28, 2023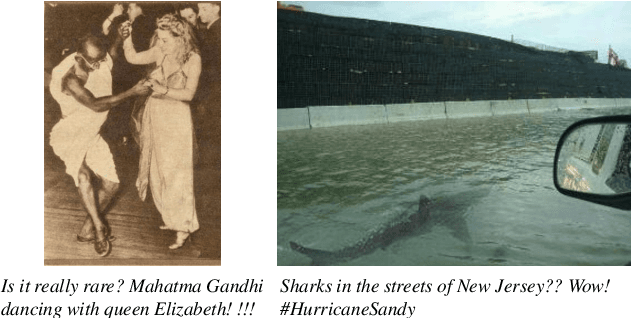
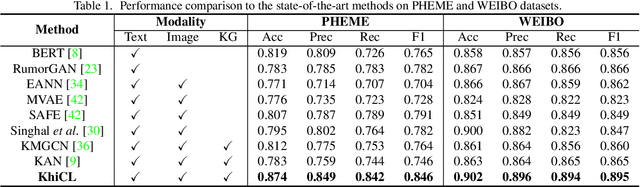
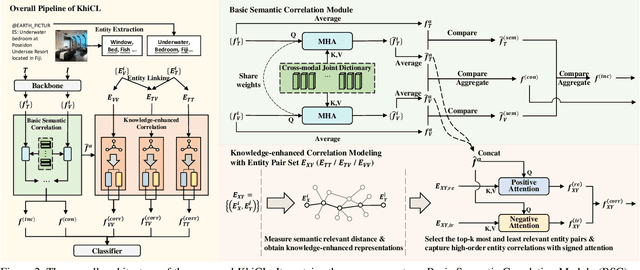
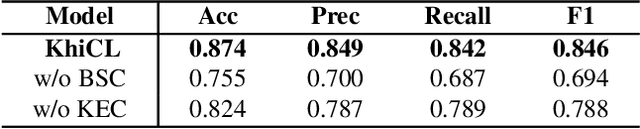
The explosive growth of rumors with text and images on social media platforms has drawn great attention. Existing studies have made significant contributions to cross-modal information interaction and fusion, but they fail to fully explore hierarchical and complex semantic correlation across different modality content, severely limiting their performance on detecting multi-modal rumor. In this work, we propose a novel knowledge-enhanced hierarchical information correlation learning approach (KhiCL) for multi-modal rumor detection by jointly modeling the basic semantic correlation and high-order knowledge-enhanced entity correlation. Specifically, KhiCL exploits cross-modal joint dictionary to transfer the heterogeneous unimodality features into the common feature space and captures the basic cross-modal semantic consistency and inconsistency by a cross-modal fusion layer. Moreover, considering the description of multi-modal content is narrated around entities, KhiCL extracts visual and textual entities from images and text, and designs a knowledge relevance reasoning strategy to find the shortest semantic relevant path between each pair of entities in external knowledge graph, and absorbs all complementary contextual knowledge of other connected entities in this path for learning knowledge-enhanced entity representations. Furthermore, KhiCL utilizes a signed attention mechanism to model the knowledge-enhanced entity consistency and inconsistency of intra-modality and inter-modality entity pairs by measuring their corresponding semantic relevant distance. Extensive experiments have demonstrated the effectiveness of the proposed method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge