"Information": models, code, and papers
Recognizable Information Bottleneck
Apr 28, 2023
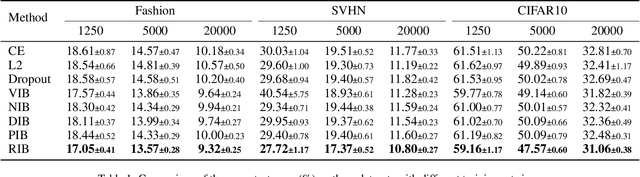
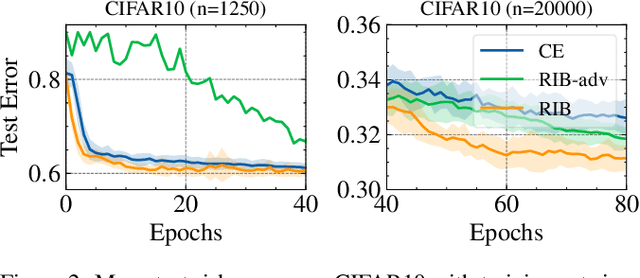

Information Bottlenecks (IBs) learn representations that generalize to unseen data by information compression. However, existing IBs are practically unable to guarantee generalization in real-world scenarios due to the vacuous generalization bound. The recent PAC-Bayes IB uses information complexity instead of information compression to establish a connection with the mutual information generalization bound. However, it requires the computation of expensive second-order curvature, which hinders its practical application. In this paper, we establish the connection between the recognizability of representations and the recent functional conditional mutual information (f-CMI) generalization bound, which is significantly easier to estimate. On this basis we propose a Recognizable Information Bottleneck (RIB) which regularizes the recognizability of representations through a recognizability critic optimized by density ratio matching under the Bregman divergence. Extensive experiments on several commonly used datasets demonstrate the effectiveness of the proposed method in regularizing the model and estimating the generalization gap.
A Circuit Complexity Formulation of Algorithmic Information Theory
Jun 25, 2023Inspired by Solomonoffs theory of inductive inference, we propose a prior based on circuit complexity. There are several advantages to this approach. First, it relies on a complexity measure that does not depend on the choice of UTM. There is one universal definition for Boolean circuits involving an universal operation such as nand with simple conversions to alternative definitions such as and, or, and not. Second, there is no analogue of the halting problem. The output value of a circuit can be calculated recursively by computer in time proportional to the number of gates, while a short program may run for a very long time. Our prior assumes that a Boolean function, or equivalently, Boolean string of fixed length, is generated by some Bayesian mixture of circuits. This model is appropriate for learning Boolean functions from partial information, a problem often encountered within machine learning as "binary classification." We argue that an inductive bias towards simple explanations as measured by circuit complexity is appropriate for this problem.
Grandma Karl is 27 years old -- research agenda for pseudonymization of research data
Aug 30, 2023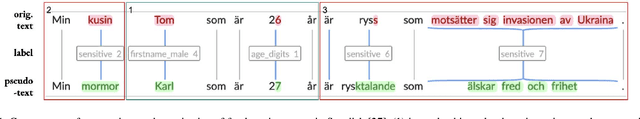
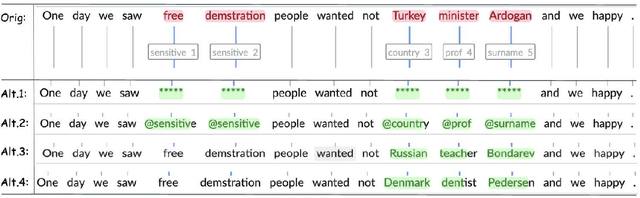
Accessibility of research data is critical for advances in many research fields, but textual data often cannot be shared due to the personal and sensitive information which it contains, e.g names or political opinions. General Data Protection Regulation (GDPR) suggests pseudonymization as a solution to secure open access to research data, but we need to learn more about pseudonymization as an approach before adopting it for manipulation of research data. This paper outlines a research agenda within pseudonymization, namely need of studies into the effects of pseudonymization on unstructured data in relation to e.g. readability and language assessment, as well as the effectiveness of pseudonymization as a way of protecting writer identity, while also exploring different ways of developing context-sensitive algorithms for detection, labelling and replacement of personal information in unstructured data. The recently granted project on pseudonymization Grandma Karl is 27 years old addresses exactly those challenges.
Enhancing Hyperedge Prediction with Context-Aware Self-Supervised Learning
Sep 11, 2023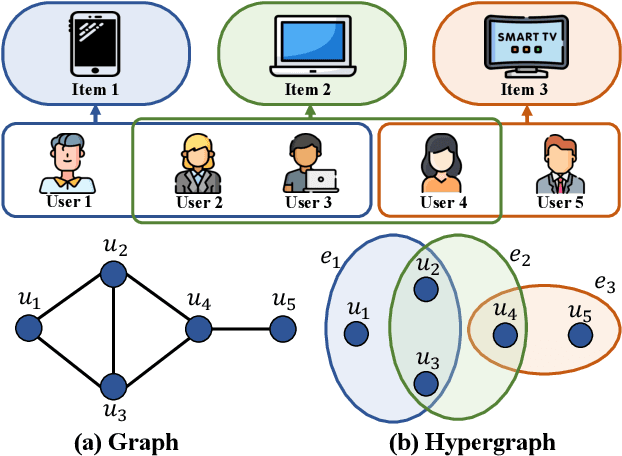
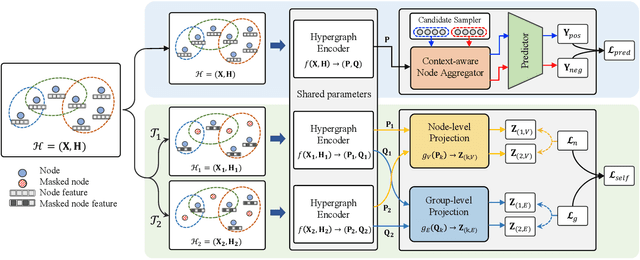
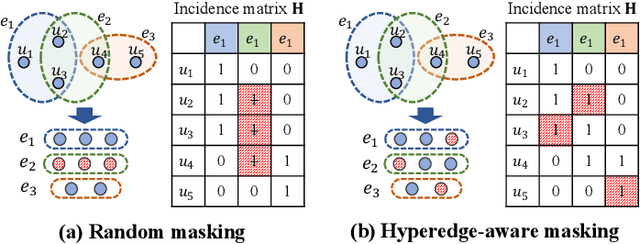
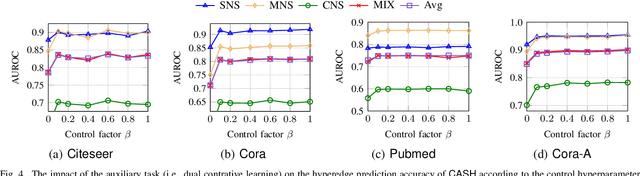
Hypergraphs can naturally model group-wise relations (e.g., a group of users who co-purchase an item) as hyperedges. Hyperedge prediction is to predict future or unobserved hyperedges, which is a fundamental task in many real-world applications (e.g., group recommendation). Despite the recent breakthrough of hyperedge prediction methods, the following challenges have been rarely studied: (C1) How to aggregate the nodes in each hyperedge candidate for accurate hyperedge prediction? and (C2) How to mitigate the inherent data sparsity problem in hyperedge prediction? To tackle both challenges together, in this paper, we propose a novel hyperedge prediction framework (CASH) that employs (1) context-aware node aggregation to precisely capture complex relations among nodes in each hyperedge for (C1) and (2) self-supervised contrastive learning in the context of hyperedge prediction to enhance hypergraph representations for (C2). Furthermore, as for (C2), we propose a hyperedge-aware augmentation method to fully exploit the latent semantics behind the original hypergraph and consider both node-level and group-level contrasts (i.e., dual contrasts) for better node and hyperedge representations. Extensive experiments on six real-world hypergraphs reveal that CASH consistently outperforms all competing methods in terms of the accuracy in hyperedge prediction and each of the proposed strategies is effective in improving the model accuracy of CASH. For the detailed information of CASH, we provide the code and datasets at: https://github.com/yy-ko/cash.
SayNav: Grounding Large Language Models for Dynamic Planning to Navigation in New Environments
Sep 11, 2023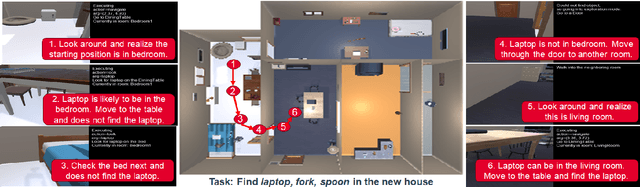
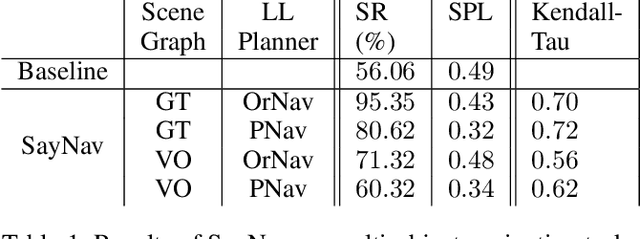
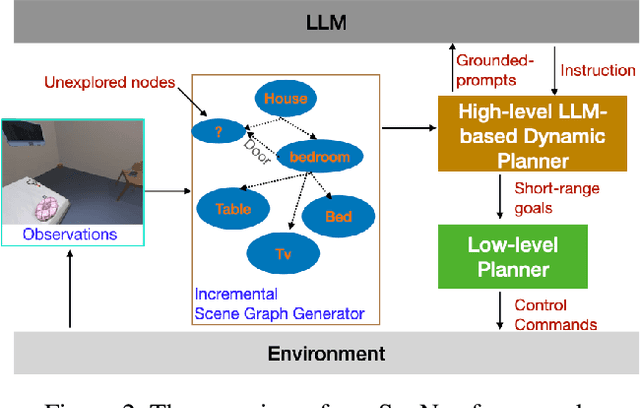
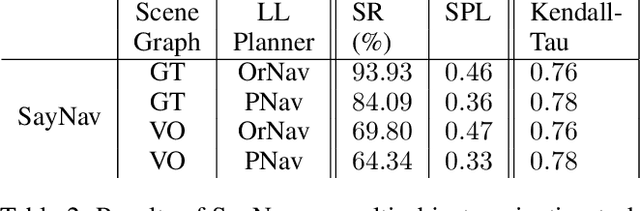
Semantic reasoning and dynamic planning capabilities are crucial for an autonomous agent to perform complex navigation tasks in unknown environments. It requires a large amount of common-sense knowledge, that humans possess, to succeed in these tasks. We present SayNav, a new approach that leverages human knowledge from Large Language Models (LLMs) for efficient generalization to complex navigation tasks in unknown large-scale environments. SayNav uses a novel grounding mechanism, that incrementally builds a 3D scene graph of the explored environment as inputs to LLMs, for generating feasible and contextually appropriate high-level plans for navigation. The LLM-generated plan is then executed by a pre-trained low-level planner, that treats each planned step as a short-distance point-goal navigation sub-task. SayNav dynamically generates step-by-step instructions during navigation and continuously refines future steps based on newly perceived information. We evaluate SayNav on a new multi-object navigation task, that requires the agent to utilize a massive amount of human knowledge to efficiently search multiple different objects in an unknown environment. SayNav outperforms an oracle based Point-nav baseline, achieving a success rate of 95.35% (vs 56.06% for the baseline), under the ideal settings on this task, highlighting its ability to generate dynamic plans for successfully locating objects in large-scale new environments. In addition, SayNav also enables efficient generalization from simulation to real environments.
Towards Content-based Pixel Retrieval in Revisited Oxford and Paris
Sep 11, 2023
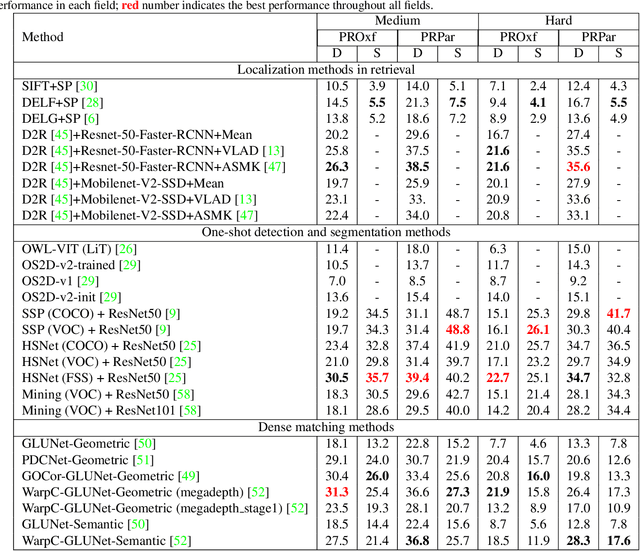
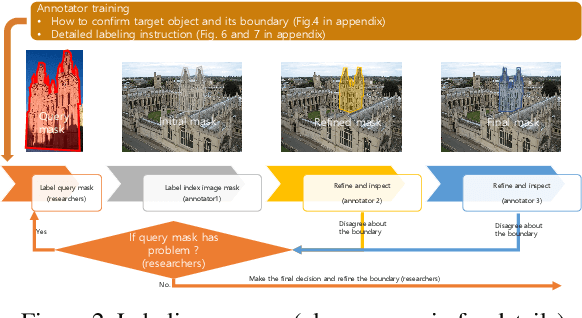
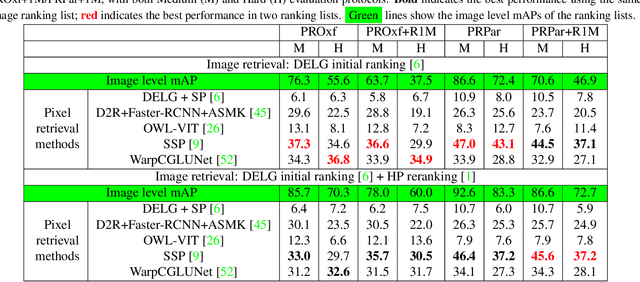
This paper introduces the first two pixel retrieval benchmarks. Pixel retrieval is segmented instance retrieval. Like semantic segmentation extends classification to the pixel level, pixel retrieval is an extension of image retrieval and offers information about which pixels are related to the query object. In addition to retrieving images for the given query, it helps users quickly identify the query object in true positive images and exclude false positive images by denoting the correlated pixels. Our user study results show pixel-level annotation can significantly improve the user experience. Compared with semantic and instance segmentation, pixel retrieval requires a fine-grained recognition capability for variable-granularity targets. To this end, we propose pixel retrieval benchmarks named PROxford and PRParis, which are based on the widely used image retrieval datasets, ROxford and RParis. Three professional annotators label 5,942 images with two rounds of double-checking and refinement. Furthermore, we conduct extensive experiments and analysis on the SOTA methods in image search, image matching, detection, segmentation, and dense matching using our pixel retrieval benchmarks. Results show that the pixel retrieval task is challenging to these approaches and distinctive from existing problems, suggesting that further research can advance the content-based pixel-retrieval and thus user search experience. The datasets can be downloaded from \href{https://github.com/anguoyuan/Pixel_retrieval-Segmented_instance_retrieval}{this link}.
Large Process Models: Business Process Management in the Age of Generative AI
Sep 11, 2023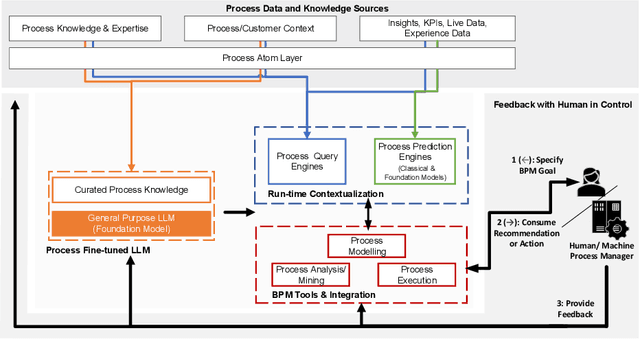
The continued success of Large Language Models (LLMs) and other generative artificial intelligence approaches highlights the advantages that large information corpora can have over rigidly defined symbolic models, but also serves as a proof-point of the challenges that purely statistics-based approaches have in terms of safety and trustworthiness. As a framework for contextualizing the potential, as well as the limitations of LLMs and other foundation model-based technologies, we propose the concept of a Large Process Model (LPM) that combines the correlation power of LLMs with the analytical precision and reliability of knowledge-based systems and automated reasoning approaches. LPMs are envisioned to directly utilize the wealth of process management experience that experts have accumulated, as well as process performance data of organizations with diverse characteristics, e.g., regarding size, region, or industry. In this vision, the proposed LPM would allow organizations to receive context-specific (tailored) process and other business models, analytical deep-dives, and improvement recommendations. As such, they would allow to substantially decrease the time and effort required for business transformation, while also allowing for deeper, more impactful, and more actionable insights than previously possible. We argue that implementing an LPM is feasible, but also highlight limitations and research challenges that need to be solved to implement particular aspects of the LPM vision.
ECG-based estimation of respiratory modulation of AV nodal conduction during atrial fibrillation
Sep 11, 2023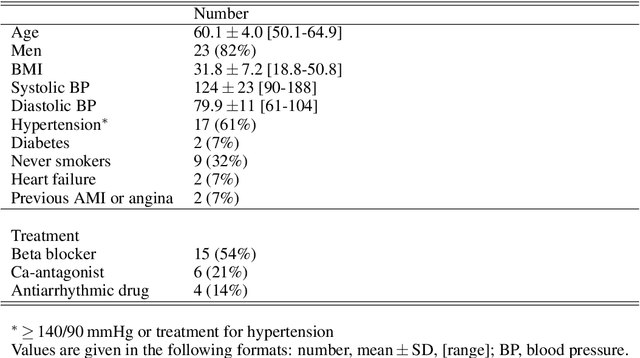
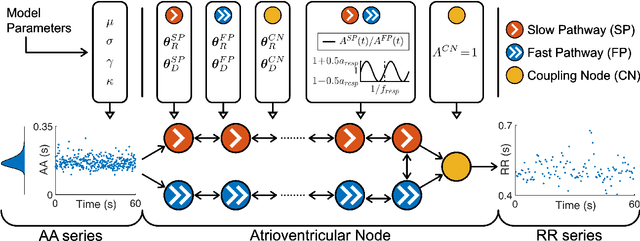
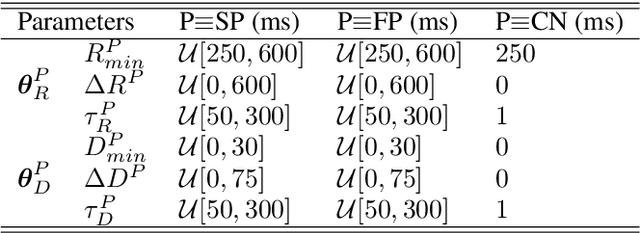
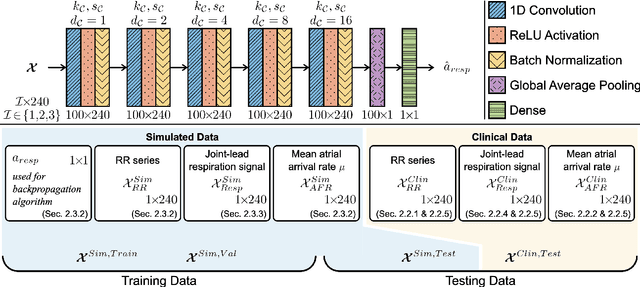
Information about autonomic nervous system (ANS) activity may be valuable for personalized atrial fibrillation (AF) treatment but is not easily accessible from the ECG. In this study, we propose a new approach for ECG-based assessment of respiratory modulation in AV nodal refractory period and conduction delay. A 1-dimensional convolutional neural network (1D-CNN) was trained to estimate respiratory modulation of AV nodal conduction properties from 1-minute segments of RR series, respiration signals, and atrial fibrillatory rates (AFR) using synthetic data that replicates clinical ECG-derived data. The synthetic data were generated using a network model of the AV node and 4 million unique model parameter sets. The 1D-CNN was then used to analyze respiratory modulation in clinical deep breathing test data of 28 patients in AF, where a ECG-derived respiration signal was extracted using a novel approach based on periodic component analysis. We demonstrated using synthetic data that the 1D-CNN can predict the respiratory modulation from RR series alone ($\rho$ = 0.805) and that the addition of either respiration signal ($\rho$ = 0.830), AFR ($\rho$ = 0.837), or both ($\rho$ = 0.855) improves the prediction. Results from analysis of clinical ECG data of 20 patients with sufficient signal quality suggest that respiratory modulation decreased in response to deep breathing for five patients, increased for five patients, and remained similar for ten patients, indicating a large inter-patient variability.
Improving Neural Ranking Models with Traditional IR Methods
Aug 29, 2023Neural ranking methods based on large transformer models have recently gained significant attention in the information retrieval community, and have been adopted by major commercial solutions. Nevertheless, they are computationally expensive to create, and require a great deal of labeled data for specialized corpora. In this paper, we explore a low resource alternative which is a bag-of-embedding model for document retrieval and find that it is competitive with large transformer models fine tuned on information retrieval tasks. Our results show that a simple combination of TF-IDF, a traditional keyword matching method, with a shallow embedding model provides a low cost path to compete well with the performance of complex neural ranking models on 3 datasets. Furthermore, adding TF-IDF measures improves the performance of large-scale fine tuned models on these tasks.
Data-Driven Information Extraction and Enrichment of Molecular Profiling Data for Cancer Cell Lines
Jul 03, 2023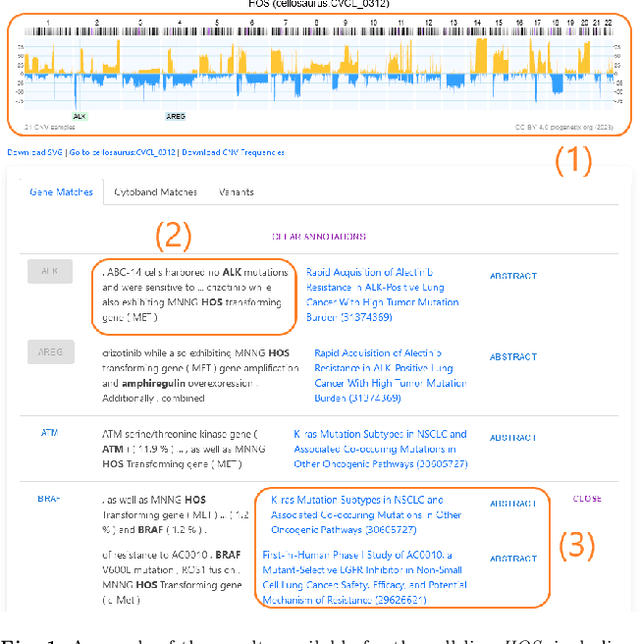
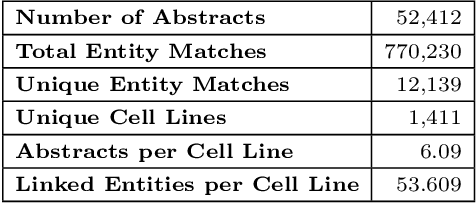

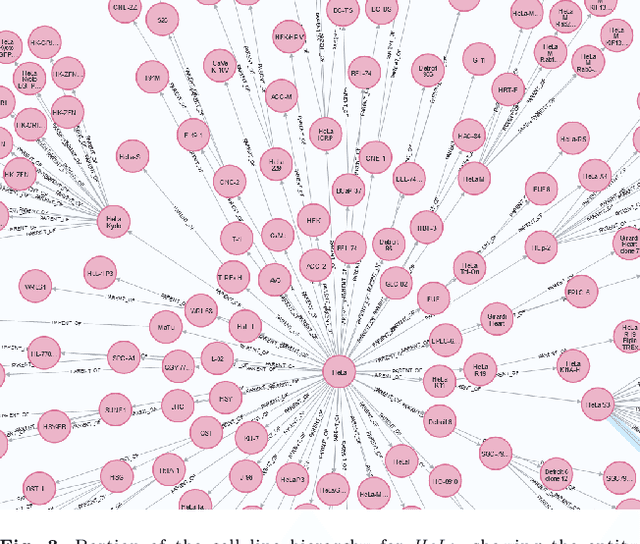
With the proliferation of research means and computational methodologies, published biomedical literature is growing exponentially in numbers and volume. As a consequence, in the fields of biological, medical and clinical research, domain experts have to sift through massive amounts of scientific text to find relevant information. However, this process is extremely tedious and slow to be performed by humans. Hence, novel computational information extraction and correlation mechanisms are required to boost meaningful knowledge extraction. In this work, we present the design, implementation and application of a novel data extraction and exploration system. This system extracts deep semantic relations between textual entities from scientific literature to enrich existing structured clinical data in the domain of cancer cell lines. We introduce a new public data exploration portal, which enables automatic linking of genomic copy number variants plots with ranked, related entities such as affected genes. Each relation is accompanied by literature-derived evidences, allowing for deep, yet rapid, literature search, using existing structured data as a springboard. Our system is publicly available on the web at https://cancercelllines.org
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge