"Information Extraction": models, code, and papers
Enhancing the Performance of Aspect-Based Sentiment Analysis Systems
Apr 04, 2024Aspect-based sentiment analysis aims to predict sentiment polarity with fine granularity. While Graph Convolutional Networks (GCNs) are widely utilized for sentimental feature extraction, their naive application for syntactic feature extraction can compromise information preservation. This study introduces an innovative edge-enhanced GCN, named SentiSys, to navigate the syntactic graph while preserving intact feature information, leading to enhanced performance. Specifically,we first integrate a bidirectional long short-term memory (Bi-LSTM) network and a self-attention-based transformer. This combination facilitates effective text encoding, preventing the loss of information and predicting long dependency text. A bidirectional GCN (Bi-GCN) with message passing is then employed to encode relationships between entities. Additionally, unnecessary information is filtered out using an aspect-specific masking technique. To validate the effectiveness of our proposed model, we conduct extensive evaluation experiments and ablation studies on four benchmark datasets. The results consistently demonstrate improved performance in aspect-based sentiment analysis when employing SentiSys. This approach successfully addresses the challenges associated with syntactic feature extraction, highlighting its potential for advancing sentiment analysis methodologies.
Automating the Information Extraction from Semi-Structured Interview Transcripts
Mar 07, 2024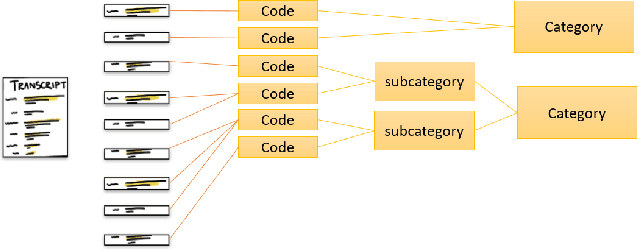
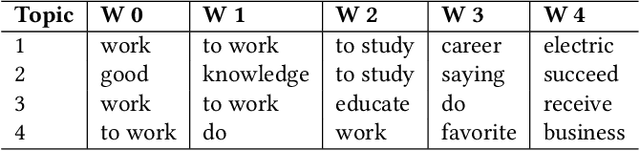
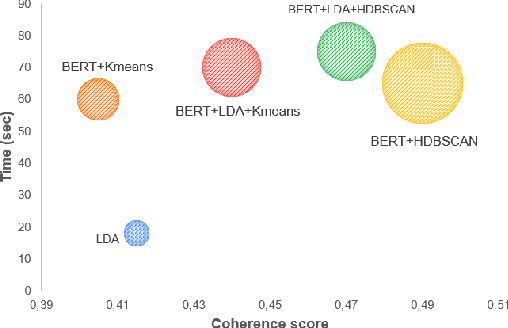
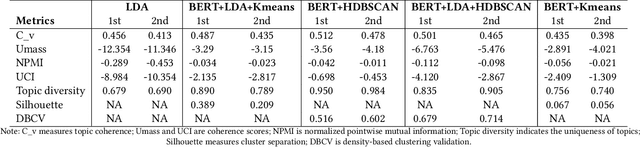
This paper explores the development and application of an automated system designed to extract information from semi-structured interview transcripts. Given the labor-intensive nature of traditional qualitative analysis methods, such as coding, there exists a significant demand for tools that can facilitate the analysis process. Our research investigates various topic modeling techniques and concludes that the best model for analyzing interview texts is a combination of BERT embeddings and HDBSCAN clustering. We present a user-friendly software prototype that enables researchers, including those without programming skills, to efficiently process and visualize the thematic structure of interview data. This tool not only facilitates the initial stages of qualitative analysis but also offers insights into the interconnectedness of topics revealed, thereby enhancing the depth of qualitative analysis.
LayoutLLM: Large Language Model Instruction Tuning for Visually Rich Document Understanding
Mar 21, 2024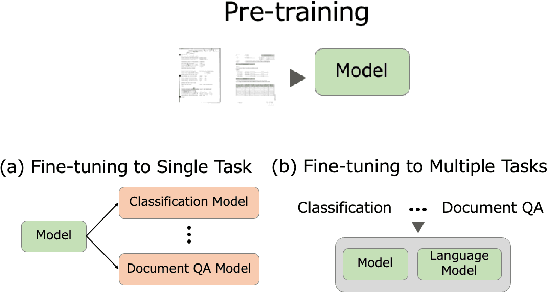

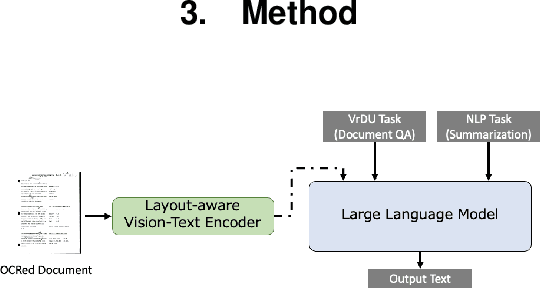
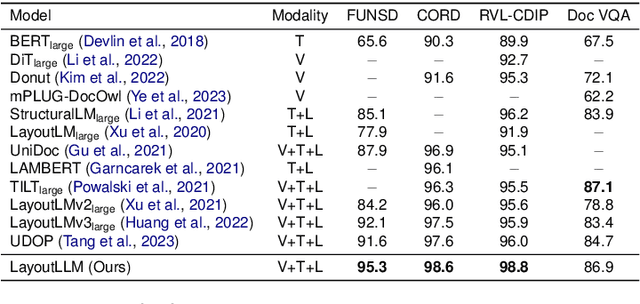
This paper proposes LayoutLLM, a more flexible document analysis method for understanding imaged documents. Visually Rich Document Understanding tasks, such as document image classification and information extraction, have gained significant attention due to their importance. Existing methods have been developed to enhance document comprehension by incorporating pre-training awareness of images, text, and layout structure. However, these methods require fine-tuning for each task and dataset, and the models are expensive to train and operate. To overcome this limitation, we propose a new LayoutLLM that integrates these with large-scale language models (LLMs). By leveraging the strengths of existing research in document image understanding and LLMs' superior language understanding capabilities, the proposed model, fine-tuned with multimodal instruction datasets, performs an understanding of document images in a single model. Our experiments demonstrate improvement over the baseline model in various document analysis tasks.
Combining Local and Global Perception for Autonomous Navigation on Nano-UAVs
Mar 18, 2024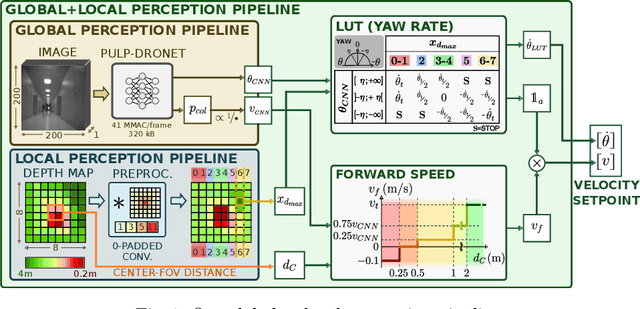
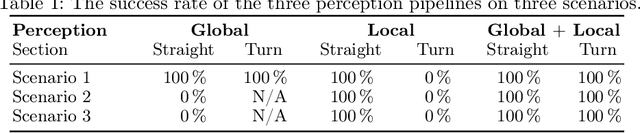

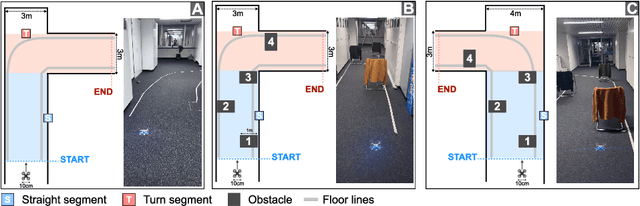
A critical challenge in deploying unmanned aerial vehicles (UAVs) for autonomous tasks is their ability to navigate in an unknown environment. This paper introduces a novel vision-depth fusion approach for autonomous navigation on nano-UAVs. We combine the visual-based PULP-Dronet convolutional neural network for semantic information extraction, i.e., serving as the global perception, with 8x8px depth maps for close-proximity maneuvers, i.e., the local perception. When tested in-field, our integration strategy highlights the complementary strengths of both visual and depth sensory information. We achieve a 100% success rate over 15 flights in a complex navigation scenario, encompassing straight pathways, static obstacle avoidance, and 90{\deg} turns.
Classifying Cancer Stage with Open-Source Clinical Large Language Models
Apr 02, 2024Cancer stage classification is important for making treatment and care management plans for oncology patients. Information on staging is often included in unstructured form in clinical, pathology, radiology and other free-text reports in the electronic health record system, requiring extensive work to parse and obtain. To facilitate the extraction of this information, previous NLP approaches rely on labeled training datasets, which are labor-intensive to prepare. In this study, we demonstrate that without any labeled training data, open-source clinical large language models (LLMs) can extract pathologic tumor-node-metastasis (pTNM) staging information from real-world pathology reports. Our experiments compare LLMs and a BERT-based model fine-tuned using the labeled data. Our findings suggest that while LLMs still exhibit subpar performance in Tumor (T) classification, with the appropriate adoption of prompting strategies, they can achieve comparable performance on Metastasis (M) classification and improved performance on Node (N) classification.
MarsSeg: Mars Surface Semantic Segmentation with Multi-level Extractor and Connector
Apr 05, 2024The segmentation and interpretation of the Martian surface play a pivotal role in Mars exploration, providing essential data for the trajectory planning and obstacle avoidance of rovers. However, the complex topography, similar surface features, and the lack of extensive annotated data pose significant challenges to the high-precision semantic segmentation of the Martian surface. To address these challenges, we propose a novel encoder-decoder based Mars segmentation network, termed MarsSeg. Specifically, we employ an encoder-decoder structure with a minimized number of down-sampling layers to preserve local details. To facilitate a high-level semantic understanding across the shadow multi-level feature maps, we introduce a feature enhancement connection layer situated between the encoder and decoder. This layer incorporates Mini Atrous Spatial Pyramid Pooling (Mini-ASPP), Polarized Self-Attention (PSA), and Strip Pyramid Pooling Module (SPPM). The Mini-ASPP and PSA are specifically designed for shadow feature enhancement, thereby enabling the expression of local details and small objects. Conversely, the SPPM is employed for deep feature enhancement, facilitating the extraction of high-level semantic category-related information. Experimental results derived from the Mars-Seg and AI4Mars datasets substantiate that the proposed MarsSeg outperforms other state-of-the-art methods in segmentation performance, validating the efficacy of each proposed component.
NLP for Knowledge Discovery and Information Extraction from Energetics Corpora
Feb 10, 2024We present a demonstration of the utility of NLP for aiding research into energetic materials and associated systems. The NLP method enables machine understanding of textual data, offering an automated route to knowledge discovery and information extraction from energetics text. We apply three established unsupervised NLP models: Latent Dirichlet Allocation, Word2Vec, and the Transformer to a large curated dataset of energetics-related scientific articles. We demonstrate that each NLP algorithm is capable of identifying energetic topics and concepts, generating a language model which aligns with Subject Matter Expert knowledge. Furthermore, we present a document classification pipeline for energetics text. Our classification pipeline achieves 59-76\% accuracy depending on the NLP model used, with the highest performing Transformer model rivaling inter-annotator agreement metrics. The NLP approaches studied in this work can identify concepts germane to energetics and therefore hold promise as a tool for accelerating energetics research efforts and energetics material development.
AutoRE: Document-Level Relation Extraction with Large Language Models
Mar 21, 2024Large Language Models (LLMs) have demonstrated exceptional abilities in comprehending and generating text, motivating numerous researchers to utilize them for Information Extraction (IE) purposes, including Relation Extraction (RE). Nonetheless, most existing methods are predominantly designed for Sentence-level Relation Extraction (SentRE) tasks, which typically encompass a restricted set of relations and triplet facts within a single sentence. Furthermore, certain approaches resort to treating relations as candidate choices integrated into prompt templates, leading to inefficient processing and suboptimal performance when tackling Document-Level Relation Extraction (DocRE) tasks, which entail handling multiple relations and triplet facts distributed across a given document, posing distinct challenges. To overcome these limitations, we introduce AutoRE, an end-to-end DocRE model that adopts a novel RE extraction paradigm named RHF (Relation-Head-Facts). Unlike existing approaches, AutoRE does not rely on the assumption of known relation options, making it more reflective of real-world scenarios. Additionally, we have developed an easily extensible RE framework using a Parameters Efficient Fine Tuning (PEFT) algorithm (QLoRA). Our experiments on the RE-DocRED dataset showcase AutoRE's best performance, achieving state-of-the-art results, surpassing TAG by 10.03% and 9.03% respectively on the dev and test set.
Supporting Mitosis Detection AI Training with Inter-Observer Eye-Gaze Consistencies
Apr 02, 2024The expansion of artificial intelligence (AI) in pathology tasks has intensified the demand for doctors' annotations in AI development. However, collecting high-quality annotations from doctors is costly and time-consuming, creating a bottleneck in AI progress. This study investigates eye-tracking as a cost-effective technology to collect doctors' behavioral data for AI training with a focus on the pathology task of mitosis detection. One major challenge in using eye-gaze data is the low signal-to-noise ratio, which hinders the extraction of meaningful information. We tackled this by levering the properties of inter-observer eye-gaze consistencies and creating eye-gaze labels from consistent eye-fixations shared by a group of observers. Our study involved 14 non-medical participants, from whom we collected eye-gaze data and generated eye-gaze labels based on varying group sizes. We assessed the efficacy of such eye-gaze labels by training Convolutional Neural Networks (CNNs) and comparing their performance to those trained with ground truth annotations and a heuristic-based baseline. Results indicated that CNNs trained with our eye-gaze labels closely followed the performance of ground-truth-based CNNs, and significantly outperformed the baseline. Although primarily focused on mitosis, we envision that insights from this study can be generalized to other medical imaging tasks.
Pipelined Biomedical Event Extraction Rivaling Joint Learning
Mar 19, 2024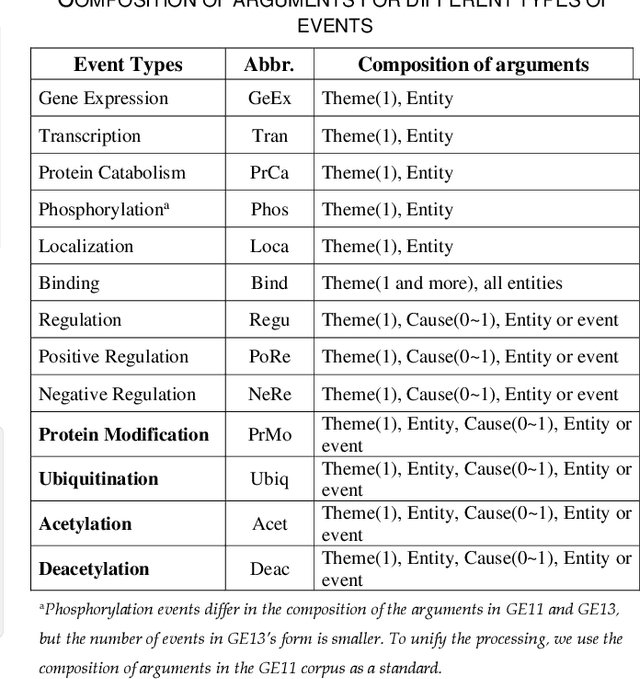
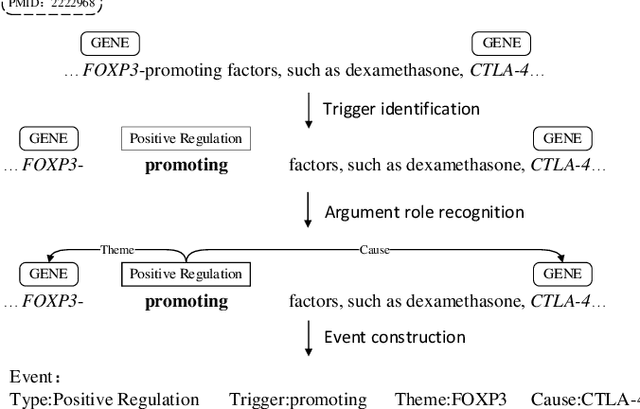
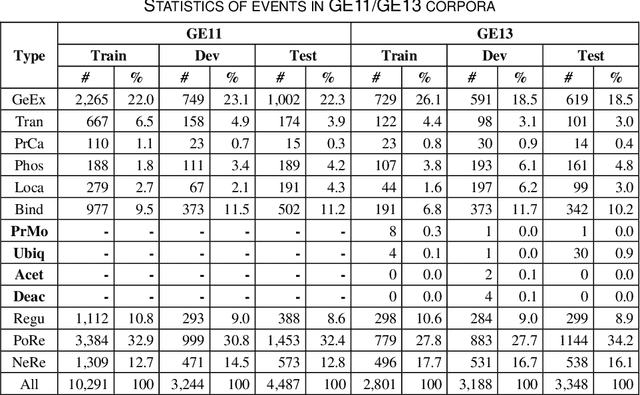
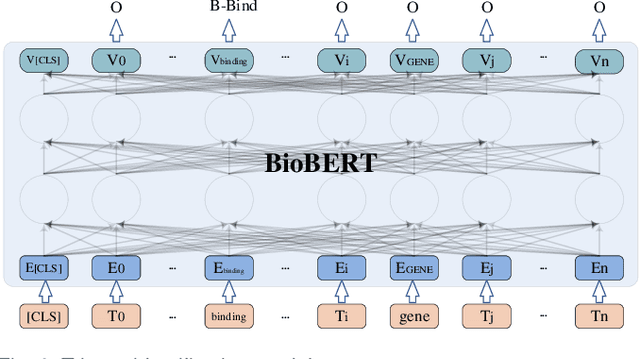
Biomedical event extraction is an information extraction task to obtain events from biomedical text, whose targets include the type, the trigger, and the respective arguments involved in an event. Traditional biomedical event extraction usually adopts a pipelined approach, which contains trigger identification, argument role recognition, and finally event construction either using specific rules or by machine learning. In this paper, we propose an n-ary relation extraction method based on the BERT pre-training model to construct Binding events, in order to capture the semantic information about an event's context and its participants. The experimental results show that our method achieves promising results on the GE11 and GE13 corpora of the BioNLP shared task with F1 scores of 63.14% and 59.40%, respectively. It demonstrates that by significantly improving theperformance of Binding events, the overall performance of the pipelined event extraction approach or even exceeds those of current joint learning methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge