"Information Extraction": models, code, and papers
Universal Information Extraction as Unified Semantic Matching
Jan 09, 2023
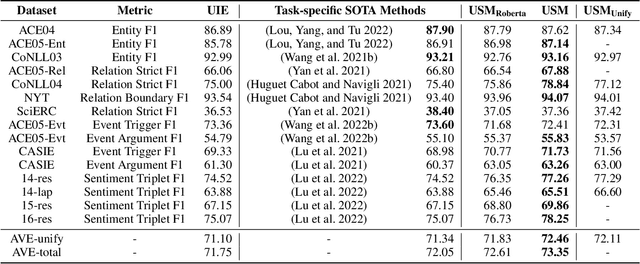
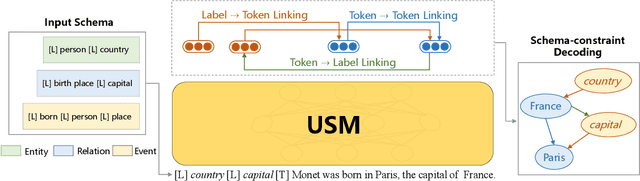
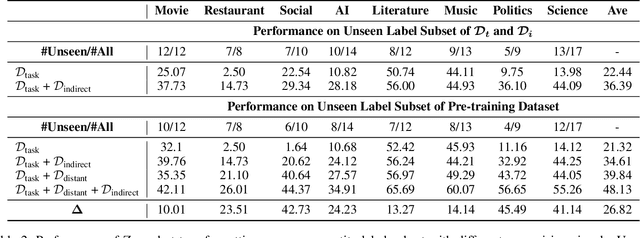
The challenge of information extraction (IE) lies in the diversity of label schemas and the heterogeneity of structures. Traditional methods require task-specific model design and rely heavily on expensive supervision, making them difficult to generalize to new schemas. In this paper, we decouple IE into two basic abilities, structuring and conceptualizing, which are shared by different tasks and schemas. Based on this paradigm, we propose to universally model various IE tasks with Unified Semantic Matching (USM) framework, which introduces three unified token linking operations to model the abilities of structuring and conceptualizing. In this way, USM can jointly encode schema and input text, uniformly extract substructures in parallel, and controllably decode target structures on demand. Empirical evaluation on 4 IE tasks shows that the proposed method achieves state-of-the-art performance under the supervised experiments and shows strong generalization ability in zero/few-shot transfer settings.
Document Layout Analysis on BaDLAD Dataset: A Comprehensive MViTv2 Based Approach
Aug 31, 2023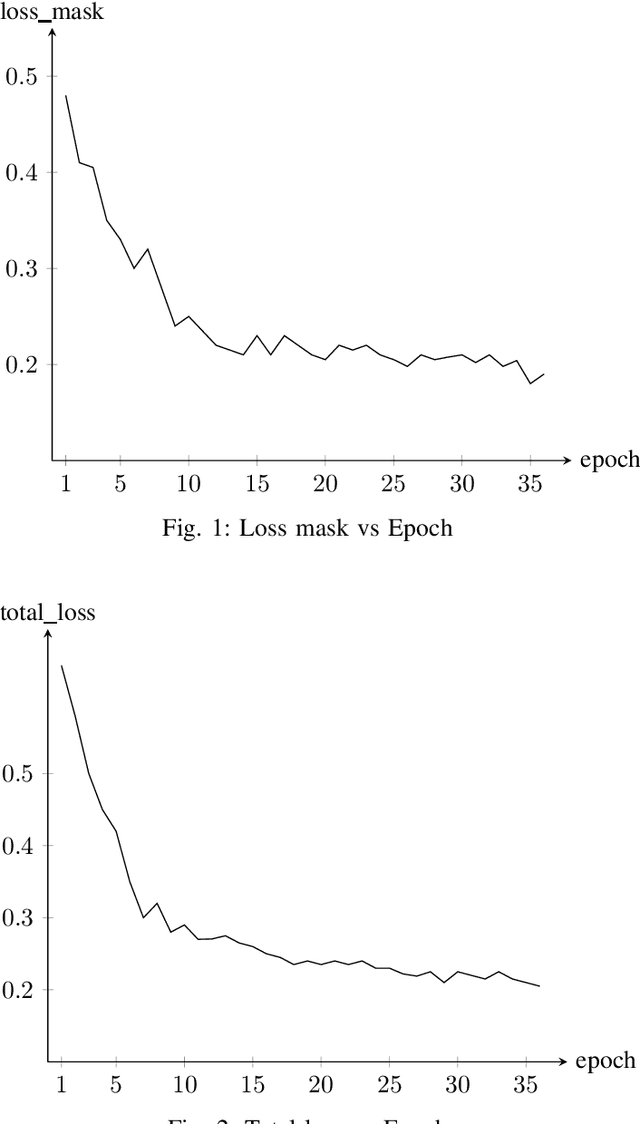
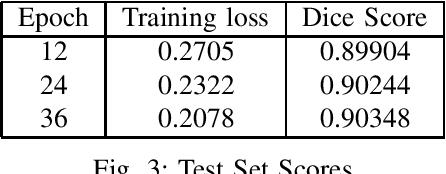
In the rapidly evolving digital era, the analysis of document layouts plays a pivotal role in automated information extraction and interpretation. In our work, we have trained MViTv2 transformer model architecture with cascaded mask R-CNN on BaDLAD dataset to extract text box, paragraphs, images and tables from a document. After training on 20365 document images for 36 epochs in a 3 phase cycle, we achieved a training loss of 0.2125 and a mask loss of 0.19. Our work extends beyond training, delving into the exploration of potential enhancement avenues. We investigate the impact of rotation and flip augmentation, the effectiveness of slicing input images pre-inference, the implications of varying the resolution of the transformer backbone, and the potential of employing a dual-pass inference to uncover missed text-boxes. Through these explorations, we observe a spectrum of outcomes, where some modifications result in tangible performance improvements, while others offer unique insights for future endeavors.
Continual Event Extraction with Semantic Confusion Rectification
Oct 24, 2023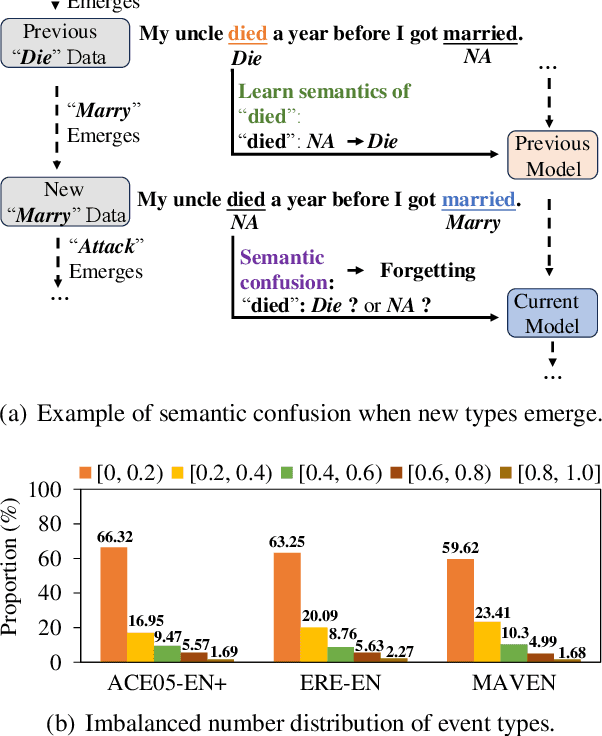
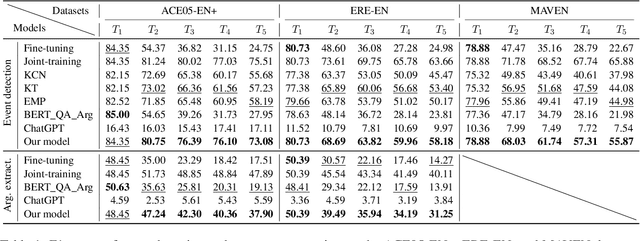
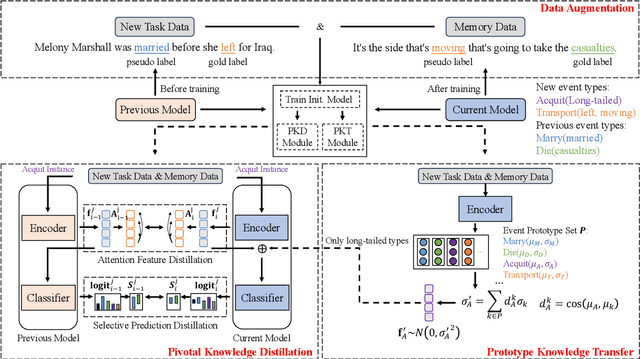
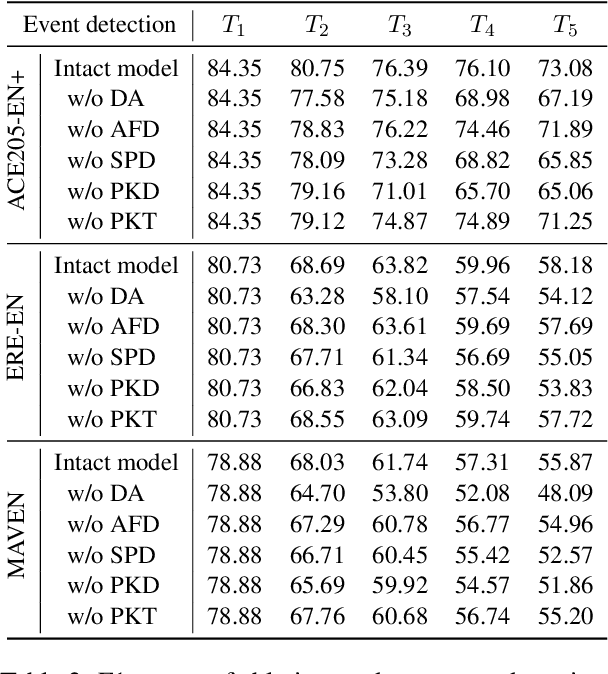
We study continual event extraction, which aims to extract incessantly emerging event information while avoiding forgetting. We observe that the semantic confusion on event types stems from the annotations of the same text being updated over time. The imbalance between event types even aggravates this issue. This paper proposes a novel continual event extraction model with semantic confusion rectification. We mark pseudo labels for each sentence to alleviate semantic confusion. We transfer pivotal knowledge between current and previous models to enhance the understanding of event types. Moreover, we encourage the model to focus on the semantics of long-tailed event types by leveraging other associated types. Experimental results show that our model outperforms state-of-the-art baselines and is proficient in imbalanced datasets.
HiNet: Novel Multi-Scenario & Multi-Task Learning with Hierarchical Information Extraction
Mar 14, 2023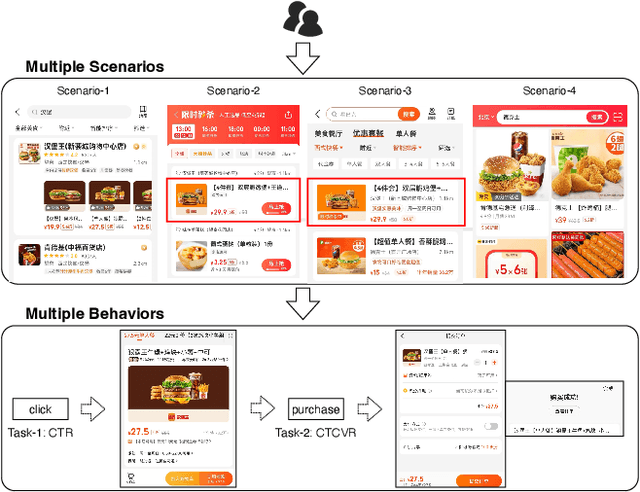
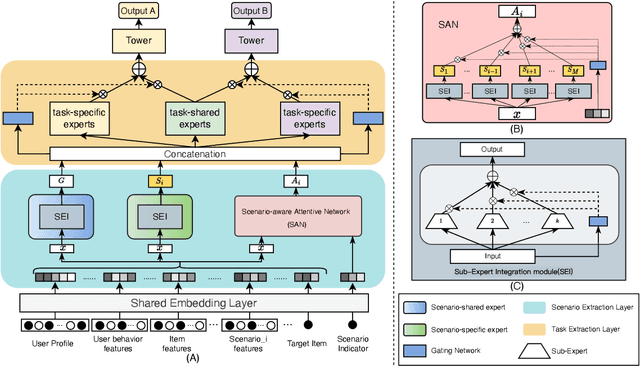
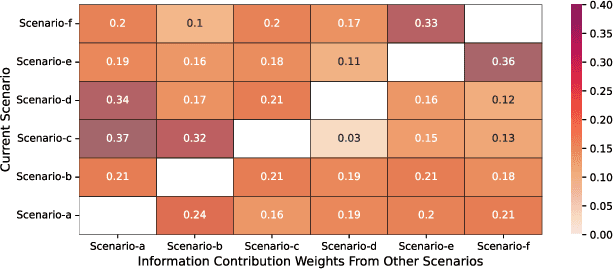
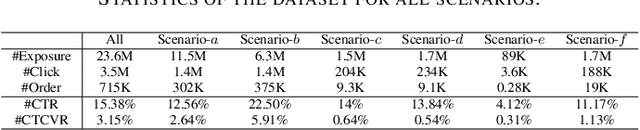
Multi-scenario & multi-task learning has been widely applied to many recommendation systems in industrial applications, wherein an effective and practical approach is to carry out multi-scenario transfer learning on the basis of the Mixture-of-Expert (MoE) architecture. However, the MoE-based method, which aims to project all information in the same feature space, cannot effectively deal with the complex relationships inherent among various scenarios and tasks, resulting in unsatisfactory performance. To tackle the problem, we propose a Hierarchical information extraction Network (HiNet) for multi-scenario and multi-task recommendation, which achieves hierarchical extraction based on coarse-to-fine knowledge transfer scheme. The multiple extraction layers of the hierarchical network enable the model to enhance the capability of transferring valuable information across scenarios while preserving specific features of scenarios and tasks. Furthermore, a novel scenario-aware attentive network module is proposed to model correlations between scenarios explicitly. Comprehensive experiments conducted on real-world industrial datasets from Meituan Meishi platform demonstrate that HiNet achieves a new state-of-the-art performance and significantly outperforms existing solutions. HiNet is currently fully deployed in two scenarios and has achieved 2.87% and 1.75% order quantity gain respectively.
C-RITNet: Set Infrared and Visible Image Fusion Free from Complementary Information Mining
Sep 13, 2023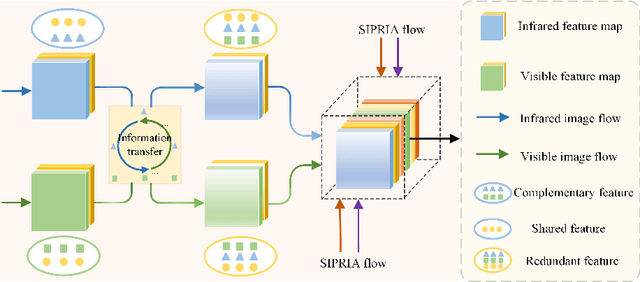
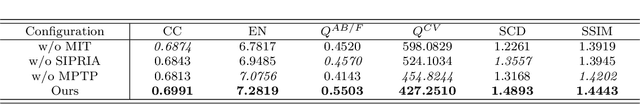
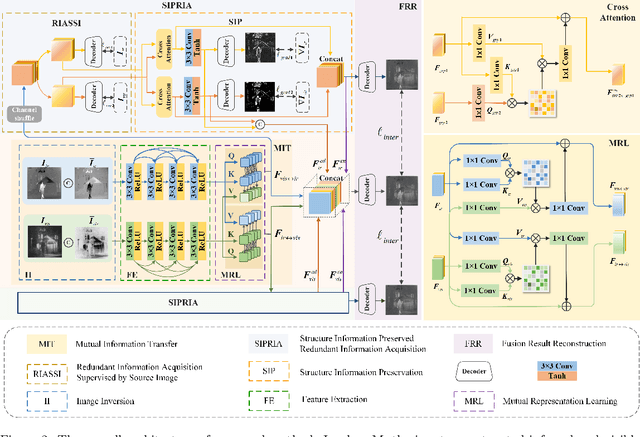
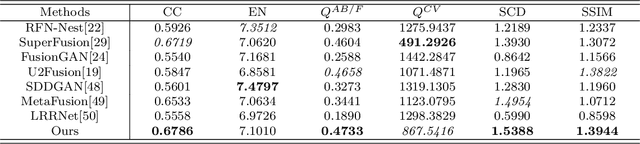
Infrared and visible image fusion (IVIF) aims to extract and integrate the complementary information in two different modalities to generate high-quality fused images with salient targets and abundant texture details. However, current image fusion methods go to great lengths to excavate complementary features, which is generally achieved through two efforts. On the one hand, the feature extraction network is expected to have excellent performance in extracting complementary information. On the other hand, complex fusion strategies are often designed to aggregate the complementary information. In other words, enabling the network to perceive and extract complementary information is extremely challenging. Complicated fusion strategies, while effective, still run the risk of losing weak edge details. To this end, this paper rethinks the IVIF outside the box, proposing a complementary-redundant information transfer network (C-RITNet). It reasonably transfers complementary information into redundant one, which integrates both the shared and complementary features from two modalities. Hence, the proposed method is able to alleviate the challenges posed by the complementary information extraction and reduce the reliance on sophisticated fusion strategies. Specifically, to skillfully sidestep aggregating complementary information in IVIF, we first design the mutual information transfer (MIT) module to mutually represent features from two modalities, roughly transferring complementary information into redundant one. Then, a redundant information acquisition supervised by source image (RIASSI) module is devised to further ensure the complementary-redundant information transfer after MIT. Meanwhile, we also propose a structure information preservation (SIP) module to guarantee that the edge structure information of the source images can be transferred to the fusion results.
Towards Generalized Open Information Extraction
Nov 29, 2022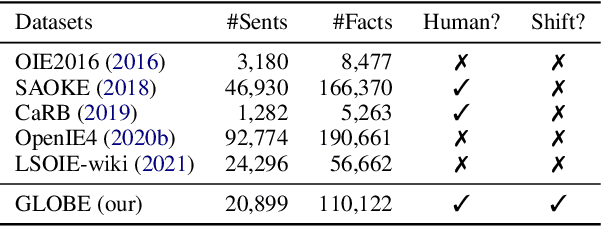
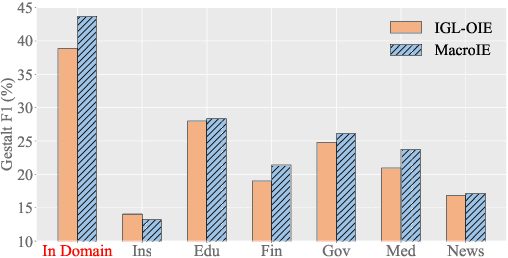
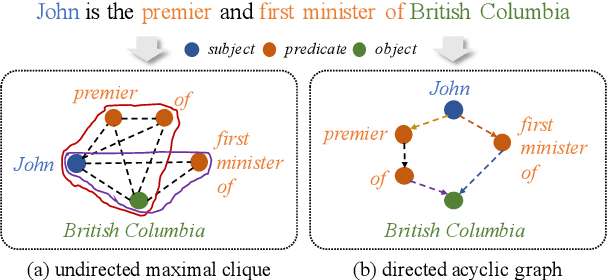
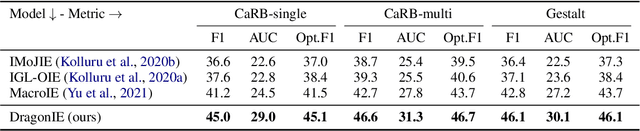
Open Information Extraction (OpenIE) facilitates the open-domain discovery of textual facts. However, the prevailing solutions evaluate OpenIE models on in-domain test sets aside from the training corpus, which certainly violates the initial task principle of domain-independence. In this paper, we propose to advance OpenIE towards a more realistic scenario: generalizing over unseen target domains with different data distributions from the source training domains, termed Generalized OpenIE. For this purpose, we first introduce GLOBE, a large-scale human-annotated multi-domain OpenIE benchmark, to examine the robustness of recent OpenIE models to domain shifts, and the relative performance degradation of up to 70% implies the challenges of generalized OpenIE. Then, we propose DragonIE, which explores a minimalist graph expression of textual fact: directed acyclic graph, to improve the OpenIE generalization. Extensive experiments demonstrate that DragonIE beats the previous methods in both in-domain and out-of-domain settings by as much as 6.0% in F1 score absolutely, but there is still ample room for improvement.
EMOFM: Ensemble MLP mOdel with Feature-based Mixers for Click-Through Rate Prediction
Oct 15, 2023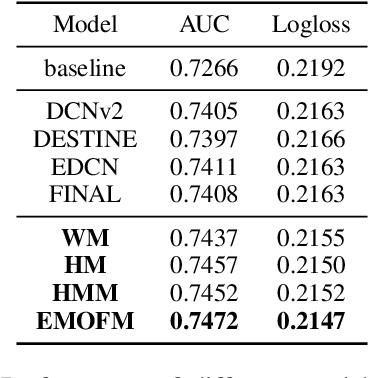
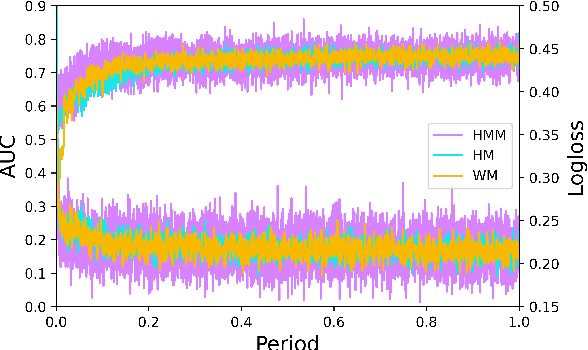
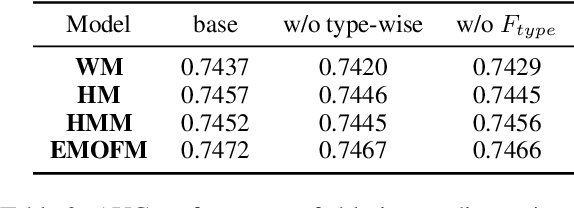
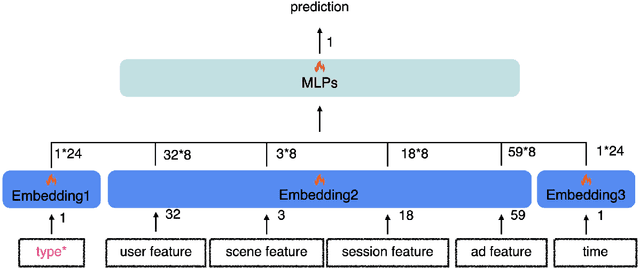
Track one of CTI competition is on click-through rate (CTR) prediction. The dataset contains millions of records and each field-wise feature in a record consists of hashed integers for privacy. For this task, the keys of network-based methods might be type-wise feature extraction and information fusion across different fields. Multi-layer perceptrons (MLPs) are able to extract field feature, but could not efficiently fuse features. Motivated by the natural fusion characteristic of cross attention and the efficiency of transformer-based structures, we propose simple plug-in mixers for field/type-wise feature fusion, and thus construct an field&type-wise ensemble model, namely EMOFM (Ensemble MLP mOdel with Feature-based Mixers). In the experiments, the proposed model is evaluated on the dataset, the optimization process is visualized and ablation studies are explored. It is shown that EMOFM outperforms compared baselines. In the end, we discuss on future work. WARNING: The comparison might not be fair enough since the proposed method is designed for this data in particular while compared methods are not. For example, EMOFM especially takes different types of interactions into consideration while others do not. Anyway, we do hope that the ideas inside our method could help other developers/learners/researchers/thinkers and so on.
Dual-Stream Knowledge-Preserving Hashing for Unsupervised Video Retrieval
Oct 12, 2023Unsupervised video hashing usually optimizes binary codes by learning to reconstruct input videos. Such reconstruction constraint spends much effort on frame-level temporal context changes without focusing on video-level global semantics that are more useful for retrieval. Hence, we address this problem by decomposing video information into reconstruction-dependent and semantic-dependent information, which disentangles the semantic extraction from reconstruction constraint. Specifically, we first design a simple dual-stream structure, including a temporal layer and a hash layer. Then, with the help of semantic similarity knowledge obtained from self-supervision, the hash layer learns to capture information for semantic retrieval, while the temporal layer learns to capture the information for reconstruction. In this way, the model naturally preserves the disentangled semantics into binary codes. Validated by comprehensive experiments, our method consistently outperforms the state-of-the-arts on three video benchmarks.
Dual Defense: Adversarial, Traceable, and Invisible Robust Watermarking against Face Swapping
Oct 25, 2023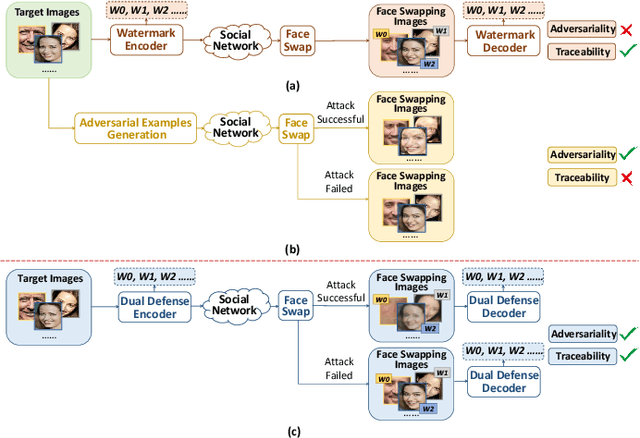
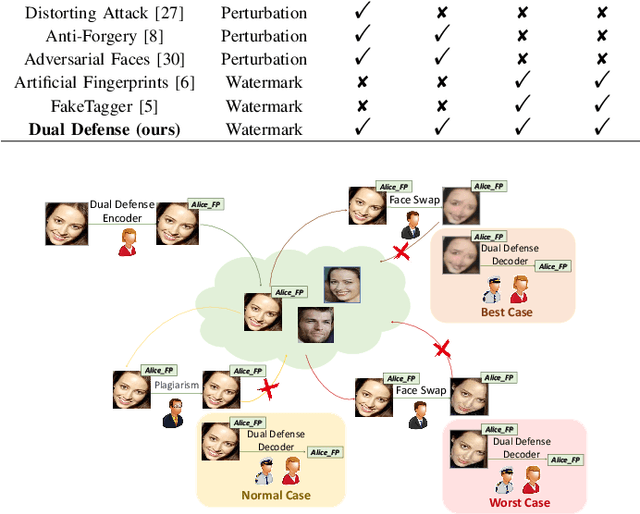
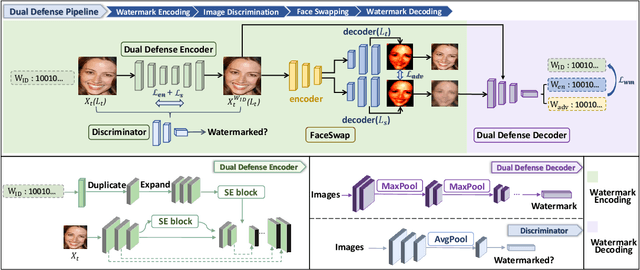
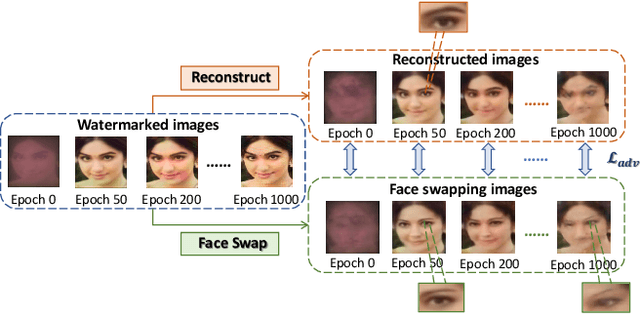
The malicious applications of deep forgery, represented by face swapping, have introduced security threats such as misinformation dissemination and identity fraud. While some research has proposed the use of robust watermarking methods to trace the copyright of facial images for post-event traceability, these methods cannot effectively prevent the generation of forgeries at the source and curb their dissemination. To address this problem, we propose a novel comprehensive active defense mechanism that combines traceability and adversariality, called Dual Defense. Dual Defense invisibly embeds a single robust watermark within the target face to actively respond to sudden cases of malicious face swapping. It disrupts the output of the face swapping model while maintaining the integrity of watermark information throughout the entire dissemination process. This allows for watermark extraction at any stage of image tracking for traceability. Specifically, we introduce a watermark embedding network based on original-domain feature impersonation attack. This network learns robust adversarial features of target facial images and embeds watermarks, seeking a well-balanced trade-off between watermark invisibility, adversariality, and traceability through perceptual adversarial encoding strategies. Extensive experiments demonstrate that Dual Defense achieves optimal overall defense success rates and exhibits promising universality in anti-face swapping tasks and dataset generalization ability. It maintains impressive adversariality and traceability in both original and robust settings, surpassing current forgery defense methods that possess only one of these capabilities, including CMUA-Watermark, Anti-Forgery, FakeTagger, or PGD methods.
On Event Individuation for Document-Level Information Extraction
Dec 19, 2022
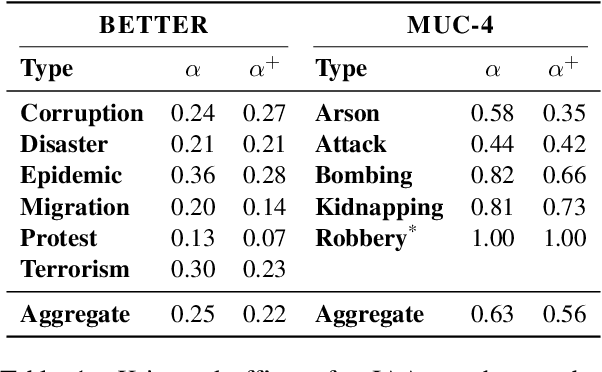

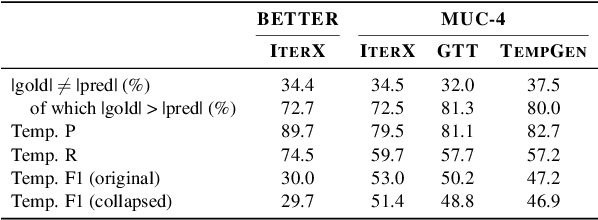
As information extraction (IE) systems have grown more capable at whole-document extraction, the classic task of \emph{template filling} has seen renewed interest as a benchmark for evaluating them. In this position paper, we call into question the suitability of template filling for this purpose. We argue that the task demands definitive answers to thorny questions of \emph{event individuation} -- the problem of distinguishing distinct events -- about which even human experts disagree. We show through annotation studies and error analysis that this raises concerns about the usefulness of template filling evaluation metrics, the quality of datasets for the task, and the ability of models to learn it. Finally, we consider possible solutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge