"Information Extraction": models, code, and papers
Natural Disaster Analysis using Satellite Imagery and Social-Media Data for Emergency Response Situations
Nov 16, 2023
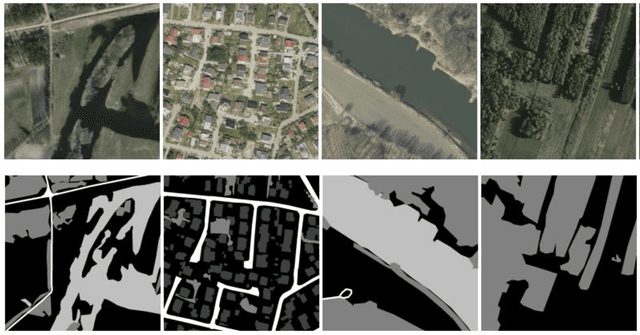


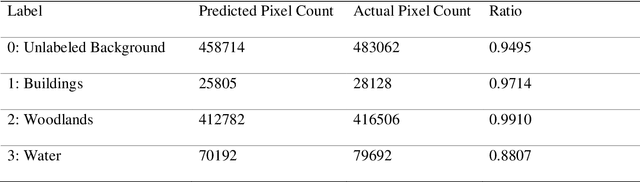
Disaster Management is one of the most promising research areas because of its significant economic, environmental and social repercussions. This research focuses on analyzing different types of data (pre and post satellite images and twitter data) related to disaster management for in-depth analysis of location-wise emergency requirements. This research has been divided into two stages, namely, satellite image analysis and twitter data analysis followed by integration using location. The first stage involves pre and post disaster satellite image analysis of the location using multi-class land cover segmentation technique based on U-Net architecture. The second stage focuses on mapping the region with essential information about the disaster situation and immediate requirements for relief operations. The severely affected regions are demarcated and twitter data is extracted using keywords respective to that location. The extraction of situational information from a large corpus of raw tweets adopts Content Word based Tweet Summarization (COWTS) technique. An integration of these modules using real-time location-based mapping and frequency analysis technique gathers multi-dimensional information in the advent of disaster occurrence such as the Kerala and Mississippi floods that were analyzed and validated as test cases. The novelty of this research lies in the application of segmented satellite images for disaster relief using highlighted land cover changes and integration of twitter data by mapping these region-specific filters for obtaining a complete overview of the disaster.
One Signal-Noise Separation based Wiener Filter for Magnetogastrogram
Nov 12, 2023Magnetogastrogram (MGG) signal frequency is about 0.05 Hz, the low-frequency environmental noise interference is serious and can be several times stronger in magnitude than the signals of interest and may severely impede the extraction of relevant information. Wiener filter is one classic denoising solution for biomagnetic applications. Since the reference channels are usually placed not far enough from the biomagnetic sources under test, they will inevitably detect the signals and the Wiener filters may produce ill-conditioned solutions. Considering the solutions to improve the signal-to-noise ratio (SNR) of Wiener filter output, there are few methods to separate the signals from the noises of the reference signal at the filter input. In this paper, a new signal processing framework called signal-noise separation based Wiener filter (SNSWF) is proposed that it separates the main noise as the input signal of the filter to improve the output SNR of Wiener filter. The filter was successfully applied to the noise suppression for MGG signal detection. Using the SNSWF, the filter SNR is 16.7 dB better than the classic Wiener filter.
CARLG: Leveraging Contextual Clues and Role Correlations for Improving Document-level Event Argument Extraction
Oct 08, 2023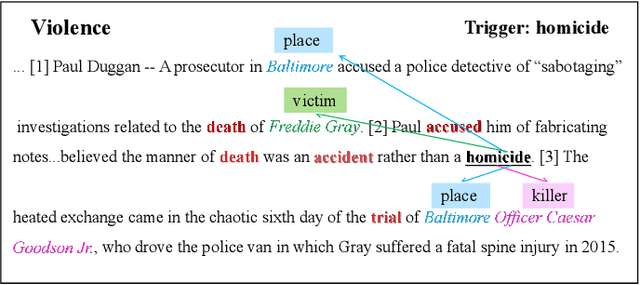
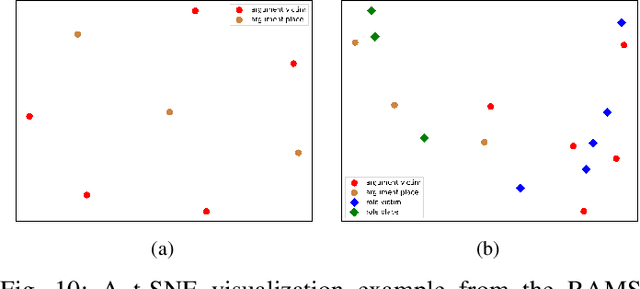
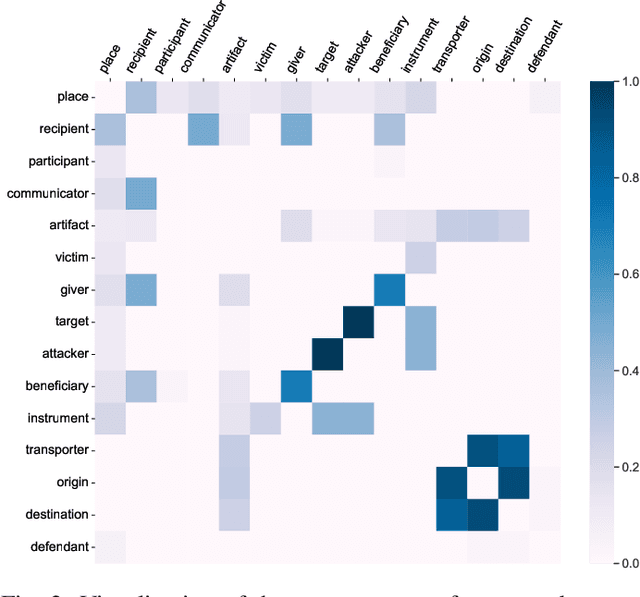
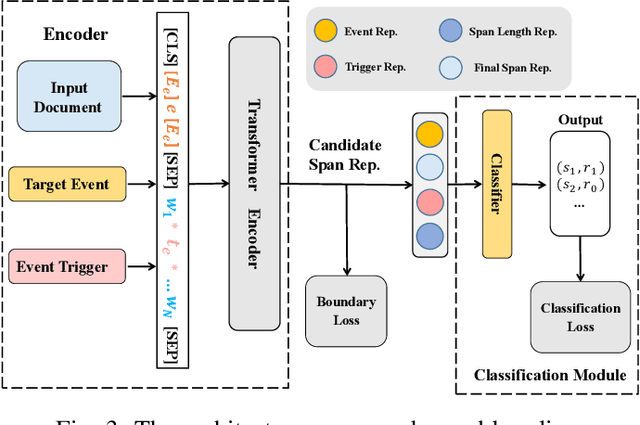
Document-level event argument extraction (EAE) is a crucial but challenging subtask in information extraction. Most existing approaches focus on the interaction between arguments and event triggers, ignoring two critical points: the information of contextual clues and the semantic correlations among argument roles. In this paper, we propose the CARLG model, which consists of two modules: Contextual Clues Aggregation (CCA) and Role-based Latent Information Guidance (RLIG), effectively leveraging contextual clues and role correlations for improving document-level EAE. The CCA module adaptively captures and integrates contextual clues by utilizing context attention weights from a pre-trained encoder. The RLIG module captures semantic correlations through role-interactive encoding and provides valuable information guidance with latent role representation. Notably, our CCA and RLIG modules are compact, transplantable and efficient, which introduce no more than 1% new parameters and can be easily equipped on other span-base methods with significant performance boost. Extensive experiments on the RAMS, WikiEvents, and MLEE datasets demonstrate the superiority of the proposed CARLG model. It outperforms previous state-of-the-art approaches by 1.26 F1, 1.22 F1, and 1.98 F1, respectively, while reducing the inference time by 31%. Furthermore, we provide detailed experimental analyses based on the performance gains and illustrate the interpretability of our model.
Modeling Entities as Semantic Points for Visual Information Extraction in the Wild
Mar 23, 2023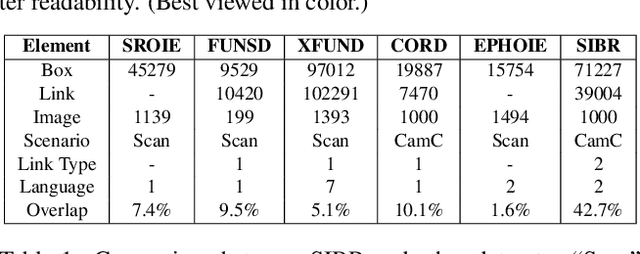
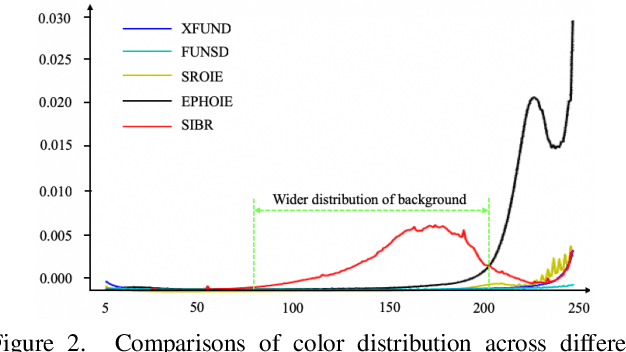
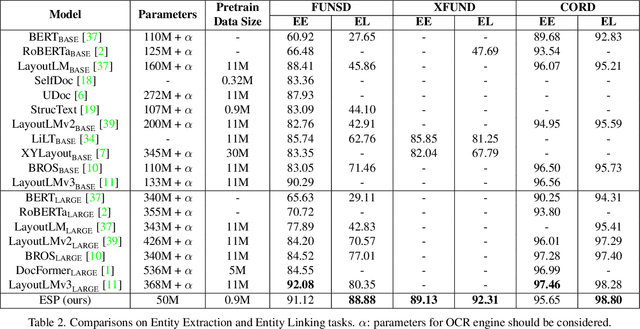
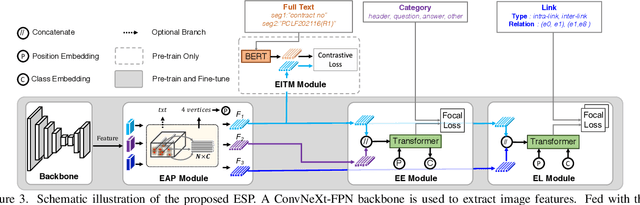
Recently, Visual Information Extraction (VIE) has been becoming increasingly important in both the academia and industry, due to the wide range of real-world applications. Previously, numerous works have been proposed to tackle this problem. However, the benchmarks used to assess these methods are relatively plain, i.e., scenarios with real-world complexity are not fully represented in these benchmarks. As the first contribution of this work, we curate and release a new dataset for VIE, in which the document images are much more challenging in that they are taken from real applications, and difficulties such as blur, partial occlusion, and printing shift are quite common. All these factors may lead to failures in information extraction. Therefore, as the second contribution, we explore an alternative approach to precisely and robustly extract key information from document images under such tough conditions. Specifically, in contrast to previous methods, which usually either incorporate visual information into a multi-modal architecture or train text spotting and information extraction in an end-to-end fashion, we explicitly model entities as semantic points, i.e., center points of entities are enriched with semantic information describing the attributes and relationships of different entities, which could largely benefit entity labeling and linking. Extensive experiments on standard benchmarks in this field as well as the proposed dataset demonstrate that the proposed method can achieve significantly enhanced performance on entity labeling and linking, compared with previous state-of-the-art models. Dataset is available at https://www.modelscope.cn/datasets/damo/SIBR/summary.
Burning the Adversarial Bridges: Robust Windows Malware Detection Against Binary-level Mutations
Oct 05, 2023Toward robust malware detection, we explore the attack surface of existing malware detection systems. We conduct root-cause analyses of the practical binary-level black-box adversarial malware examples. Additionally, we uncover the sensitivity of volatile features within the detection engines and exhibit their exploitability. Highlighting volatile information channels within the software, we introduce three software pre-processing steps to eliminate the attack surface, namely, padding removal, software stripping, and inter-section information resetting. Further, to counter the emerging section injection attacks, we propose a graph-based section-dependent information extraction scheme for software representation. The proposed scheme leverages aggregated information within various sections in the software to enable robust malware detection and mitigate adversarial settings. Our experimental results show that traditional malware detection models are ineffective against adversarial threats. However, the attack surface can be largely reduced by eliminating the volatile information. Therefore, we propose simple-yet-effective methods to mitigate the impacts of binary manipulation attacks. Overall, our graph-based malware detection scheme can accurately detect malware with an area under the curve score of 88.32\% and a score of 88.19% under a combination of binary manipulation attacks, exhibiting the efficiency of our proposed scheme.
An Information Extraction Study: Take In Mind the Tokenization!
Apr 01, 2023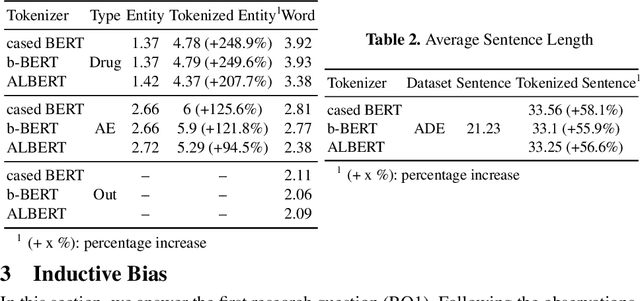
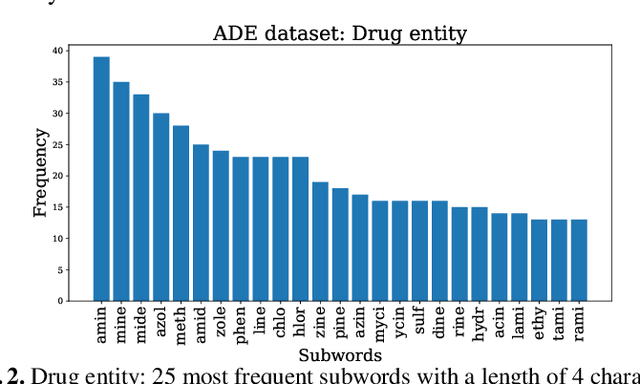
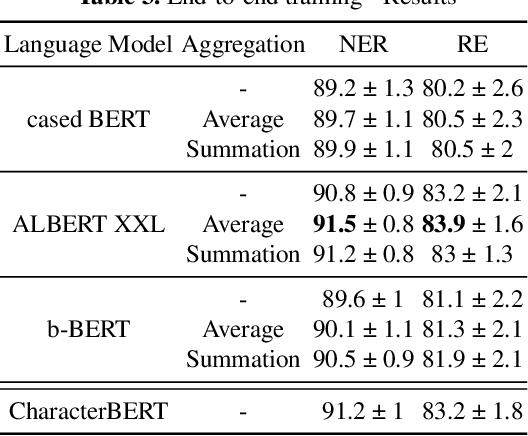
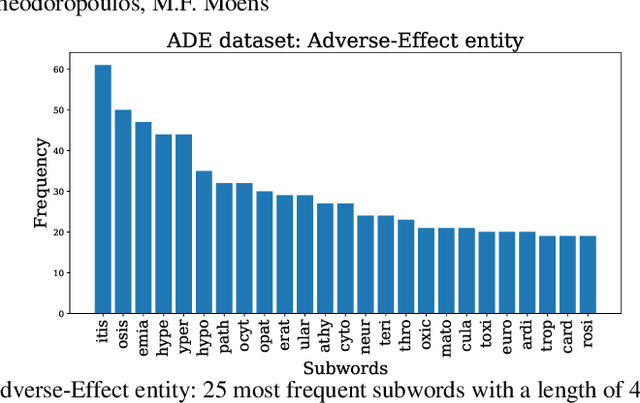
Current research on the advantages and trade-offs of using characters, instead of tokenized text, as input for deep learning models, has evolved substantially. New token-free models remove the traditional tokenization step; however, their efficiency remains unclear. Moreover, the effect of tokenization is relatively unexplored in sequence tagging tasks. To this end, we investigate the impact of tokenization when extracting information from documents and present a comparative study and analysis of subword-based and character-based models. Specifically, we study Information Extraction (IE) from biomedical texts. The main outcome is twofold: tokenization patterns can introduce inductive bias that results in state-of-the-art performance, and the character-based models produce promising results; thus, transitioning to token-free IE models is feasible.
SpACNN-LDVAE: Spatial Attention Convolutional Latent Dirichlet Variational Autoencoder for Hyperspectral Pixel Unmixing
Nov 17, 2023The Hyperspectral Unxming problem is to find the pure spectral signal of the underlying materials (endmembers) and their proportions (abundances). The proposed method builds upon the recently proposed method, Latent Dirichlet Variational Autoencoder (LDVAE). It assumes that abundances can be encoded as Dirichlet Distributions while mixed pixels and endmembers are represented by Multivariate Normal Distributions. However, LDVAE does not leverage spatial information present in an HSI; we propose an Isotropic CNN encoder with spatial attention to solve the hyperspectral unmixing problem. We evaluated our model on Samson, Hydice Urban, Cuprite, and OnTech-HSI-Syn-21 datasets. Our model also leverages the transfer learning paradigm for Cuprite Dataset, where we train the model on synthetic data and evaluate it on real-world data. We are able to observe the improvement in the results for the endmember extraction and abundance estimation by incorporating the spatial information. Code can be found at https://github.com/faisalqureshi/cnn-ldvae
Information Redundancy and Biases in Public Document Information Extraction Benchmarks
Apr 28, 2023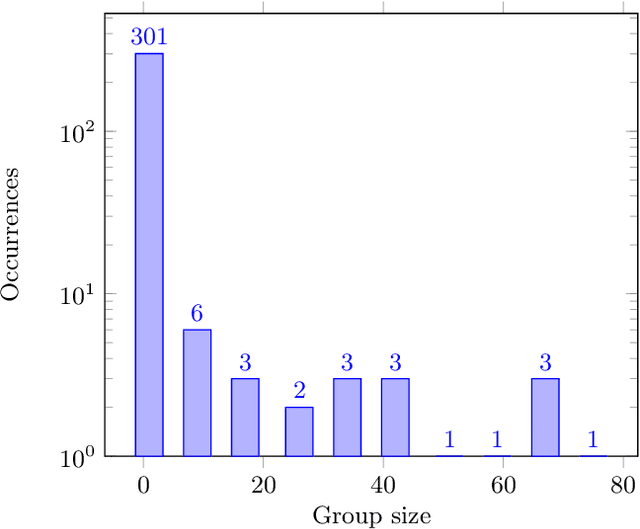
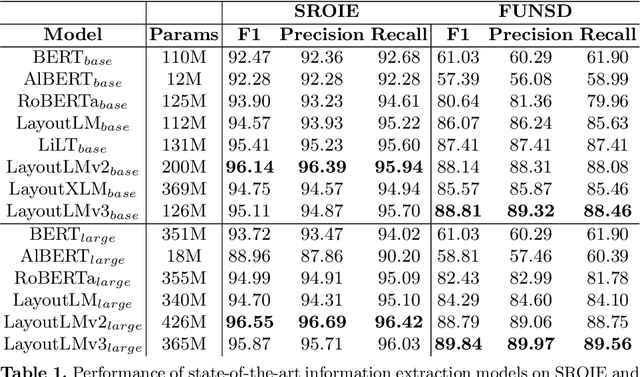

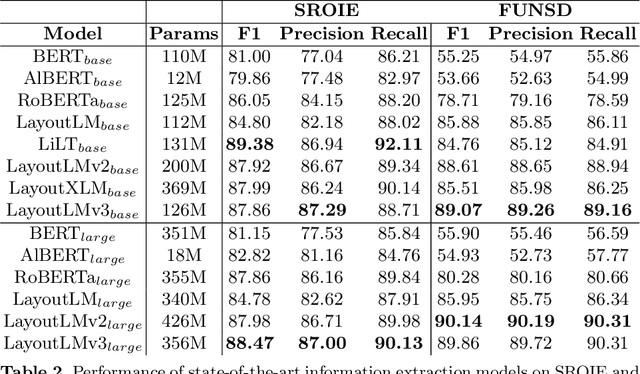
Advances in the Visually-rich Document Understanding (VrDU) field and particularly the Key-Information Extraction (KIE) task are marked with the emergence of efficient Transformer-based approaches such as the LayoutLM models. Despite the good performance of KIE models when fine-tuned on public benchmarks, they still struggle to generalize on complex real-life use-cases lacking sufficient document annotations. Our research highlighted that KIE standard benchmarks such as SROIE and FUNSD contain significant similarity between training and testing documents and can be adjusted to better evaluate the generalization of models. In this work, we designed experiments to quantify the information redundancy in public benchmarks, revealing a 75% template replication in SROIE official test set and 16% in FUNSD. We also proposed resampling strategies to provide benchmarks more representative of the generalization ability of models. We showed that models not suited for document analysis struggle on the adjusted splits dropping on average 10,5% F1 score on SROIE and 3.5% on FUNSD compared to multi-modal models dropping only 7,5% F1 on SROIE and 0.5% F1 on FUNSD.
From Large Language Models to Knowledge Graphs for Biomarker Discovery in Cancer
Oct 12, 2023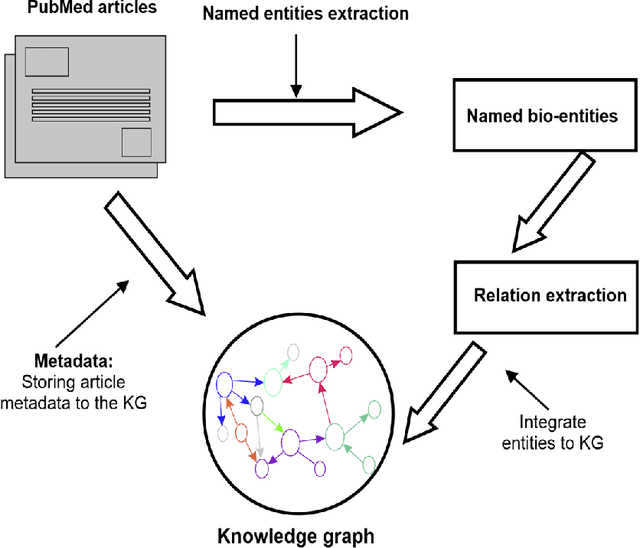
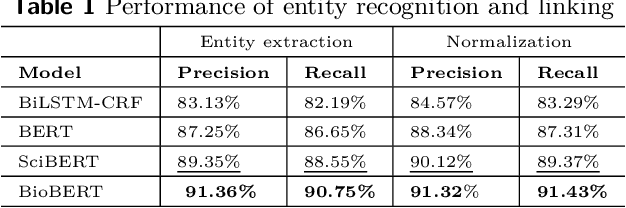
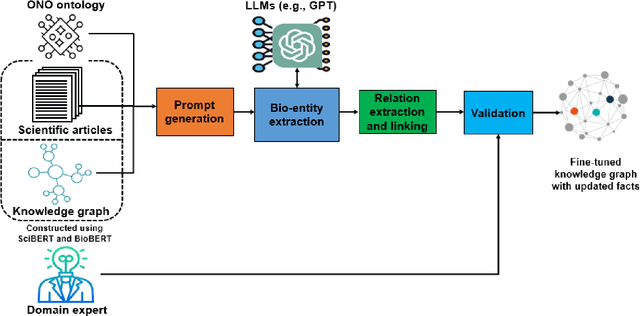
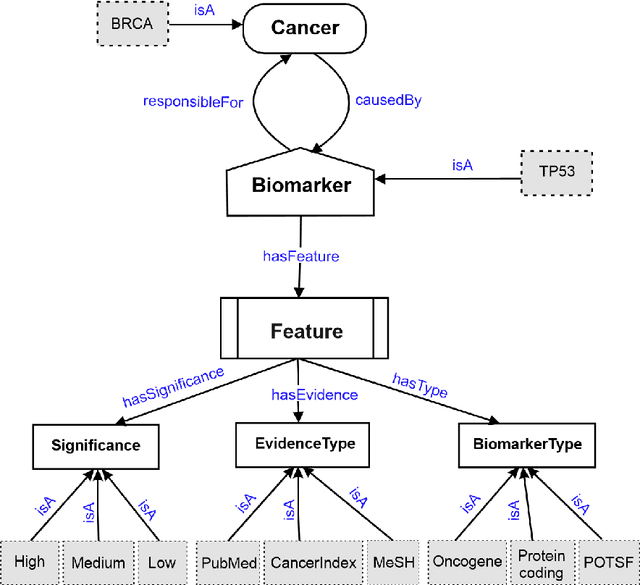
Domain experts often rely on up-to-date knowledge for apprehending and disseminating specific biological processes that help them design strategies to develop prevention and therapeutic decision-making. A challenging scenario for artificial intelligence (AI) is using biomedical data (e.g., texts, imaging, omics, and clinical) to provide diagnosis and treatment recommendations for cancerous conditions. Data and knowledge about cancer, drugs, genes, proteins, and their mechanism is spread across structured (knowledge bases (KBs)) and unstructured (e.g., scientific articles) sources. A large-scale knowledge graph (KG) can be constructed by integrating these data, followed by extracting facts about semantically interrelated entities and relations. Such KGs not only allow exploration and question answering (QA) but also allow domain experts to deduce new knowledge. However, exploring and querying large-scale KGs is tedious for non-domain users due to a lack of understanding of the underlying data assets and semantic technologies. In this paper, we develop a domain KG to leverage cancer-specific biomarker discovery and interactive QA. For this, a domain ontology called OncoNet Ontology (ONO) is developed to enable semantic reasoning for validating gene-disease relations. The KG is then enriched by harmonizing the ONO, controlled vocabularies, and additional biomedical concepts from scientific articles by employing BioBERT- and SciBERT-based information extraction (IE) methods. Further, since the biomedical domain is evolving, where new findings often replace old ones, without employing up-to-date findings, there is a high chance an AI system exhibits concept drift while providing diagnosis and treatment. Therefore, we finetuned the KG using large language models (LLMs) based on more recent articles and KBs that might not have been seen by the named entity recognition models.
Automated clinical coding using off-the-shelf large language models
Oct 10, 2023The task of assigning diagnostic ICD codes to patient hospital admissions is typically performed by expert human coders. Efforts towards automated ICD coding are dominated by supervised deep learning models. However, difficulties in learning to predict the large number of rare codes remain a barrier to adoption in clinical practice. In this work, we leverage off-the-shelf pre-trained generative large language models (LLMs) to develop a practical solution that is suitable for zero-shot and few-shot code assignment. Unsupervised pre-training alone does not guarantee precise knowledge of the ICD ontology and specialist clinical coding task, therefore we frame the task as information extraction, providing a description of each coded concept and asking the model to retrieve related mentions. For efficiency, rather than iterating over all codes, we leverage the hierarchical nature of the ICD ontology to sparsely search for relevant codes. Then, in a second stage, which we term 'meta-refinement', we utilise GPT-4 to select a subset of the relevant labels as predictions. We validate our method using Llama-2, GPT-3.5 and GPT-4 on the CodiEsp dataset of ICD-coded clinical case documents. Our tree-search method achieves state-of-the-art performance on rarer classes, achieving the best macro-F1 of 0.225, whilst achieving slightly lower micro-F1 of 0.157, compared to 0.216 and 0.219 respectively from PLM-ICD. To the best of our knowledge, this is the first method for automated ICD coding requiring no task-specific learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge