"Information Extraction": models, code, and papers
Less is More: Learning Reference Knowledge Using No-Reference Image Quality Assessment
Dec 01, 2023Image Quality Assessment (IQA) with reference images have achieved great success by imitating the human vision system, in which the image quality is effectively assessed by comparing the query image with its pristine reference image. However, for the images in the wild, it is quite difficult to access accurate reference images. We argue that it is possible to learn reference knowledge under the No-Reference Image Quality Assessment (NR-IQA) setting, which is effective and efficient empirically. Concretely, by innovatively introducing a novel feature distillation method in IQA, we propose a new framework to learn comparative knowledge from non-aligned reference images. And then, to achieve fast convergence and avoid overfitting, we further propose an inductive bias regularization. Such a framework not only solves the congenital defects of NR-IQA but also improves the feature extraction framework, enabling it to express more abundant quality information. Surprisingly, our method utilizes less input while obtaining a more significant improvement compared to the teacher models. Extensive experiments on eight standard NR-IQA datasets demonstrate the superior performance to the state-of-the-art NR-IQA methods, i.e., achieving the PLCC values of 0.917 (vs. 0.884 in LIVEC) and 0.686 (vs. 0.661 in LIVEFB).
Sub-network Discovery and Soft-masking for Continual Learning of Mixed Tasks
Oct 13, 2023Continual learning (CL) has two main objectives: preventing catastrophic forgetting (CF) and encouraging knowledge transfer (KT). The existing literature mainly focused on overcoming CF. Some work has also been done on KT when the tasks are similar. To our knowledge, only one method has been proposed to learn a sequence of mixed tasks. However, these techniques still suffer from CF and/or limited KT. This paper proposes a new CL method to achieve both. It overcomes CF by isolating the knowledge of each task via discovering a subnetwork for it. A soft-masking mechanism is also proposed to preserve the previous knowledge and to enable the new task to leverage the past knowledge to achieve KT. Experiments using classification, generation, information extraction, and their mixture (i.e., heterogeneous tasks) show that the proposed method consistently outperforms strong baselines.
* https://github.com/ZixuanKe/PyContinual
DocParser: End-to-end OCR-free Information Extraction from Visually Rich Documents
May 01, 2023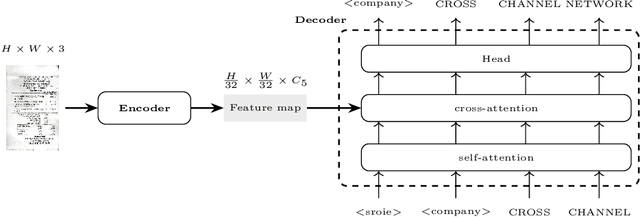
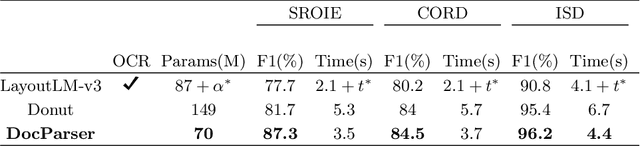
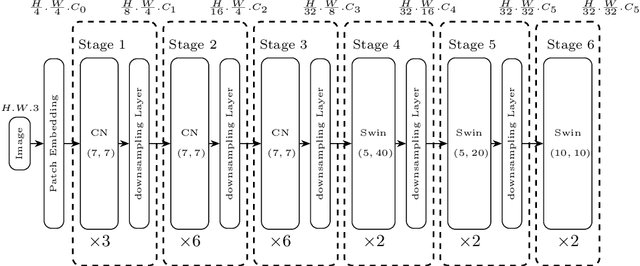

Information Extraction from visually rich documents is a challenging task that has gained a lot of attention in recent years due to its importance in several document-control based applications and its widespread commercial value. The majority of the research work conducted on this topic to date follow a two-step pipeline. First, they read the text using an off-the-shelf Optical Character Recognition (OCR) engine, then, they extract the fields of interest from the obtained text. The main drawback of these approaches is their dependence on an external OCR system, which can negatively impact both performance and computational speed. Recent OCR-free methods were proposed to address the previous issues. Inspired by their promising results, we propose in this paper an OCR-free end-to-end information extraction model named DocParser. It differs from prior end-to-end approaches by its ability to better extract discriminative character features. DocParser achieves state-of-the-art results on various datasets, while still being faster than previous works.
A Knowledge Graph-Based Search Engine for Robustly Finding Doctors and Locations in the Healthcare Domain
Oct 08, 2023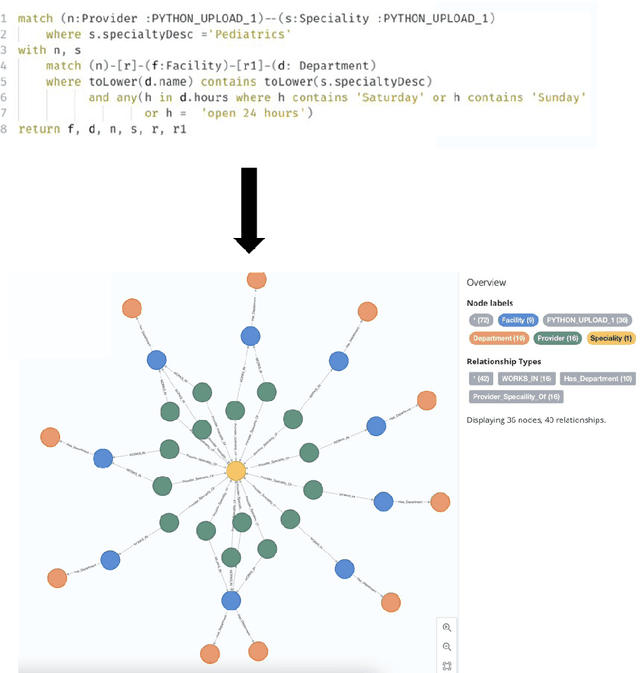
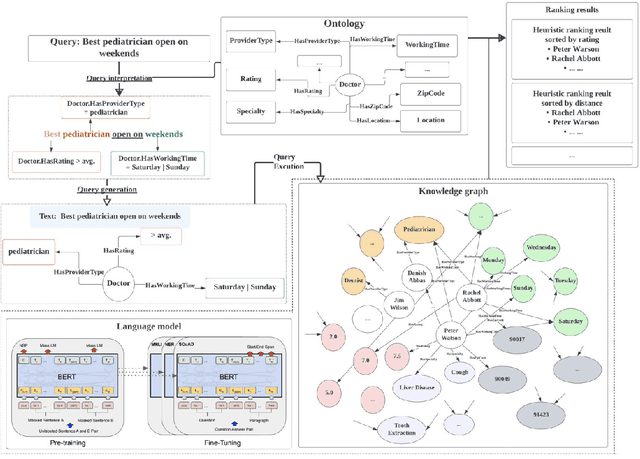
Efficiently finding doctors and locations is an important search problem for patients in the healthcare domain, for which traditional information retrieval methods tend not to work optimally. In the last ten years, knowledge graphs (KGs) have emerged as a powerful way to combine the benefits of gleaning insights from semi-structured data using semantic modeling, natural language processing techniques like information extraction, and robust querying using structured query languages like SPARQL and Cypher. In this short paper, we present a KG-based search engine architecture for robustly finding doctors and locations in the healthcare domain. Early results demonstrate that our approach can lead to significantly higher coverage for complex queries without degrading quality.
CAW-coref: Conjunction-Aware Word-level Coreference Resolution
Oct 09, 2023State-of-the-art coreference resolutions systems depend on multiple LLM calls per document and are thus prohibitively expensive for many use cases (e.g., information extraction with large corpora). The leading word-level coreference system (WL-coref) attains 96.6% of these SOTA systems' performance while being much more efficient. In this work, we identify a routine yet important failure case of WL-coref: dealing with conjoined mentions such as 'Tom and Mary'. We offer a simple yet effective solution that improves the performance on the OntoNotes test set by 0.9% F1, shrinking the gap between efficient word-level coreference resolution and expensive SOTA approaches by 34.6%. Our Conjunction-Aware Word-level coreference model (CAW-coref) and code is available at https://github.com/KarelDO/wl-coref.
Cooperative Dual Attention for Audio-Visual Speech Enhancement with Facial Cues
Nov 24, 2023In this work, we focus on leveraging facial cues beyond the lip region for robust Audio-Visual Speech Enhancement (AVSE). The facial region, encompassing the lip region, reflects additional speech-related attributes such as gender, skin color, nationality, etc., which contribute to the effectiveness of AVSE. However, static and dynamic speech-unrelated attributes also exist, causing appearance changes during speech. To address these challenges, we propose a Dual Attention Cooperative Framework, DualAVSE, to ignore speech-unrelated information, capture speech-related information with facial cues, and dynamically integrate it with the audio signal for AVSE. Specifically, we introduce a spatial attention-based visual encoder to capture and enhance visual speech information beyond the lip region, incorporating global facial context and automatically ignoring speech-unrelated information for robust visual feature extraction. Additionally, a dynamic visual feature fusion strategy is introduced by integrating a temporal-dimensional self-attention module, enabling the model to robustly handle facial variations. The acoustic noise in the speaking process is variable, impacting audio quality. Therefore, a dynamic fusion strategy for both audio and visual features is introduced to address this issue. By integrating cooperative dual attention in the visual encoder and audio-visual fusion strategy, our model effectively extracts beneficial speech information from both audio and visual cues for AVSE. Thorough analysis and comparison on different datasets, including normal and challenging cases with unreliable or absent visual information, consistently show our model outperforming existing methods across multiple metrics.
Leveraging deep active learning to identify low-resource mobility functioning information in public clinical notes
Nov 27, 2023Function is increasingly recognized as an important indicator of whole-person health, although it receives little attention in clinical natural language processing research. We introduce the first public annotated dataset specifically on the Mobility domain of the International Classification of Functioning, Disability and Health (ICF), aiming to facilitate automatic extraction and analysis of functioning information from free-text clinical notes. We utilize the National NLP Clinical Challenges (n2c2) research dataset to construct a pool of candidate sentences using keyword expansion. Our active learning approach, using query-by-committee sampling weighted by density representativeness, selects informative sentences for human annotation. We train BERT and CRF models, and use predictions from these models to guide the selection of new sentences for subsequent annotation iterations. Our final dataset consists of 4,265 sentences with a total of 11,784 entities, including 5,511 Action entities, 5,328 Mobility entities, 306 Assistance entities, and 639 Quantification entities. The inter-annotator agreement (IAA), averaged over all entity types, is 0.72 for exact matching and 0.91 for partial matching. We also train and evaluate common BERT models and state-of-the-art Nested NER models. The best F1 scores are 0.84 for Action, 0.7 for Mobility, 0.62 for Assistance, and 0.71 for Quantification. Empirical results demonstrate promising potential of NER models to accurately extract mobility functioning information from clinical text. The public availability of our annotated dataset will facilitate further research to comprehensively capture functioning information in electronic health records (EHRs).
Rebuild City Buildings from Off-Nadir Aerial Images with Offset-Building Model (OBM)
Oct 25, 2023Accurate measurement of the offset from roof-to-footprint in very-high-resolution remote sensing imagery is crucial for urban information extraction tasks. With the help of deep learning, existing methods typically rely on two-stage CNN models to extract regions of interest on building feature maps. At the first stage, a Region Proposal Network (RPN) is applied to extract thousands of ROIs (Region of Interests) which will post-imported into a Region-based Convolutional Neural Networks (RCNN) to extract wanted information. However, because of inflexible RPN, these methods often lack effective user interaction, encounter difficulties in instance correspondence, and struggle to keep up with the advancements in general artificial intelligence. This paper introduces an interactive Transformer model combined with a prompt encoder to precisely extract building segmentation as well as the offset vectors from roofs to footprints. In our model, a powerful module, namely ROAM, was tailored for common problems in predicting roof-to-footprint offsets. We tested our model's feasibility on the publicly available BONAI dataset, achieving a significant reduction in Prompt-Instance-Level offset errors ranging from 14.6% to 16.3%. Additionally, we developed a Distance-NMS algorithm tailored for large-scale building offsets, significantly enhancing the accuracy of predicted building offset angles and lengths in a straightforward and efficient manner. To further validate the model's robustness, we created a new test set using 0.5m remote sensing imagery from Huizhou, China, for inference testing. Our code, training methods, and the updated dataset will be accessable at https://github.com/likaiucas.
End-to-End Document Classification and Key Information Extraction using Assignment Optimization
Jun 01, 2023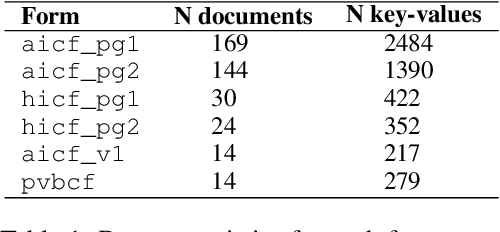
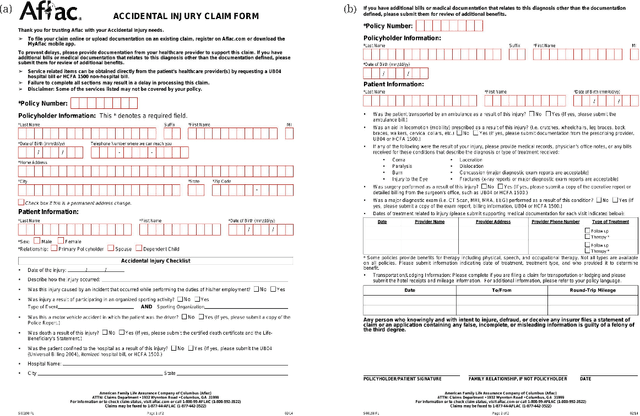

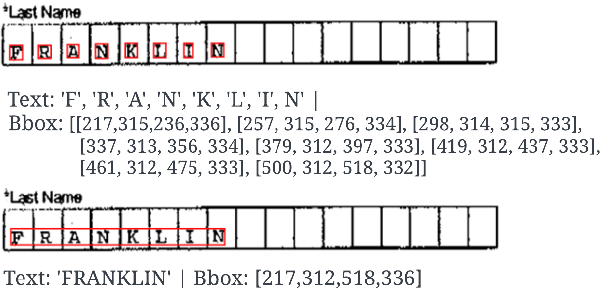
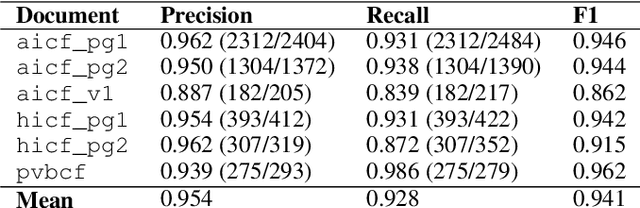
We propose end-to-end document classification and key information extraction (KIE) for automating document processing in forms. Through accurate document classification we harness known information from templates to enhance KIE from forms. We use text and layout encoding with a cosine similarity measure to classify visually-similar documents. We then demonstrate a novel application of mixed integer programming by using assignment optimization to extract key information from documents. Our approach is validated on an in-house dataset of noisy scanned forms. The best performing document classification approach achieved 0.97 f1 score. A mean f1 score of 0.94 for the KIE task suggests there is significant potential in applying optimization techniques. Abation results show that the method relies on document preprocessing techniques to mitigate Type II errors and achieve optimal performance.
Extracting individual variable information for their decoupling, direct mutual information and multi-feature Granger causality
Nov 22, 2023Working with multiple variables they usually contain difficult to control complex dependencies. This article proposes extraction of their individual information, e.g. $\overline{X|Y}$ as random variable containing information from $X$, but with removed information about $Y$, by using $(x,y) \leftrightarrow (\bar{x}=\textrm{CDF}_{X|Y=y}(x),y)$ reversible normalization. One application can be decoupling of individual information of variables: reversibly transform $(X_1,\ldots,X_n)\leftrightarrow(\tilde{X}_1,\ldots \tilde{X}_n)$ together containing the same information, but being independent: $\forall_{i\neq j} \tilde{X}_i\perp \tilde{X}_j, \tilde{X}_i\perp X_j$. It requires detailed models of complex conditional probability distributions - it is generally a difficult task, but here can be done through multiple dependency reducing iterations, using imperfect methods (here HCR: Hierarchical Correlation Reconstruction). It could be also used for direct mutual information - evaluating direct information transfer: without use of intermediate variables. For causality direction there is discussed multi-feature Granger causality, e.g. to trace various types of individual information transfers between such decoupled variables, including propagation time (delay).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge