"Information Extraction": models, code, and papers
Applications of Sequential Learning for Medical Image Classification
Sep 26, 2023Purpose: The aim of this work is to develop a neural network training framework for continual training of small amounts of medical imaging data and create heuristics to assess training in the absence of a hold-out validation or test set. Materials and Methods: We formulated a retrospective sequential learning approach that would train and consistently update a model on mini-batches of medical images over time. We address problems that impede sequential learning such as overfitting, catastrophic forgetting, and concept drift through PyTorch convolutional neural networks (CNN) and publicly available Medical MNIST and NIH Chest X-Ray imaging datasets. We begin by comparing two methods for a sequentially trained CNN with and without base pre-training. We then transition to two methods of unique training and validation data recruitment to estimate full information extraction without overfitting. Lastly, we consider an example of real-life data that shows how our approach would see mainstream research implementation. Results: For the first experiment, both approaches successfully reach a ~95% accuracy threshold, although the short pre-training step enables sequential accuracy to plateau in fewer steps. The second experiment comparing two methods showed better performance with the second method which crosses the ~90% accuracy threshold much sooner. The final experiment showed a slight advantage with a pre-training step that allows the CNN to cross ~60% threshold much sooner than without pre-training. Conclusion: We have displayed sequential learning as a serviceable multi-classification technique statistically comparable to traditional CNNs that can acquire data in small increments feasible for clinically realistic scenarios.
Informed Named Entity Recognition Decoding for Generative Language Models
Aug 15, 2023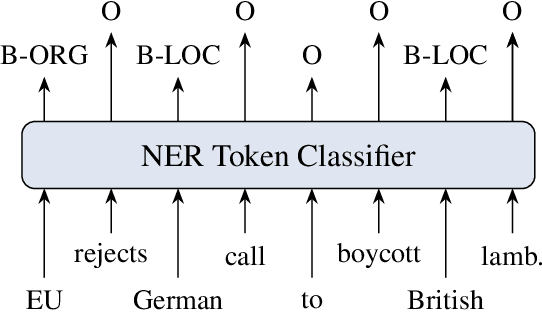

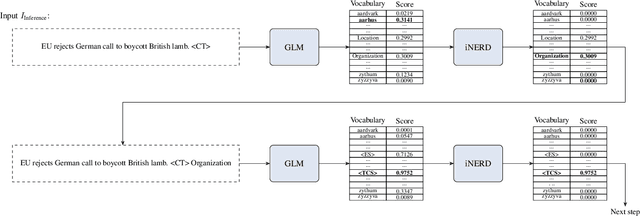
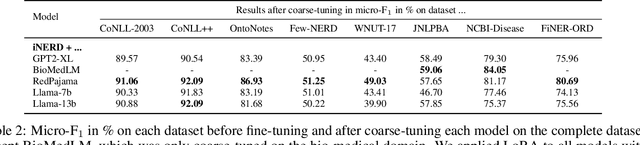
Ever-larger language models with ever-increasing capabilities are by now well-established text processing tools. Alas, information extraction tasks such as named entity recognition are still largely unaffected by this progress as they are primarily based on the previous generation of encoder-only transformer models. Here, we propose a simple yet effective approach, Informed Named Entity Recognition Decoding (iNERD), which treats named entity recognition as a generative process. It leverages the language understanding capabilities of recent generative models in a future-proof manner and employs an informed decoding scheme incorporating the restricted nature of information extraction into open-ended text generation, improving performance and eliminating any risk of hallucinations. We coarse-tune our model on a merged named entity corpus to strengthen its performance, evaluate five generative language models on eight named entity recognition datasets, and achieve remarkable results, especially in an environment with an unknown entity class set, demonstrating the adaptability of the approach.
A Knowledge-enhanced Two-stage Generative Framework for Medical Dialogue Information Extraction
Jul 30, 2023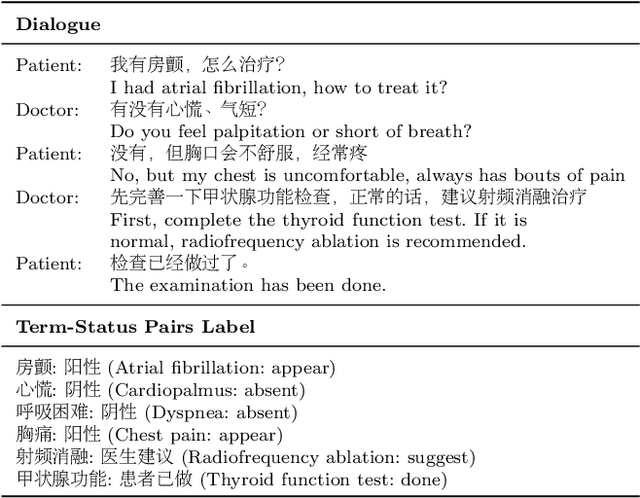
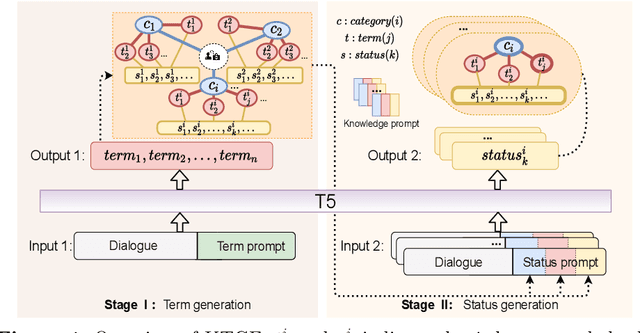
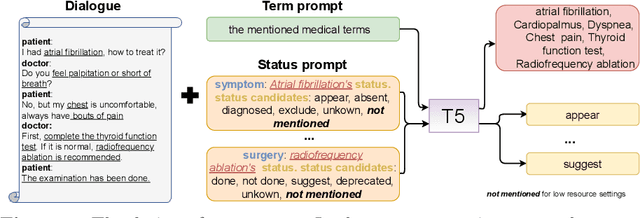
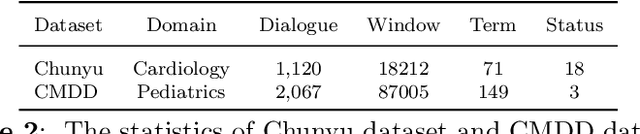
This paper focuses on term-status pair extraction from medical dialogues (MD-TSPE), which is essential in diagnosis dialogue systems and the automatic scribe of electronic medical records (EMRs). In the past few years, works on MD-TSPE have attracted increasing research attention, especially after the remarkable progress made by generative methods. However, these generative methods output a whole sequence consisting of term-status pairs in one stage and ignore integrating prior knowledge, which demands a deeper understanding to model the relationship between terms and infer the status of each term. This paper presents a knowledge-enhanced two-stage generative framework (KTGF) to address the above challenges. Using task-specific prompts, we employ a single model to complete the MD-TSPE through two phases in a unified generative form: we generate all terms the first and then generate the status of each generated term. In this way, the relationship between terms can be learned more effectively from the sequence containing only terms in the first phase, and our designed knowledge-enhanced prompt in the second phase can leverage the category and status candidates of the generated term for status generation. Furthermore, our proposed special status ``not mentioned" makes more terms available and enriches the training data in the second phase, which is critical in the low-resource setting. The experiments on the Chunyu and CMDD datasets show that the proposed method achieves superior results compared to the state-of-the-art models in the full training and low-resource settings.
OceanBench: The Sea Surface Height Edition
Sep 27, 2023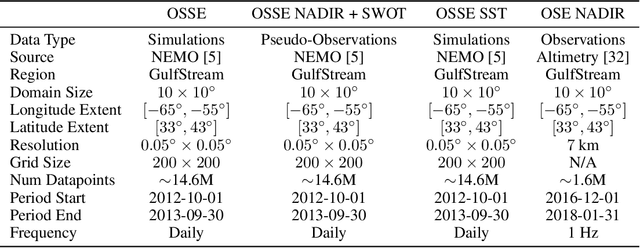
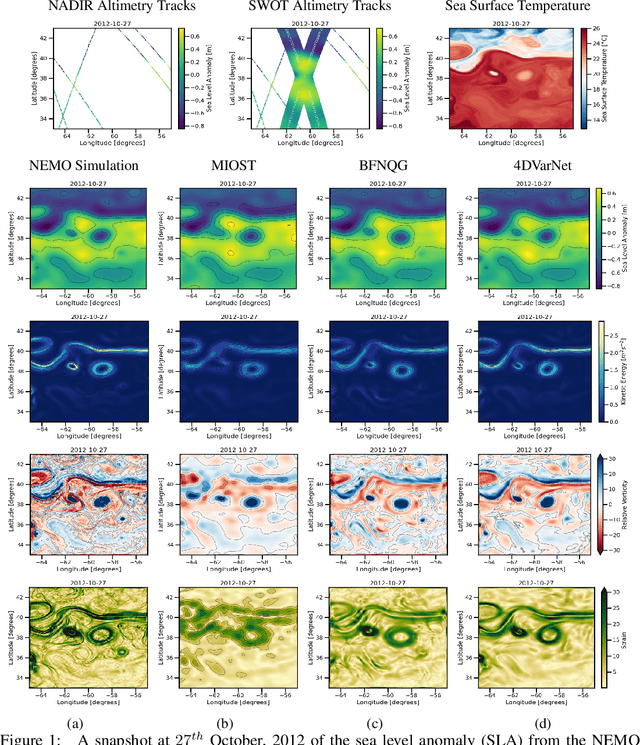
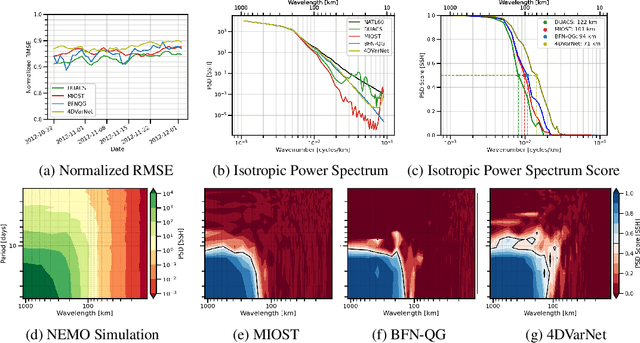

The ocean profoundly influences human activities and plays a critical role in climate regulation. Our understanding has improved over the last decades with the advent of satellite remote sensing data, allowing us to capture essential quantities over the globe, e.g., sea surface height (SSH). However, ocean satellite data presents challenges for information extraction due to their sparsity and irregular sampling, signal complexity, and noise. Machine learning (ML) techniques have demonstrated their capabilities in dealing with large-scale, complex signals. Therefore we see an opportunity for ML models to harness the information contained in ocean satellite data. However, data representation and relevant evaluation metrics can be the defining factors when determining the success of applied ML. The processing steps from the raw observation data to a ML-ready state and from model outputs to interpretable quantities require domain expertise, which can be a significant barrier to entry for ML researchers. OceanBench is a unifying framework that provides standardized processing steps that comply with domain-expert standards. It provides plug-and-play data and pre-configured pipelines for ML researchers to benchmark their models and a transparent configurable framework for researchers to customize and extend the pipeline for their tasks. In this work, we demonstrate the OceanBench framework through a first edition dedicated to SSH interpolation challenges. We provide datasets and ML-ready benchmarking pipelines for the long-standing problem of interpolating observations from simulated ocean satellite data, multi-modal and multi-sensor fusion issues, and transfer-learning to real ocean satellite observations. The OceanBench framework is available at github.com/jejjohnson/oceanbench and the dataset registry is available at github.com/quentinf00/oceanbench-data-registry.
Black-Box Analysis: GPTs Across Time in Legal Textual Entailment Task
Sep 11, 2023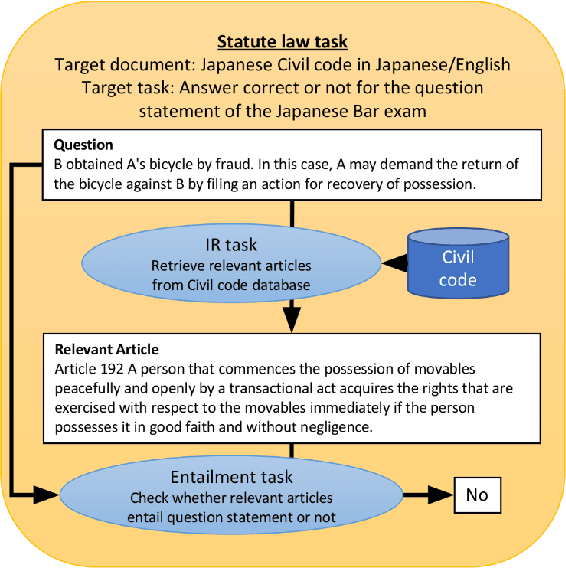
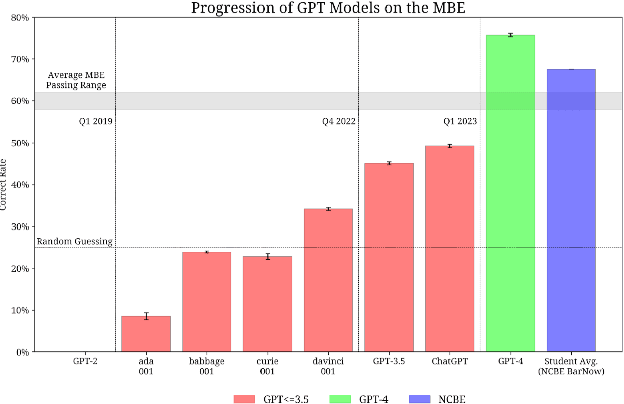
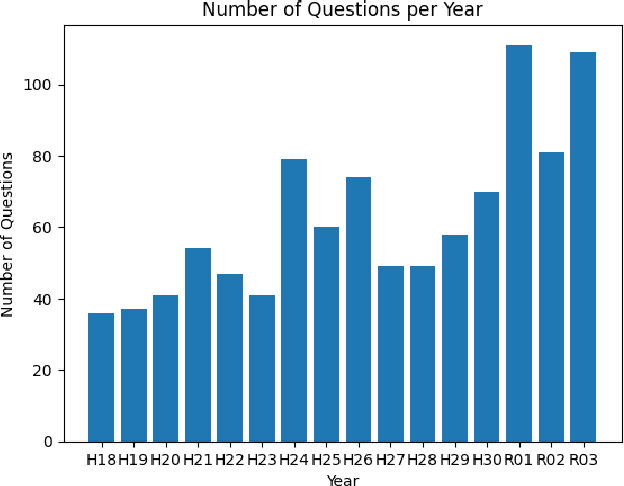
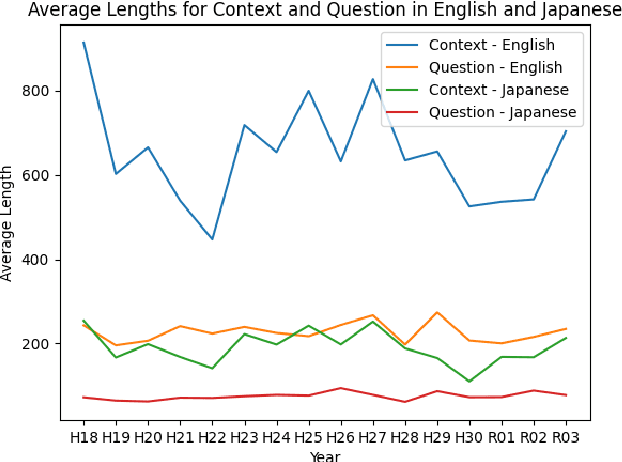
The evolution of Generative Pre-trained Transformer (GPT) models has led to significant advancements in various natural language processing applications, particularly in legal textual entailment. We present an analysis of GPT-3.5 (ChatGPT) and GPT-4 performances on COLIEE Task 4 dataset, a prominent benchmark in this domain. The study encompasses data from Heisei 18 (2006) to Reiwa 3 (2021), exploring the models' abilities to discern entailment relationships within Japanese statute law across different periods. Our preliminary experimental results unveil intriguing insights into the models' strengths and weaknesses in handling legal textual entailment tasks, as well as the patterns observed in model performance. In the context of proprietary models with undisclosed architectures and weights, black-box analysis becomes crucial for evaluating their capabilities. We discuss the influence of training data distribution and the implications on the models' generalizability. This analysis serves as a foundation for future research, aiming to optimize GPT-based models and enable their successful adoption in legal information extraction and entailment applications.
Copyright Violations and Large Language Models
Oct 20, 2023Language models may memorize more than just facts, including entire chunks of texts seen during training. Fair use exemptions to copyright laws typically allow for limited use of copyrighted material without permission from the copyright holder, but typically for extraction of information from copyrighted materials, rather than {\em verbatim} reproduction. This work explores the issue of copyright violations and large language models through the lens of verbatim memorization, focusing on possible redistribution of copyrighted text. We present experiments with a range of language models over a collection of popular books and coding problems, providing a conservative characterization of the extent to which language models can redistribute these materials. Overall, this research highlights the need for further examination and the potential impact on future developments in natural language processing to ensure adherence to copyright regulations. Code is at \url{https://github.com/coastalcph/CopyrightLLMs}.
Towards Safer Operations: An Expert-involved Dataset of High-Pressure Gas Incidents for Preventing Future Failures
Oct 18, 2023This paper introduces a new IncidentAI dataset for safety prevention. Different from prior corpora that usually contain a single task, our dataset comprises three tasks: named entity recognition, cause-effect extraction, and information retrieval. The dataset is annotated by domain experts who have at least six years of practical experience as high-pressure gas conservation managers. We validate the contribution of the dataset in the scenario of safety prevention. Preliminary results on the three tasks show that NLP techniques are beneficial for analyzing incident reports to prevent future failures. The dataset facilitates future research in NLP and incident management communities. The access to the dataset is also provided (the IncidentAI dataset is available at: https://github.com/Cinnamon/incident-ai-dataset).
Optimising Human-Machine Collaboration for Efficient High-Precision Information Extraction from Text Documents
Feb 18, 2023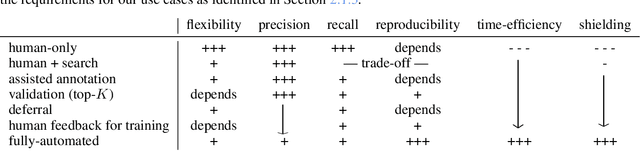
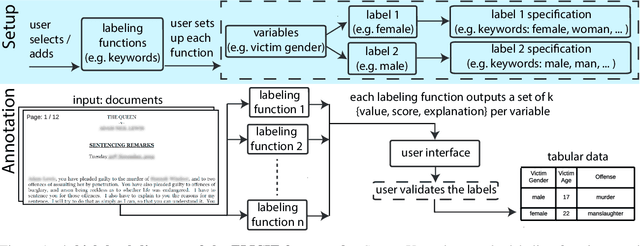
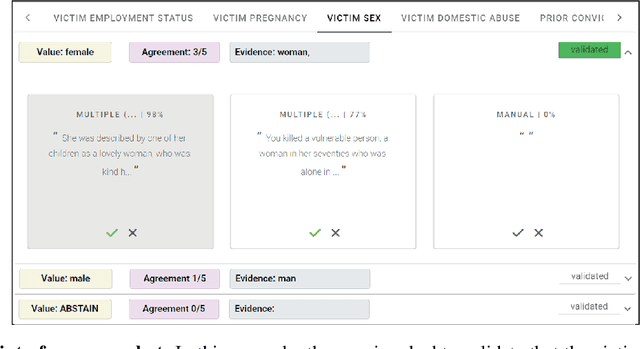
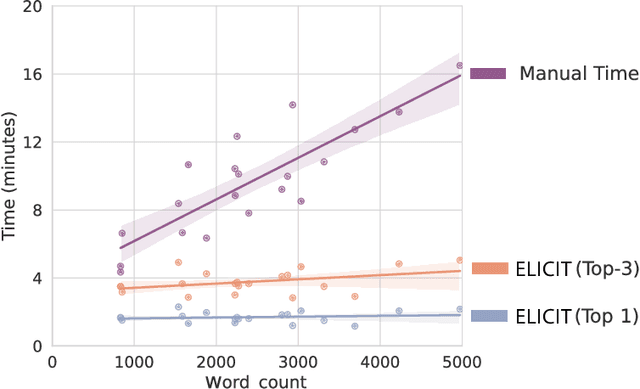
While humans can extract information from unstructured text with high precision and recall, this is often too time-consuming to be practical. Automated approaches, on the other hand, produce nearly-immediate results, but may not be reliable enough for high-stakes applications where precision is essential. In this work, we consider the benefits and drawbacks of various human-only, human-machine, and machine-only information extraction approaches. We argue for the utility of a human-in-the-loop approach in applications where high precision is required, but purely manual extraction is infeasible. We present a framework and an accompanying tool for information extraction using weak-supervision labelling with human validation. We demonstrate our approach on three criminal justice datasets. We find that the combination of computer speed and human understanding yields precision comparable to manual annotation while requiring only a fraction of time, and significantly outperforms fully automated baselines in terms of precision.
Zero-Shot Information Extraction via Chatting with ChatGPT
Feb 20, 2023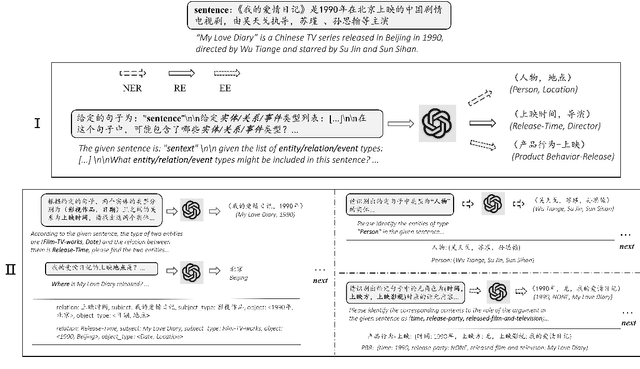
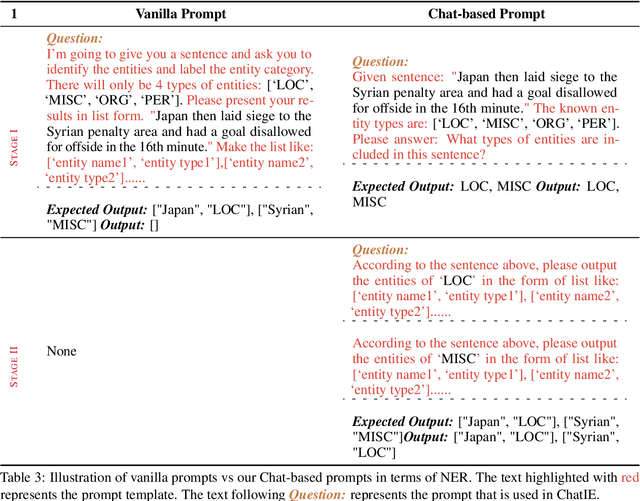
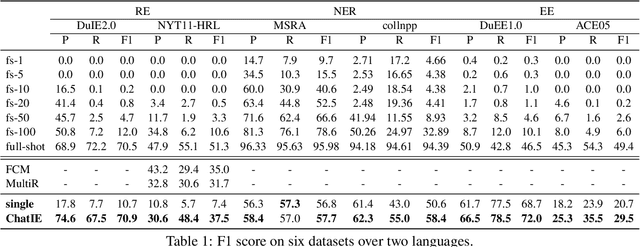


Zero-shot information extraction (IE) aims to build IE systems from the unannotated text. It is challenging due to involving little human intervention. Challenging but worthwhile, zero-shot IE reduces the time and effort that data labeling takes. Recent efforts on large language models (LLMs, e.g., GPT-3, ChatGPT) show promising performance on zero-shot settings, thus inspiring us to explore prompt-based methods. In this work, we ask whether strong IE models can be constructed by directly prompting LLMs. Specifically, we transform the zero-shot IE task into a multi-turn question-answering problem with a two-stage framework (ChatIE). With the power of ChatGPT, we extensively evaluate our framework on three IE tasks: entity-relation triple extract, named entity recognition, and event extraction. Empirical results on six datasets across two languages show that ChatIE achieves impressive performance and even surpasses some full-shot models on several datasets (e.g., NYT11-HRL). We believe that our work could shed light on building IE models with limited resources.
LasUIE: Unifying Information Extraction with Latent Adaptive Structure-aware Generative Language Model
Apr 13, 2023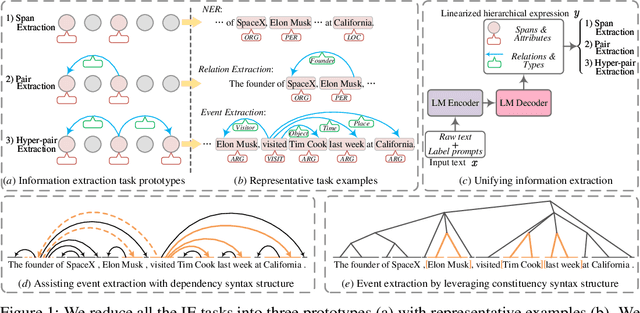
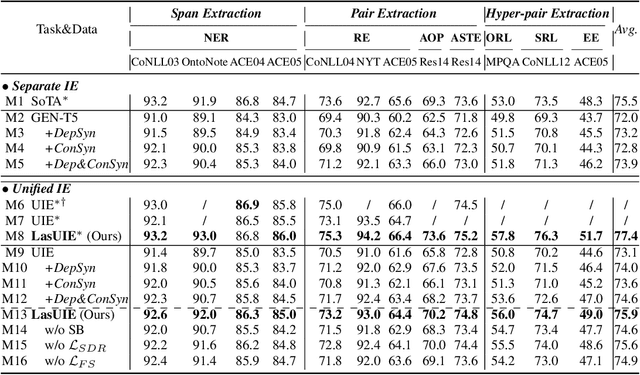
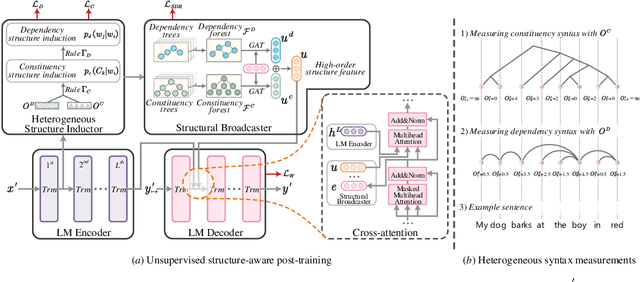
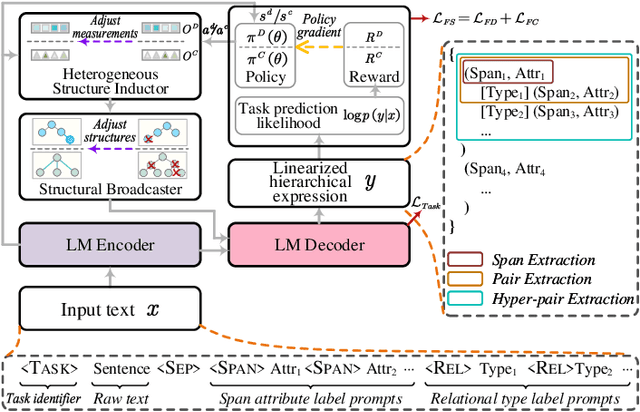
Universally modeling all typical information extraction tasks (UIE) with one generative language model (GLM) has revealed great potential by the latest study, where various IE predictions are unified into a linearized hierarchical expression under a GLM. Syntactic structure information, a type of effective feature which has been extensively utilized in IE community, should also be beneficial to UIE. In this work, we propose a novel structure-aware GLM, fully unleashing the power of syntactic knowledge for UIE. A heterogeneous structure inductor is explored to unsupervisedly induce rich heterogeneous structural representations by post-training an existing GLM. In particular, a structural broadcaster is devised to compact various latent trees into explicit high-order forests, helping to guide a better generation during decoding. We finally introduce a task-oriented structure fine-tuning mechanism, further adjusting the learned structures to most coincide with the end-task's need. Over 12 IE benchmarks across 7 tasks our system shows significant improvements over the baseline UIE system. Further in-depth analyses show that our GLM learns rich task-adaptive structural bias that greatly resolves the UIE crux, the long-range dependence issue and boundary identifying. Source codes are open at https://github.com/ChocoWu/LasUIE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge