"Information Extraction": models, code, and papers
LLMs as Counterfactual Explanation Modules: Can ChatGPT Explain Black-box Text Classifiers?
Sep 23, 2023Large language models (LLMs) are increasingly being used for tasks beyond text generation, including complex tasks such as data labeling, information extraction, etc. With the recent surge in research efforts to comprehend the full extent of LLM capabilities, in this work, we investigate the role of LLMs as counterfactual explanation modules, to explain decisions of black-box text classifiers. Inspired by causal thinking, we propose a pipeline for using LLMs to generate post-hoc, model-agnostic counterfactual explanations in a principled way via (i) leveraging the textual understanding capabilities of the LLM to identify and extract latent features, and (ii) leveraging the perturbation and generation capabilities of the same LLM to generate a counterfactual explanation by perturbing input features derived from the extracted latent features. We evaluate three variants of our framework, with varying degrees of specificity, on a suite of state-of-the-art LLMs, including ChatGPT and LLaMA 2. We evaluate the effectiveness and quality of the generated counterfactual explanations, over a variety of text classification benchmarks. Our results show varied performance of these models in different settings, with a full two-step feature extraction based variant outperforming others in most cases. Our pipeline can be used in automated explanation systems, potentially reducing human effort.
RexUIE: A Recursive Method with Explicit Schema Instructor for Universal Information Extraction
Apr 28, 2023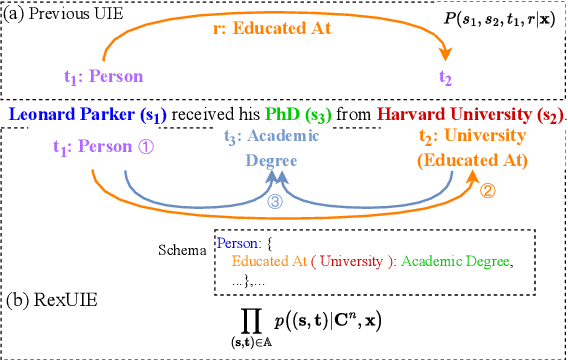
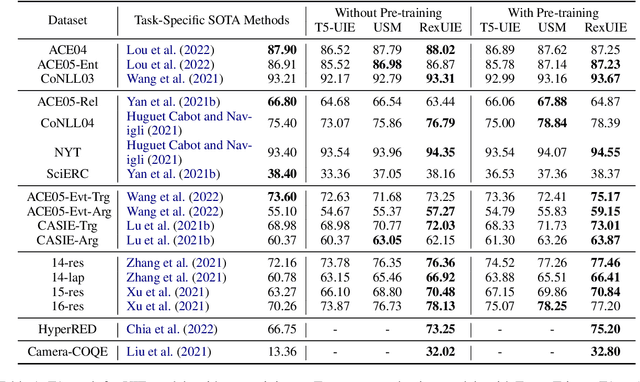
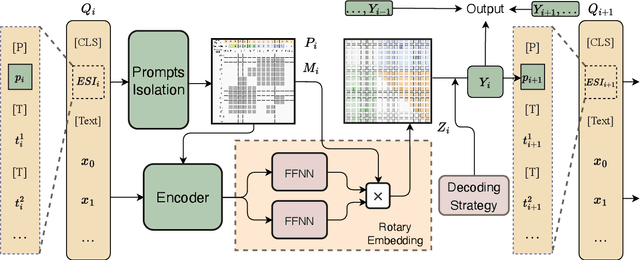
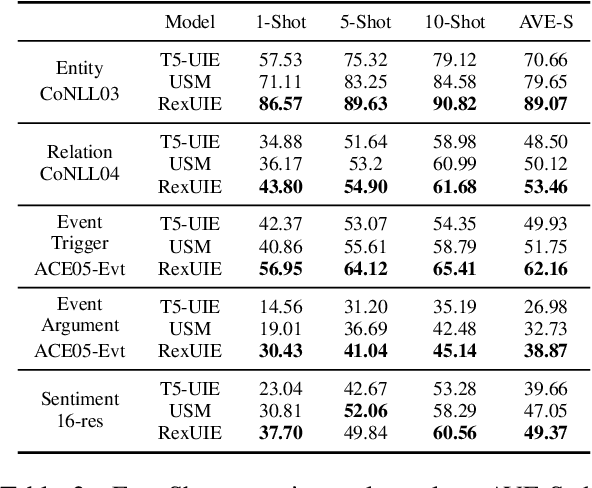
Universal Information Extraction (UIE) is an area of interest due to the challenges posed by varying targets, heterogeneous structures, and demand-specific schemas. However, previous works have only achieved limited success by unifying a few tasks, such as Named Entity Recognition (NER) and Relation Extraction (RE), which fall short of being authentic UIE models particularly when extracting other general schemas such as quadruples and quintuples. Additionally, these models used an implicit structural schema instructor, which could lead to incorrect links between types, hindering the model's generalization and performance in low-resource scenarios. In this paper, we redefine the authentic UIE with a formal formulation that encompasses almost all extraction schemas. To the best of our knowledge, we are the first to introduce UIE for any kind of schemas. In addition, we propose RexUIE, which is a Recursive Method with Explicit Schema Instructor for UIE. To avoid interference between different types, we reset the position ids and attention mask matrices. RexUIE shows strong performance under both full-shot and few-shot settings and achieves State-of-the-Art results on the tasks of extracting complex schemas.
Taking a PEEK into YOLOv5 for Satellite Component Recognition via Entropy-based Visual Explanations
Nov 03, 2023The escalating risk of collisions and the accumulation of space debris in Low Earth Orbit (LEO) has reached critical concern due to the ever increasing number of spacecraft. Addressing this crisis, especially in dealing with non-cooperative and unidentified space debris, is of paramount importance. This paper contributes to efforts in enabling autonomous swarms of small chaser satellites for target geometry determination and safe flight trajectory planning for proximity operations in LEO. Our research explores on-orbit use of the You Only Look Once v5 (YOLOv5) object detection model trained to detect satellite components. While this model has shown promise, its inherent lack of interpretability hinders human understanding, a critical aspect of validating algorithms for use in safety-critical missions. To analyze the decision processes, we introduce Probabilistic Explanations for Entropic Knowledge extraction (PEEK), a method that utilizes information theoretic analysis of the latent representations within the hidden layers of the model. Through both synthetic in hardware-in-the-loop experiments, PEEK illuminates the decision-making processes of the model, helping identify its strengths, limitations and biases.
Teamwork Is Not Always Good: An Empirical Study of Classifier Drift in Class-incremental Information Extraction
May 26, 2023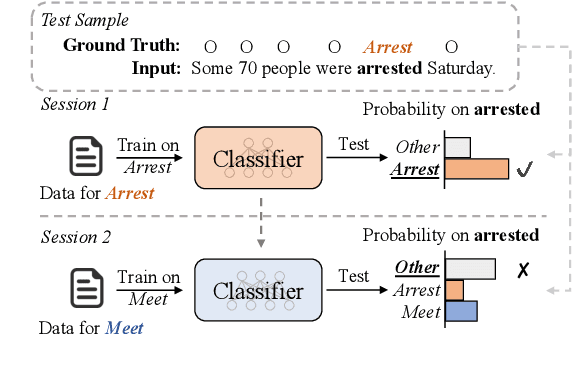
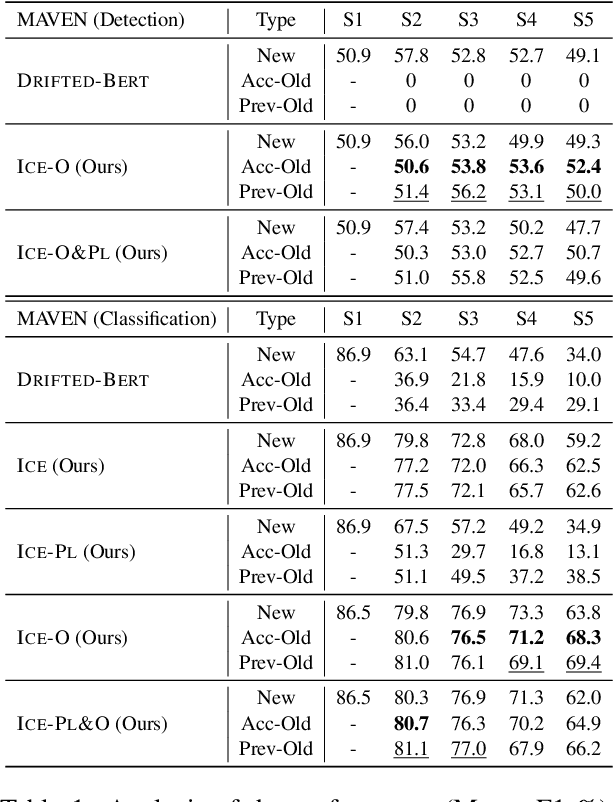
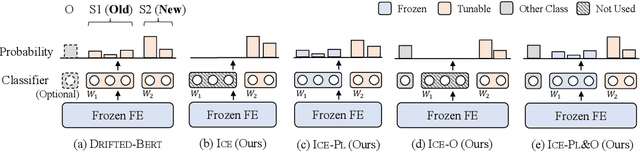
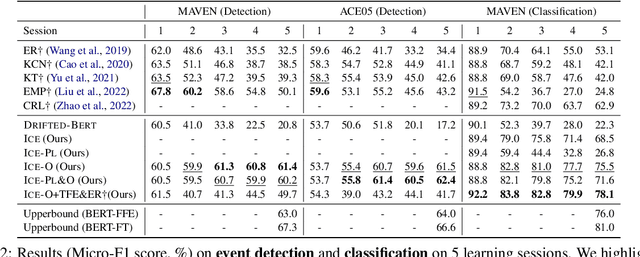
Class-incremental learning (CIL) aims to develop a learning system that can continually learn new classes from a data stream without forgetting previously learned classes. When learning classes incrementally, the classifier must be constantly updated to incorporate new classes, and the drift in decision boundary may lead to severe forgetting. This fundamental challenge, however, has not yet been studied extensively, especially in the setting where no samples from old classes are stored for rehearsal. In this paper, we take a closer look at how the drift in the classifier leads to forgetting, and accordingly, design four simple yet (super-) effective solutions to alleviate the classifier drift: an Individual Classifiers with Frozen Feature Extractor (ICE) framework where we individually train a classifier for each learning session, and its three variants ICE-PL, ICE-O, and ICE-PL&O which further take the logits of previously learned classes from old sessions or a constant logit of an Other class as a constraint to the learning of new classifiers. Extensive experiments and analysis on 6 class-incremental information extraction tasks demonstrate that our solutions, especially ICE-O, consistently show significant improvement over the previous state-of-the-art approaches with up to 44.7% absolute F-score gain, providing a strong baseline and insights for future research on class-incremental learning.
Applications of Sequential Learning for Medical Image Classification
Sep 26, 2023Purpose: The aim of this work is to develop a neural network training framework for continual training of small amounts of medical imaging data and create heuristics to assess training in the absence of a hold-out validation or test set. Materials and Methods: We formulated a retrospective sequential learning approach that would train and consistently update a model on mini-batches of medical images over time. We address problems that impede sequential learning such as overfitting, catastrophic forgetting, and concept drift through PyTorch convolutional neural networks (CNN) and publicly available Medical MNIST and NIH Chest X-Ray imaging datasets. We begin by comparing two methods for a sequentially trained CNN with and without base pre-training. We then transition to two methods of unique training and validation data recruitment to estimate full information extraction without overfitting. Lastly, we consider an example of real-life data that shows how our approach would see mainstream research implementation. Results: For the first experiment, both approaches successfully reach a ~95% accuracy threshold, although the short pre-training step enables sequential accuracy to plateau in fewer steps. The second experiment comparing two methods showed better performance with the second method which crosses the ~90% accuracy threshold much sooner. The final experiment showed a slight advantage with a pre-training step that allows the CNN to cross ~60% threshold much sooner than without pre-training. Conclusion: We have displayed sequential learning as a serviceable multi-classification technique statistically comparable to traditional CNNs that can acquire data in small increments feasible for clinically realistic scenarios.
StructChart: Perception, Structuring, Reasoning for Visual Chart Understanding
Sep 20, 2023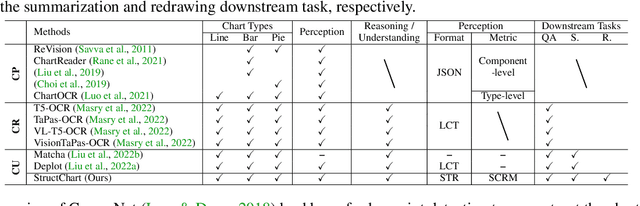
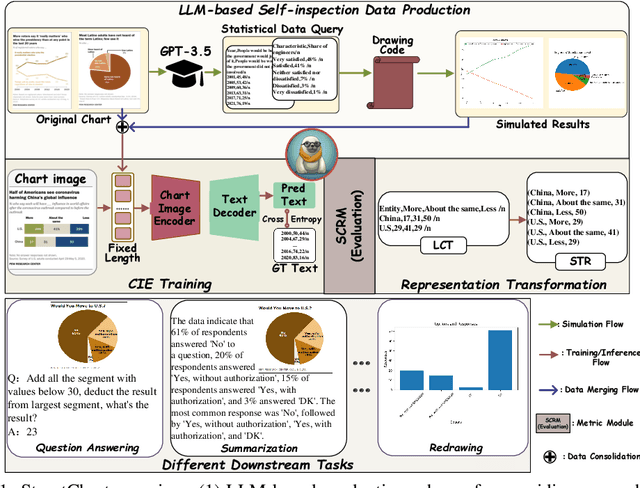
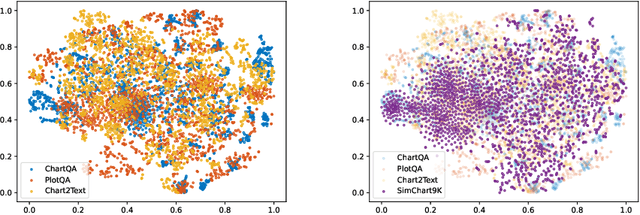
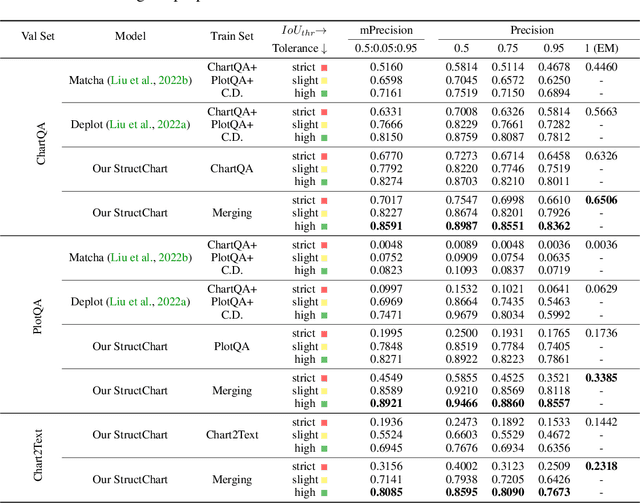
Charts are common in literature across different scientific fields, conveying rich information easily accessible to readers. Current chart-related tasks focus on either chart perception which refers to extracting information from the visual charts, or performing reasoning given the extracted data, e.g. in a tabular form. In this paper, we aim to establish a unified and label-efficient learning paradigm for joint perception and reasoning tasks, which can be generally applicable to different downstream tasks, beyond the question-answering task as specifically studied in peer works. Specifically, StructChart first reformulates the chart information from the popular tubular form (specifically linearized CSV) to the proposed Structured Triplet Representations (STR), which is more friendly for reducing the task gap between chart perception and reasoning due to the employed structured information extraction for charts. We then propose a Structuring Chart-oriented Representation Metric (SCRM) to quantitatively evaluate the performance for the chart perception task. To enrich the dataset for training, we further explore the possibility of leveraging the Large Language Model (LLM), enhancing the chart diversity in terms of both chart visual style and its statistical information. Extensive experiments are conducted on various chart-related tasks, demonstrating the effectiveness and promising potential for a unified chart perception-reasoning paradigm to push the frontier of chart understanding.
OceanBench: The Sea Surface Height Edition
Sep 27, 2023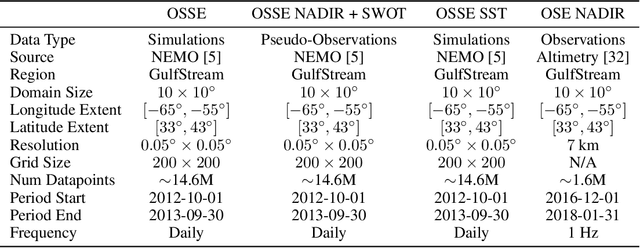
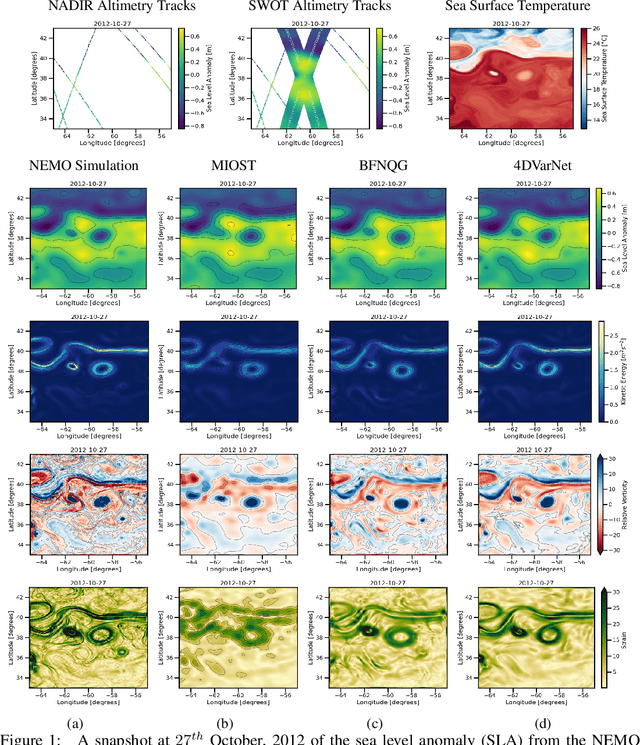
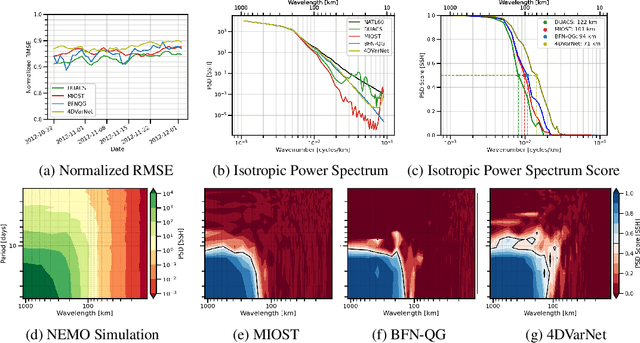

The ocean profoundly influences human activities and plays a critical role in climate regulation. Our understanding has improved over the last decades with the advent of satellite remote sensing data, allowing us to capture essential quantities over the globe, e.g., sea surface height (SSH). However, ocean satellite data presents challenges for information extraction due to their sparsity and irregular sampling, signal complexity, and noise. Machine learning (ML) techniques have demonstrated their capabilities in dealing with large-scale, complex signals. Therefore we see an opportunity for ML models to harness the information contained in ocean satellite data. However, data representation and relevant evaluation metrics can be the defining factors when determining the success of applied ML. The processing steps from the raw observation data to a ML-ready state and from model outputs to interpretable quantities require domain expertise, which can be a significant barrier to entry for ML researchers. OceanBench is a unifying framework that provides standardized processing steps that comply with domain-expert standards. It provides plug-and-play data and pre-configured pipelines for ML researchers to benchmark their models and a transparent configurable framework for researchers to customize and extend the pipeline for their tasks. In this work, we demonstrate the OceanBench framework through a first edition dedicated to SSH interpolation challenges. We provide datasets and ML-ready benchmarking pipelines for the long-standing problem of interpolating observations from simulated ocean satellite data, multi-modal and multi-sensor fusion issues, and transfer-learning to real ocean satellite observations. The OceanBench framework is available at github.com/jejjohnson/oceanbench and the dataset registry is available at github.com/quentinf00/oceanbench-data-registry.
Copyright Violations and Large Language Models
Oct 20, 2023Language models may memorize more than just facts, including entire chunks of texts seen during training. Fair use exemptions to copyright laws typically allow for limited use of copyrighted material without permission from the copyright holder, but typically for extraction of information from copyrighted materials, rather than {\em verbatim} reproduction. This work explores the issue of copyright violations and large language models through the lens of verbatim memorization, focusing on possible redistribution of copyrighted text. We present experiments with a range of language models over a collection of popular books and coding problems, providing a conservative characterization of the extent to which language models can redistribute these materials. Overall, this research highlights the need for further examination and the potential impact on future developments in natural language processing to ensure adherence to copyright regulations. Code is at \url{https://github.com/coastalcph/CopyrightLLMs}.
Towards Safer Operations: An Expert-involved Dataset of High-Pressure Gas Incidents for Preventing Future Failures
Oct 18, 2023This paper introduces a new IncidentAI dataset for safety prevention. Different from prior corpora that usually contain a single task, our dataset comprises three tasks: named entity recognition, cause-effect extraction, and information retrieval. The dataset is annotated by domain experts who have at least six years of practical experience as high-pressure gas conservation managers. We validate the contribution of the dataset in the scenario of safety prevention. Preliminary results on the three tasks show that NLP techniques are beneficial for analyzing incident reports to prevent future failures. The dataset facilitates future research in NLP and incident management communities. The access to the dataset is also provided (the IncidentAI dataset is available at: https://github.com/Cinnamon/incident-ai-dataset).
Evaluating ChatGPT's Information Extraction Capabilities: An Assessment of Performance, Explainability, Calibration, and Faithfulness
Apr 23, 2023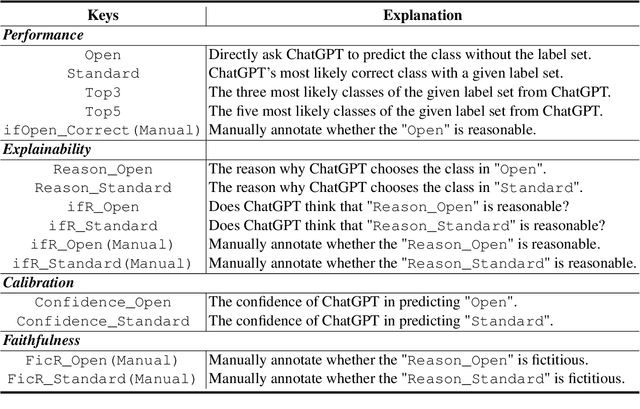
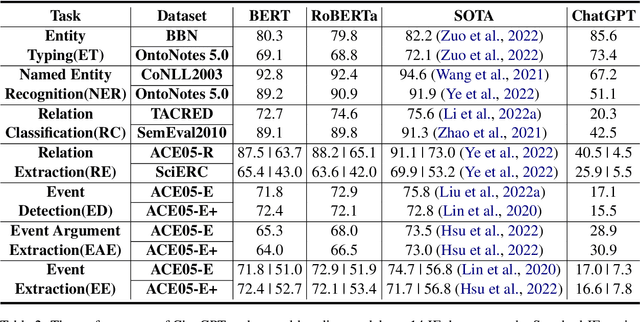
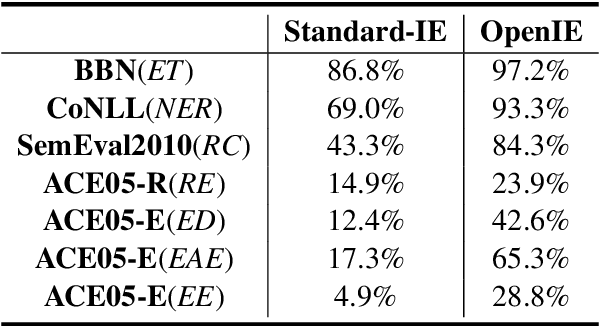
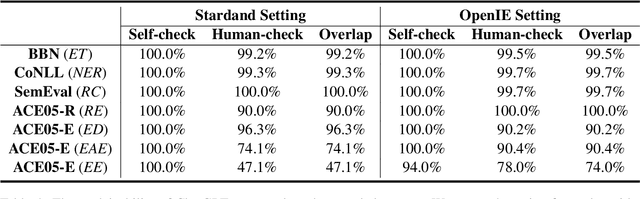
The capability of Large Language Models (LLMs) like ChatGPT to comprehend user intent and provide reasonable responses has made them extremely popular lately. In this paper, we focus on assessing the overall ability of ChatGPT using 7 fine-grained information extraction (IE) tasks. Specially, we present the systematically analysis by measuring ChatGPT's performance, explainability, calibration, and faithfulness, and resulting in 15 keys from either the ChatGPT or domain experts. Our findings reveal that ChatGPT's performance in Standard-IE setting is poor, but it surprisingly exhibits excellent performance in the OpenIE setting, as evidenced by human evaluation. In addition, our research indicates that ChatGPT provides high-quality and trustworthy explanations for its decisions. However, there is an issue of ChatGPT being overconfident in its predictions, which resulting in low calibration. Furthermore, ChatGPT demonstrates a high level of faithfulness to the original text in the majority of cases. We manually annotate and release the test sets of 7 fine-grained IE tasks contains 14 datasets to further promote the research. The datasets and code are available at https://github.com/pkuserc/ChatGPT_for_IE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge