"Information Extraction": models, code, and papers
Modeling Entities as Semantic Points for Visual Information Extraction in the Wild
Mar 29, 2023
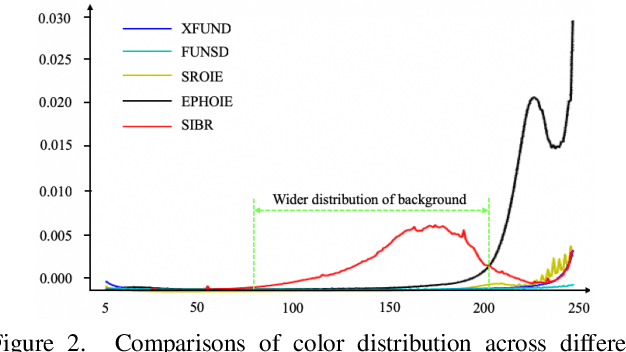
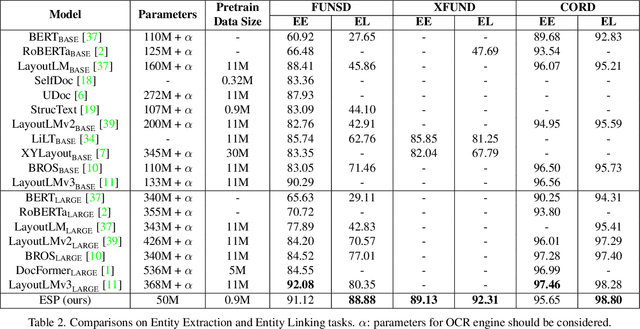
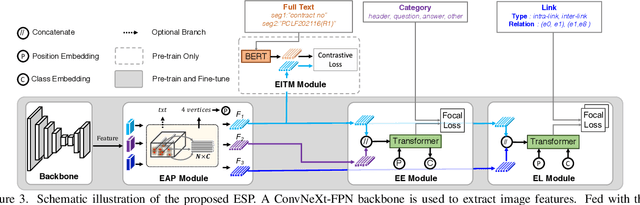
Recently, Visual Information Extraction (VIE) has been becoming increasingly important in both the academia and industry, due to the wide range of real-world applications. Previously, numerous works have been proposed to tackle this problem. However, the benchmarks used to assess these methods are relatively plain, i.e., scenarios with real-world complexity are not fully represented in these benchmarks. As the first contribution of this work, we curate and release a new dataset for VIE, in which the document images are much more challenging in that they are taken from real applications, and difficulties such as blur, partial occlusion, and printing shift are quite common. All these factors may lead to failures in information extraction. Therefore, as the second contribution, we explore an alternative approach to precisely and robustly extract key information from document images under such tough conditions. Specifically, in contrast to previous methods, which usually either incorporate visual information into a multi-modal architecture or train text spotting and information extraction in an end-to-end fashion, we explicitly model entities as semantic points, i.e., center points of entities are enriched with semantic information describing the attributes and relationships of different entities, which could largely benefit entity labeling and linking. Extensive experiments on standard benchmarks in this field as well as the proposed dataset demonstrate that the proposed method can achieve significantly enhanced performance on entity labeling and linking, compared with previous state-of-the-art models. Dataset is available at https://www.modelscope.cn/datasets/damo/SIBR/summary.
From Large Language Models to Knowledge Graphs for Biomarker Discovery in Cancer
Oct 12, 2023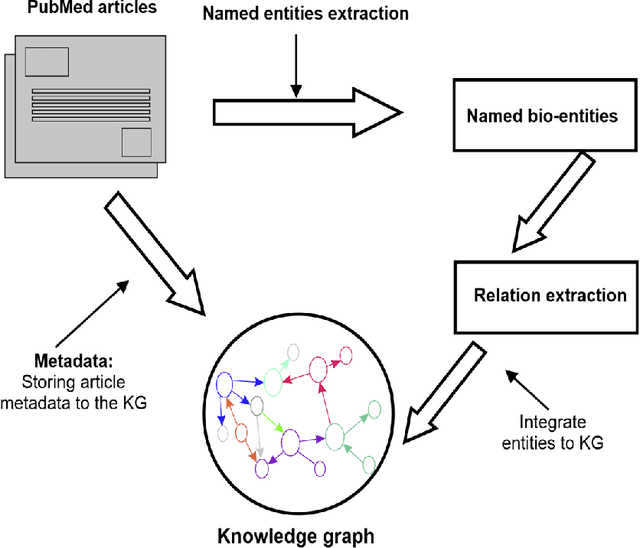
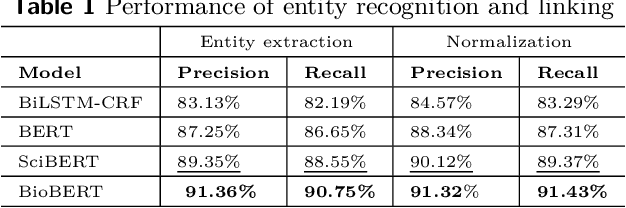
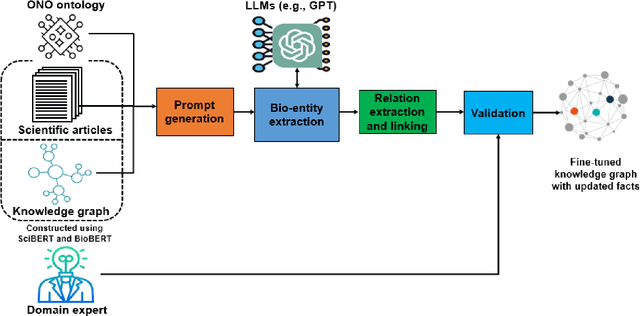
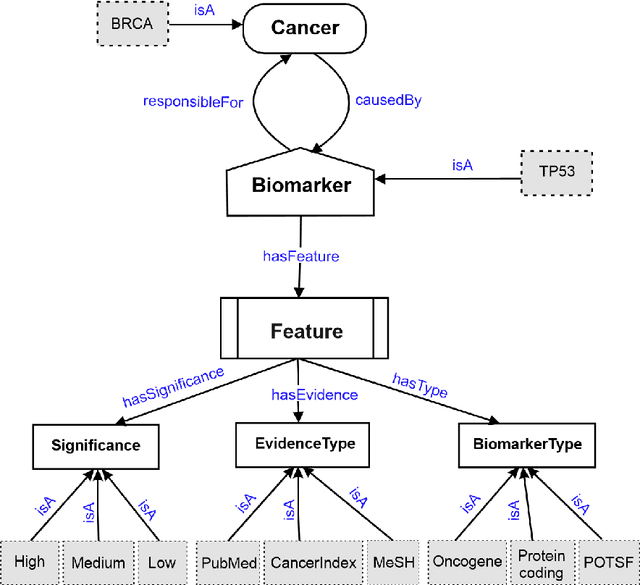
Domain experts often rely on up-to-date knowledge for apprehending and disseminating specific biological processes that help them design strategies to develop prevention and therapeutic decision-making. A challenging scenario for artificial intelligence (AI) is using biomedical data (e.g., texts, imaging, omics, and clinical) to provide diagnosis and treatment recommendations for cancerous conditions. Data and knowledge about cancer, drugs, genes, proteins, and their mechanism is spread across structured (knowledge bases (KBs)) and unstructured (e.g., scientific articles) sources. A large-scale knowledge graph (KG) can be constructed by integrating these data, followed by extracting facts about semantically interrelated entities and relations. Such KGs not only allow exploration and question answering (QA) but also allow domain experts to deduce new knowledge. However, exploring and querying large-scale KGs is tedious for non-domain users due to a lack of understanding of the underlying data assets and semantic technologies. In this paper, we develop a domain KG to leverage cancer-specific biomarker discovery and interactive QA. For this, a domain ontology called OncoNet Ontology (ONO) is developed to enable semantic reasoning for validating gene-disease relations. The KG is then enriched by harmonizing the ONO, controlled vocabularies, and additional biomedical concepts from scientific articles by employing BioBERT- and SciBERT-based information extraction (IE) methods. Further, since the biomedical domain is evolving, where new findings often replace old ones, without employing up-to-date findings, there is a high chance an AI system exhibits concept drift while providing diagnosis and treatment. Therefore, we finetuned the KG using large language models (LLMs) based on more recent articles and KBs that might not have been seen by the named entity recognition models.
Automated clinical coding using off-the-shelf large language models
Oct 10, 2023The task of assigning diagnostic ICD codes to patient hospital admissions is typically performed by expert human coders. Efforts towards automated ICD coding are dominated by supervised deep learning models. However, difficulties in learning to predict the large number of rare codes remain a barrier to adoption in clinical practice. In this work, we leverage off-the-shelf pre-trained generative large language models (LLMs) to develop a practical solution that is suitable for zero-shot and few-shot code assignment. Unsupervised pre-training alone does not guarantee precise knowledge of the ICD ontology and specialist clinical coding task, therefore we frame the task as information extraction, providing a description of each coded concept and asking the model to retrieve related mentions. For efficiency, rather than iterating over all codes, we leverage the hierarchical nature of the ICD ontology to sparsely search for relevant codes. Then, in a second stage, which we term 'meta-refinement', we utilise GPT-4 to select a subset of the relevant labels as predictions. We validate our method using Llama-2, GPT-3.5 and GPT-4 on the CodiEsp dataset of ICD-coded clinical case documents. Our tree-search method achieves state-of-the-art performance on rarer classes, achieving the best macro-F1 of 0.225, whilst achieving slightly lower micro-F1 of 0.157, compared to 0.216 and 0.219 respectively from PLM-ICD. To the best of our knowledge, this is the first method for automated ICD coding requiring no task-specific learning.
Inspire the Large Language Model by External Knowledge on BioMedical Named Entity Recognition
Sep 21, 2023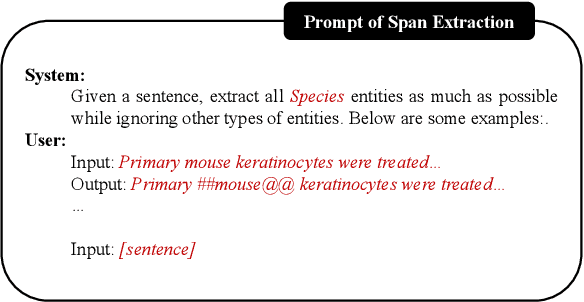
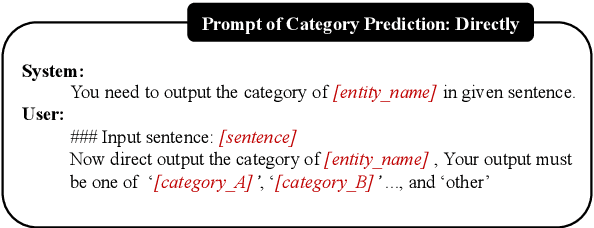
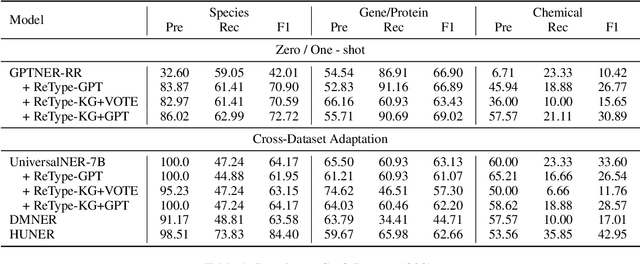

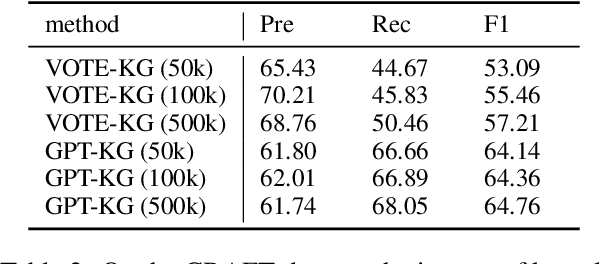
Large language models (LLMs) have demonstrated dominating performance in many NLP tasks, especially on generative tasks. However, they often fall short in some information extraction tasks, particularly those requiring domain-specific knowledge, such as Biomedical Named Entity Recognition (NER). In this paper, inspired by Chain-of-thought, we leverage the LLM to solve the Biomedical NER step-by-step: break down the NER task into entity span extraction and entity type determination. Additionally, for entity type determination, we inject entity knowledge to address the problem that LLM's lack of domain knowledge when predicting entity category. Experimental results show a significant improvement in our two-step BioNER approach compared to previous few-shot LLM baseline. Additionally, the incorporation of external knowledge significantly enhances entity category determination performance.
Cross-modal Prompts: Adapting Large Pre-trained Models for Audio-Visual Downstream Tasks
Nov 09, 2023In recent years, the deployment of large-scale pre-trained models in audio-visual downstream tasks has yielded remarkable outcomes. However, these models, primarily trained on single-modality unconstrained datasets, still encounter challenges in feature extraction for multi-modal tasks, leading to suboptimal performance. This limitation arises due to the introduction of irrelevant modality-specific information during encoding, which adversely affects the performance of downstream tasks. To address this challenge, this paper proposes a novel Dual-Guided Spatial-Channel-Temporal (DG-SCT) attention mechanism. This mechanism leverages audio and visual modalities as soft prompts to dynamically adjust the parameters of pre-trained models based on the current multi-modal input features. Specifically, the DG-SCT module incorporates trainable cross-modal interaction layers into pre-trained audio-visual encoders, allowing adaptive extraction of crucial information from the current modality across spatial, channel, and temporal dimensions, while preserving the frozen parameters of large-scale pre-trained models. Experimental evaluations demonstrate that our proposed model achieves state-of-the-art results across multiple downstream tasks, including AVE, AVVP, AVS, and AVQA. Furthermore, our model exhibits promising performance in challenging few-shot and zero-shot scenarios. The source code and pre-trained models are available at https://github.com/haoyi-duan/DG-SCT.
Weakly Supervised Reasoning by Neuro-Symbolic Approaches
Sep 19, 2023Deep learning has largely improved the performance of various natural language processing (NLP) tasks. However, most deep learning models are black-box machinery, and lack explicit interpretation. In this chapter, we will introduce our recent progress on neuro-symbolic approaches to NLP, which combines different schools of AI, namely, symbolism and connectionism. Generally, we will design a neural system with symbolic latent structures for an NLP task, and apply reinforcement learning or its relaxation to perform weakly supervised reasoning in the downstream task. Our framework has been successfully applied to various tasks, including table query reasoning, syntactic structure reasoning, information extraction reasoning, and rule reasoning. For each application, we will introduce the background, our approach, and experimental results.
Information Redundancy and Biases in Public Document Information Extraction Benchmarks
Apr 28, 2023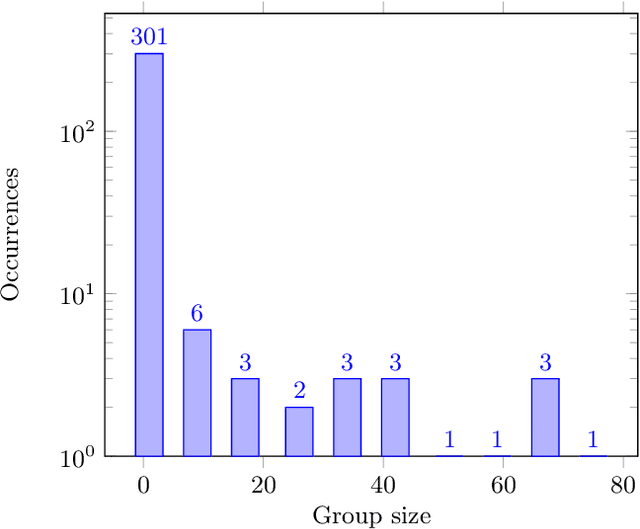
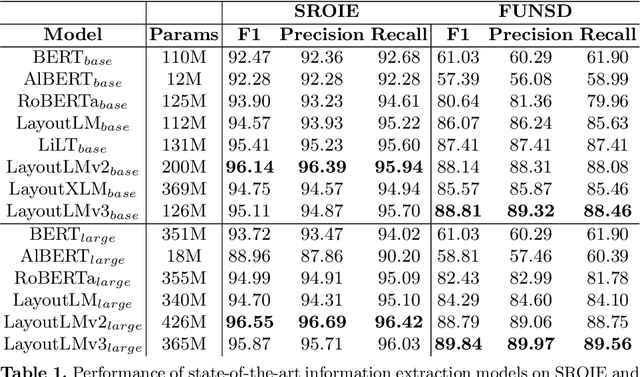

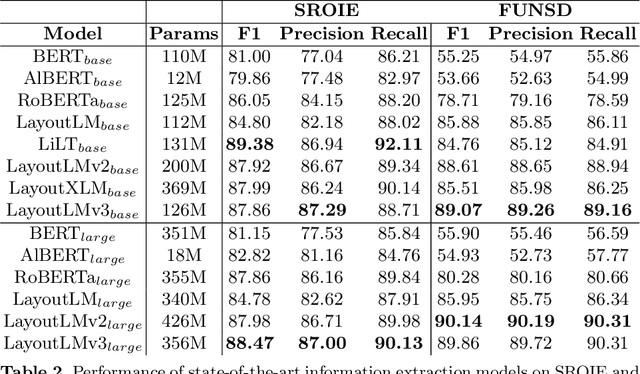
Advances in the Visually-rich Document Understanding (VrDU) field and particularly the Key-Information Extraction (KIE) task are marked with the emergence of efficient Transformer-based approaches such as the LayoutLM models. Despite the good performance of KIE models when fine-tuned on public benchmarks, they still struggle to generalize on complex real-life use-cases lacking sufficient document annotations. Our research highlighted that KIE standard benchmarks such as SROIE and FUNSD contain significant similarity between training and testing documents and can be adjusted to better evaluate the generalization of models. In this work, we designed experiments to quantify the information redundancy in public benchmarks, revealing a 75% template replication in SROIE official test set and 16% in FUNSD. We also proposed resampling strategies to provide benchmarks more representative of the generalization ability of models. We showed that models not suited for document analysis struggle on the adjusted splits dropping on average 10,5% F1 score on SROIE and 3.5% on FUNSD compared to multi-modal models dropping only 7,5% F1 on SROIE and 0.5% F1 on FUNSD.
An Information Extraction Study: Take In Mind the Tokenization!
Apr 01, 2023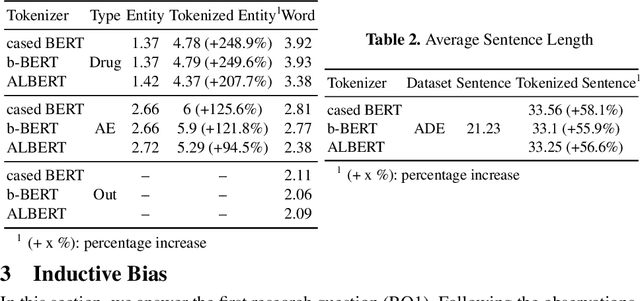
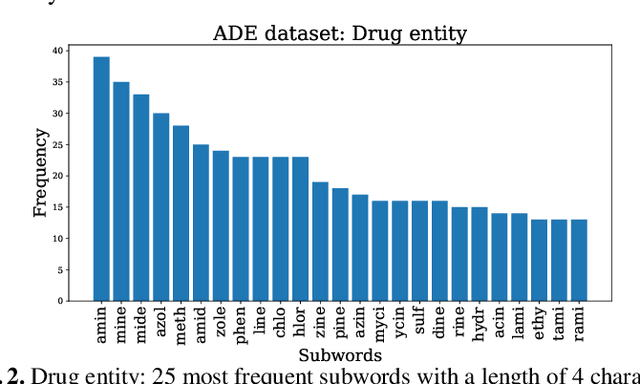
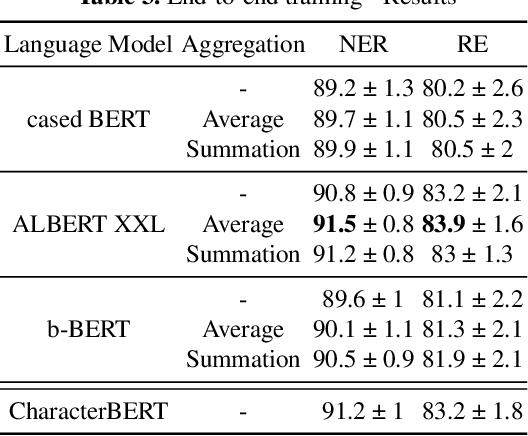
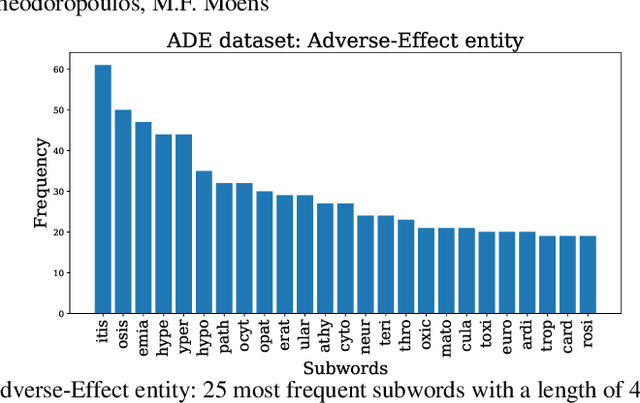
Current research on the advantages and trade-offs of using characters, instead of tokenized text, as input for deep learning models, has evolved substantially. New token-free models remove the traditional tokenization step; however, their efficiency remains unclear. Moreover, the effect of tokenization is relatively unexplored in sequence tagging tasks. To this end, we investigate the impact of tokenization when extracting information from documents and present a comparative study and analysis of subword-based and character-based models. Specifically, we study Information Extraction (IE) from biomedical texts. The main outcome is twofold: tokenization patterns can introduce inductive bias that results in state-of-the-art performance, and the character-based models produce promising results; thus, transitioning to token-free IE models is feasible.
OmniEvent: A Comprehensive, Fair, and Easy-to-Use Toolkit for Event Understanding
Sep 25, 2023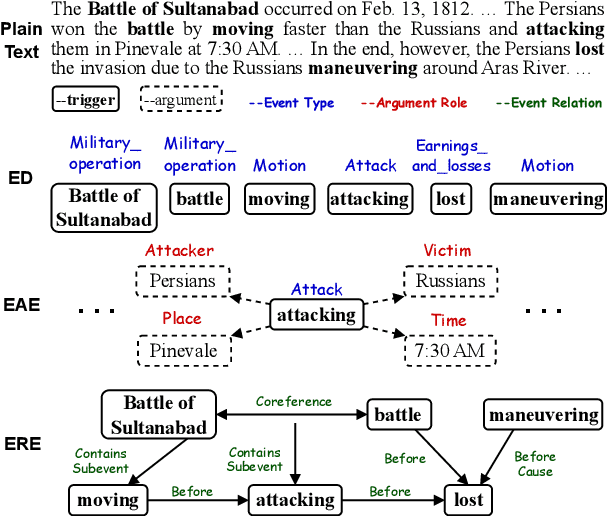
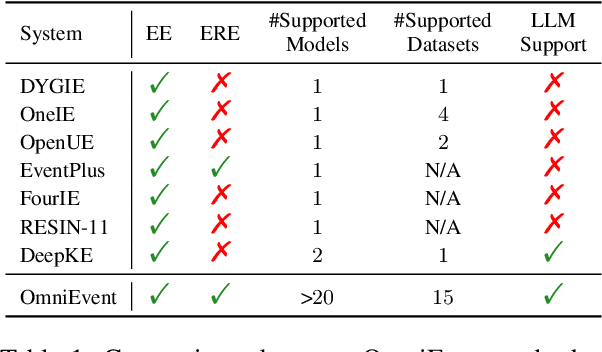
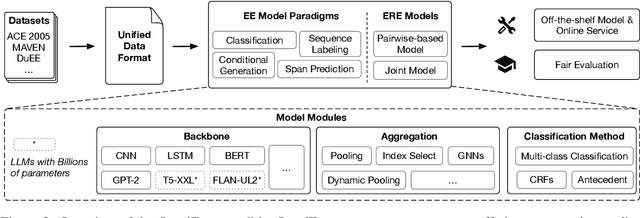
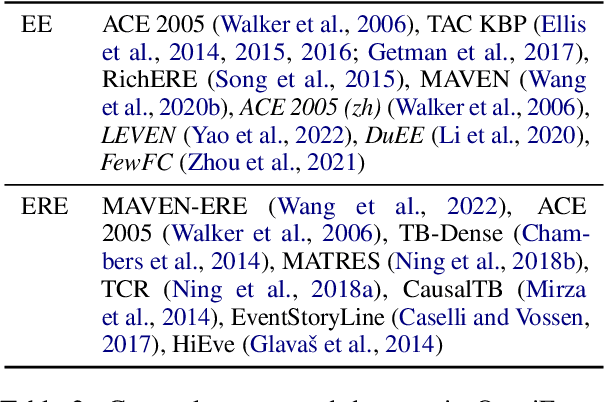
Event understanding aims at understanding the content and relationship of events within texts, which covers multiple complicated information extraction tasks: event detection, event argument extraction, and event relation extraction. To facilitate related research and application, we present an event understanding toolkit OmniEvent, which features three desiderata: (1) Comprehensive. OmniEvent supports mainstream modeling paradigms of all the event understanding tasks and the processing of 15 widely-used English and Chinese datasets. (2) Fair. OmniEvent carefully handles the inconspicuous evaluation pitfalls reported in Peng et al. (2023), which ensures fair comparisons between different models. (3) Easy-to-use. OmniEvent is designed to be easily used by users with varying needs. We provide off-the-shelf models that can be directly deployed as web services. The modular framework also enables users to easily implement and evaluate new event understanding models with OmniEvent. The toolkit (https://github.com/THU-KEG/OmniEvent) is publicly released along with the demonstration website and video (https://omnievent.xlore.cn/).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge