"Information Extraction": models, code, and papers
BioInstruct: Instruction Tuning of Large Language Models for Biomedical Natural Language Processing
Oct 30, 2023Large language models (LLMs) has achieved a great success in many natural language processing (NLP) tasks. This is achieved by pretraining of LLMs on vast amount of data and then instruction tuning to specific domains. However, only a few instructions in the biomedical domain have been published. To address this issue, we introduce BioInstruct, a customized task-specific instruction dataset containing more than 25,000 examples. This dataset was generated attractively by prompting a GPT-4 language model with a three-seed-sample of 80 human-curated instructions. By fine-tuning LLMs using the BioInstruct dataset, we aim to optimize the LLM's performance in biomedical natural language processing (BioNLP). We conducted instruction tuning on the LLaMA LLMs (1\&2, 7B\&13B) and evaluated them on BioNLP applications, including information extraction, question answering, and text generation. We also evaluated how instructions contributed to model performance using multi-tasking learning principles.
Deciphering 'What' and 'Where' Visual Pathways from Spectral Clustering of Layer-Distributed Neural Representations
Dec 11, 2023We present an approach for analyzing grouping information contained within a neural network's activations, permitting extraction of spatial layout and semantic segmentation from the behavior of large pre-trained vision models. Unlike prior work, our method conducts a wholistic analysis of a network's activation state, leveraging features from all layers and obviating the need to guess which part of the model contains relevant information. Motivated by classic spectral clustering, we formulate this analysis in terms of an optimization objective involving a set of affinity matrices, each formed by comparing features within a different layer. Solving this optimization problem using gradient descent allows our technique to scale from single images to dataset-level analysis, including, in the latter, both intra- and inter-image relationships. Analyzing a pre-trained generative transformer provides insight into the computational strategy learned by such models. Equating affinity with key-query similarity across attention layers yields eigenvectors encoding scene spatial layout, whereas defining affinity by value vector similarity yields eigenvectors encoding object identity. This result suggests that key and query vectors coordinate attentional information flow according to spatial proximity (a `where' pathway), while value vectors refine a semantic category representation (a `what' pathway).
Disentangling Extraction and Reasoning in Multi-hop Spatial Reasoning
Oct 25, 2023Spatial reasoning over text is challenging as the models not only need to extract the direct spatial information from the text but also reason over those and infer implicit spatial relations. Recent studies highlight the struggles even large language models encounter when it comes to performing spatial reasoning over text. In this paper, we explore the potential benefits of disentangling the processes of information extraction and reasoning in models to address this challenge. To explore this, we design various models that disentangle extraction and reasoning(either symbolic or neural) and compare them with state-of-the-art(SOTA) baselines with no explicit design for these parts. Our experimental results consistently demonstrate the efficacy of disentangling, showcasing its ability to enhance models' generalizability within realistic data domains.
Investigating Deep-Learning NLP for Automating the Extraction of Oncology Efficacy Endpoints from Scientific Literature
Nov 03, 2023Benchmarking drug efficacy is a critical step in clinical trial design and planning. The challenge is that much of the data on efficacy endpoints is stored in scientific papers in free text form, so extraction of such data is currently a largely manual task. Our objective is to automate this task as much as possible. In this study we have developed and optimised a framework to extract efficacy endpoints from text in scientific papers, using a machine learning approach. Our machine learning model predicts 25 classes associated with efficacy endpoints and leads to high F1 scores (harmonic mean of precision and recall) of 96.4% on the test set, and 93.9% and 93.7% on two case studies. These methods were evaluated against - and showed strong agreement with - subject matter experts and show significant promise in the future of automating the extraction of clinical endpoints from free text. Clinical information extraction from text data is currently a laborious manual task which scales poorly and is prone to human error. Demonstrating the ability to extract efficacy endpoints automatically shows great promise for accelerating clinical trial design moving forwards.
Data-Free Distillation of Language Model by Text-to-Text Transfer
Nov 03, 2023Data-Free Knowledge Distillation (DFKD) plays a vital role in compressing the model when original training data is unavailable. Previous works for DFKD in NLP mainly focus on distilling encoder-only structures like BERT on classification tasks, which overlook the notable progress of generative language modeling. In this work, we propose a novel DFKD framework, namely DFKD-T$^{3}$, where the pretrained generative language model can also serve as a controllable data generator for model compression. This novel framework DFKD-T$^{3}$ leads to an end-to-end learnable text-to-text framework to transform the general domain corpus to compression-friendly task data, targeting to improve both the \textit{specificity} and \textit{diversity}. Extensive experiments show that our method can boost the distillation performance in various downstream tasks such as sentiment analysis, linguistic acceptability, and information extraction. Furthermore, we show that the generated texts can be directly used for distilling other language models and outperform the SOTA methods, making our method more appealing in a general DFKD setting. Our code is available at https://gitee.com/mindspore/models/tree/master/research/nlp/DFKD\_T3.
Dance of Channel and Sequence: An Efficient Attention-Based Approach for Multivariate Time Series Forecasting
Dec 11, 2023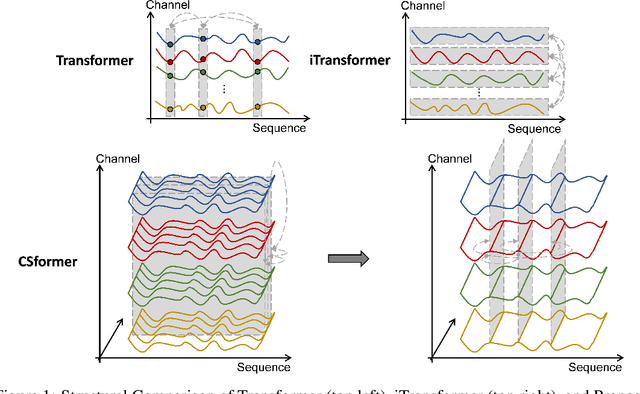

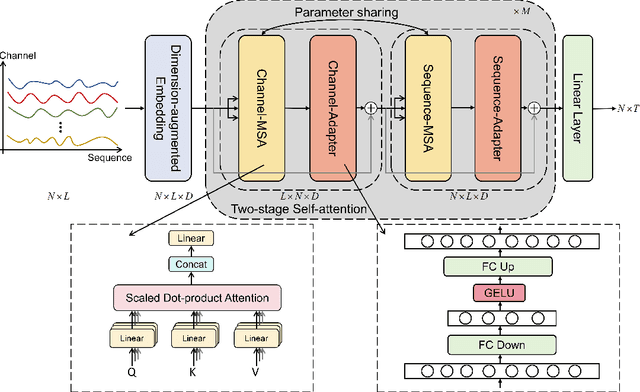
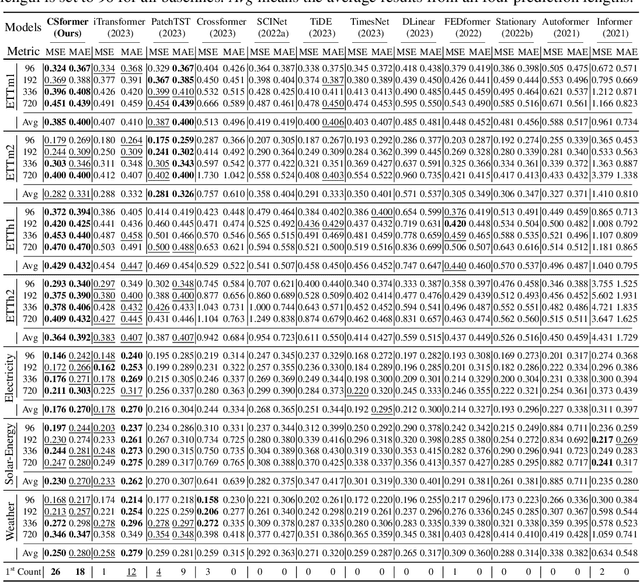
In recent developments, predictive models for multivariate time series analysis have exhibited commendable performance through the adoption of the prevalent principle of channel independence. Nevertheless, it is imperative to acknowledge the intricate interplay among channels, which fundamentally influences the outcomes of multivariate predictions. Consequently, the notion of channel independence, while offering utility to a certain extent, becomes increasingly impractical, leading to information degradation. In response to this pressing concern, we present CSformer, an innovative framework characterized by a meticulously engineered two-stage self-attention mechanism. This mechanism is purposefully designed to enable the segregated extraction of sequence-specific and channel-specific information, while sharing parameters to promote synergy and mutual reinforcement between sequences and channels. Simultaneously, we introduce sequence adapters and channel adapters, ensuring the model's ability to discern salient features across various dimensions. Rigorous experimentation, spanning multiple real-world datasets, underscores the robustness of our approach, consistently establishing its position at the forefront of predictive performance across all datasets. This augmentation substantially enhances the capacity for feature extraction inherent to multivariate time series data, facilitating a more comprehensive exploitation of the available information.
Self-Verification Improves Few-Shot Clinical Information Extraction
May 30, 2023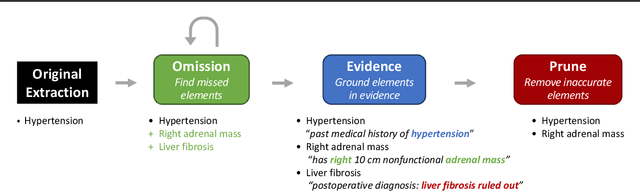
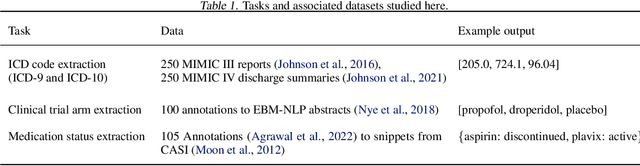
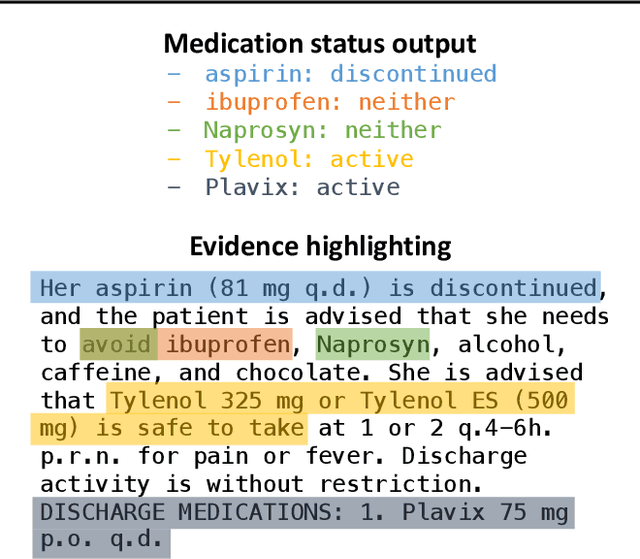


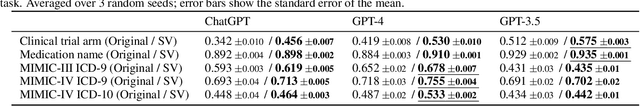
Extracting patient information from unstructured text is a critical task in health decision-support and clinical research. Large language models (LLMs) have shown the potential to accelerate clinical curation via few-shot in-context learning, in contrast to supervised learning which requires much more costly human annotations. However, despite drastic advances in modern LLMs such as GPT-4, they still struggle with issues regarding accuracy and interpretability, especially in mission-critical domains such as health. Here, we explore a general mitigation framework using self-verification, which leverages the LLM to provide provenance for its own extraction and check its own outputs. This is made possible by the asymmetry between verification and generation, where the latter is often much easier than the former. Experimental results show that our method consistently improves accuracy for various LLMs in standard clinical information extraction tasks. Additionally, self-verification yields interpretations in the form of a short text span corresponding to each output, which makes it very efficient for human experts to audit the results, paving the way towards trustworthy extraction of clinical information in resource-constrained scenarios. To facilitate future research in this direction, we release our code and prompts.
InstructIE: A Chinese Instruction-based Information Extraction Dataset
May 19, 2023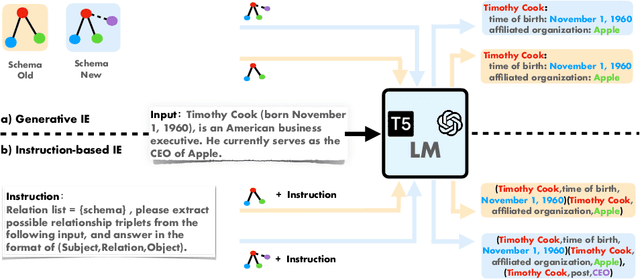
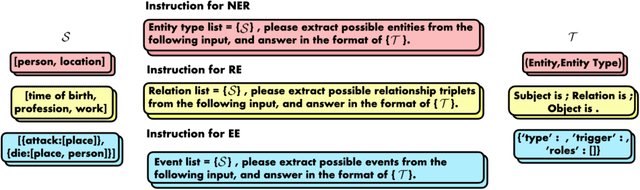
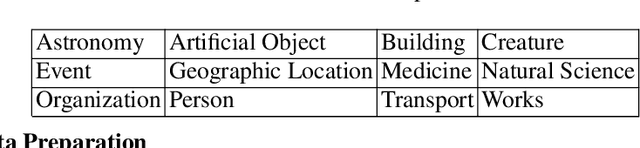

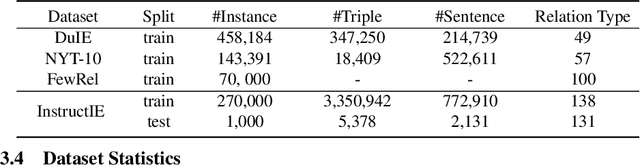
We introduce a new Information Extraction (IE) task dubbed Instruction-based IE, which aims to ask the system to follow specific instructions or guidelines to extract information. To facilitate research in this area, we construct a dataset called InstructIE, consisting of 270,000 weakly supervised data from Chinese Wikipedia and 1,000 high-quality crowdsourced annotated instances. We further evaluate the performance of various baseline models on the InstructIE dataset. The results reveal that although current models exhibit promising performance, there is still room for improvement. Furthermore, we conduct a comprehensive case study analysis, underlining the challenges inherent in the Instruction-based IE task. Code and dataset are available at https://github.com/zjunlp/DeepKE/tree/main/example/llm.
Bridging the Gap between Multi-focus and Multi-modal: A Focused Integration Framework for Multi-modal Image Fusion
Nov 03, 2023Multi-modal image fusion (MMIF) integrates valuable information from different modality images into a fused one. However, the fusion of multiple visible images with different focal regions and infrared images is a unprecedented challenge in real MMIF applications. This is because of the limited depth of the focus of visible optical lenses, which impedes the simultaneous capture of the focal information within the same scene. To address this issue, in this paper, we propose a MMIF framework for joint focused integration and modalities information extraction. Specifically, a semi-sparsity-based smoothing filter is introduced to decompose the images into structure and texture components. Subsequently, a novel multi-scale operator is proposed to fuse the texture components, capable of detecting significant information by considering the pixel focus attributes and relevant data from various modal images. Additionally, to achieve an effective capture of scene luminance and reasonable contrast maintenance, we consider the distribution of energy information in the structural components in terms of multi-directional frequency variance and information entropy. Extensive experiments on existing MMIF datasets, as well as the object detection and depth estimation tasks, consistently demonstrate that the proposed algorithm can surpass the state-of-the-art methods in visual perception and quantitative evaluation. The code is available at https://github.com/ixilai/MFIF-MMIF.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge