"Information Extraction": models, code, and papers
Image Restoration with Point Spread Function Regularization and Active Learning
Oct 31, 2023Large-scale astronomical surveys can capture numerous images of celestial objects, including galaxies and nebulae. Analysing and processing these images can reveal intricate internal structures of these objects, allowing researchers to conduct comprehensive studies on their morphology, evolution, and physical properties. However, varying noise levels and point spread functions can hamper the accuracy and efficiency of information extraction from these images. To mitigate these effects, we propose a novel image restoration algorithm that connects a deep learning-based restoration algorithm with a high-fidelity telescope simulator. During the training stage, the simulator generates images with different levels of blur and noise to train the neural network based on the quality of restored images. After training, the neural network can directly restore images obtained by the telescope, as represented by the simulator. We have tested the algorithm using real and simulated observation data and have found that it effectively enhances fine structures in blurry images and increases the quality of observation images. This algorithm can be applied to large-scale sky survey data, such as data obtained by LSST, Euclid, and CSST, to further improve the accuracy and efficiency of information extraction, promoting advances in the field of astronomical research.
Optimized measurements of chaotic dynamical systems via the information bottleneck
Nov 08, 2023Deterministic chaos permits a precise notion of a "perfect measurement" as one that, when obtained repeatedly, captures all of the information created by the system's evolution with minimal redundancy. Finding an optimal measurement is challenging, and has generally required intimate knowledge of the dynamics in the few cases where it has been done. We establish an equivalence between a perfect measurement and a variant of the information bottleneck. As a consequence, we can employ machine learning to optimize measurement processes that efficiently extract information from trajectory data. We obtain approximately optimal measurements for multiple chaotic maps and lay the necessary groundwork for efficient information extraction from general time series.
ChiMed-GPT: A Chinese Medical Large Language Model with Full Training Regime and Better Alignment to Human Preferences
Nov 23, 2023
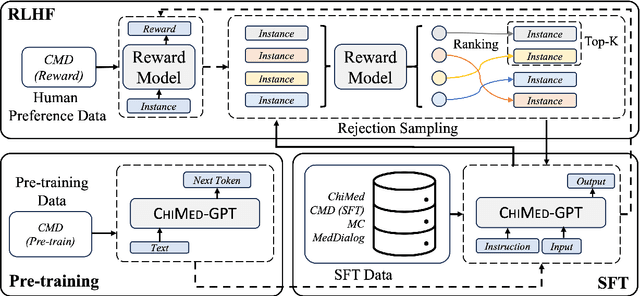

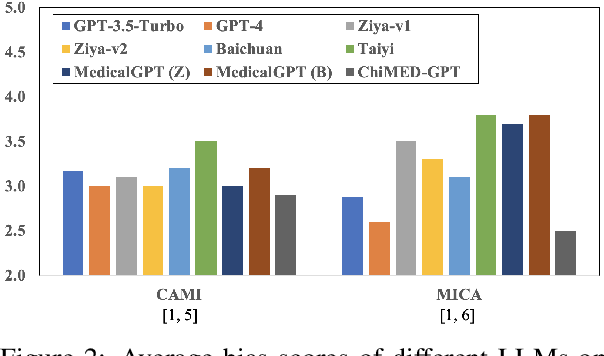
Recently, the increasing demand for superior medical services has highlighted the discrepancies in the medical infrastructure. With big data, especially texts, forming the foundation of medical services, there is an exigent need for effective natural language processing (NLP) solutions tailored to the healthcare domain. Conventional approaches leveraging pre-trained models present promising results in this domain and current large language models (LLMs) offer advanced foundation for medical text processing. However, most medical LLMs are trained only with supervised fine-tuning (SFT), even though it efficiently empowers LLMs to understand and respond to medical instructions but is ineffective in learning domain knowledge and aligning with human preference. Another engineering barrier that prevents current medical LLM from better text processing ability is their restricted context length (e.g., 2,048 tokens), making it hard for the LLMs to process long context, which is frequently required in the medical domain. In this work, we propose ChiMed-GPT, a new benchmark LLM designed explicitly for Chinese medical domain, with enlarged context length to 4,096 tokens and undergoes a comprehensive training regime with pre-training, SFT, and RLHF. Evaluations on real-world tasks including information extraction, question answering, and dialogue generation demonstrate ChiMed-GPT's superior performance over general domain LLMs. Furthermore, we analyze possible biases through prompting ChiMed-GPT to perform attitude scales regarding discrimination of patients, so as to contribute to further responsible development of LLMs in the medical domain. The code and model are released at https://github.com/synlp/ChiMed-GPT.
An Attention-Based Denoising Framework for Personality Detection in Social Media Texts
Nov 16, 2023In social media networks, users produce a large amount of text content anytime, providing researchers with a valuable approach to digging for personality-related information. Personality detection based on user-generated texts is a universal method that can be used to build user portraits. The presence of noise in social media texts hinders personality detection. However, previous studies have not fully addressed this challenge. Inspired by the scanning reading technique, we propose an attention-based information extraction mechanism (AIEM) for long texts, which is applied to quickly locate valuable pieces of information, and focus more attention on the deep semantics of key pieces. Then, we provide a novel attention-based denoising framework (ADF) for personality detection tasks and achieve state-of-the-art performance on two commonly used datasets. Notably, we obtain an average accuracy improvement of 10.2% on the gold standard Twitter-Myers-Briggs Type Indicator (Twitter-MBTI) dataset. We made our code publicly available on GitHub. We shed light on how AIEM works to magnify personality-related signals.
CMFDFormer: Transformer-based Copy-Move Forgery Detection with Continual Learning
Nov 22, 2023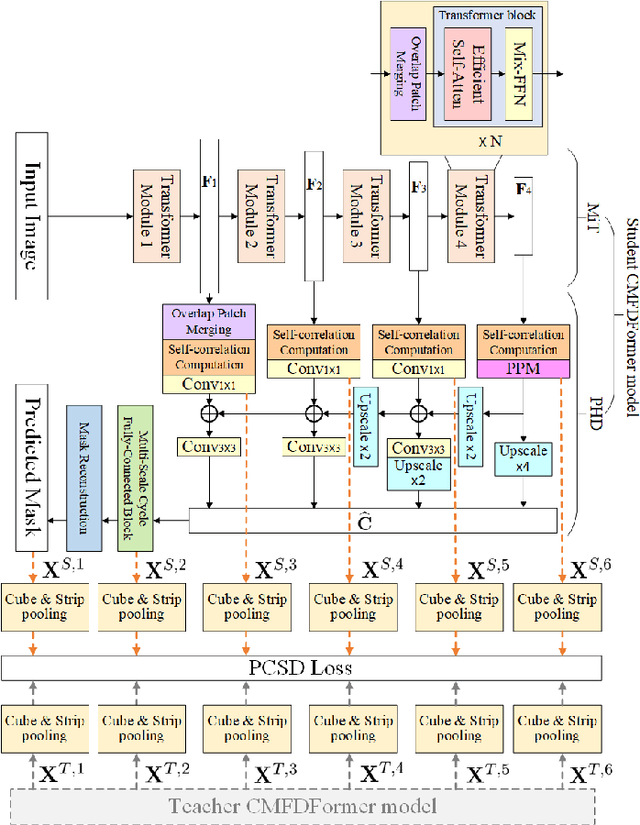
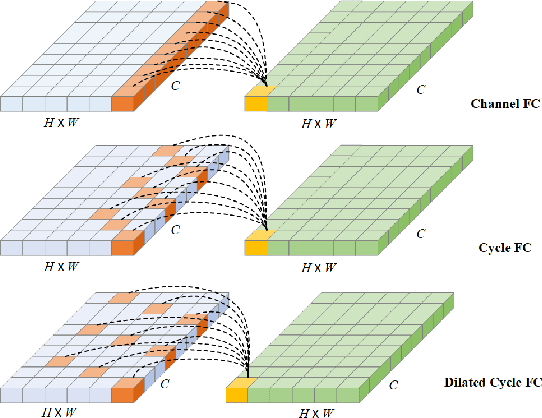
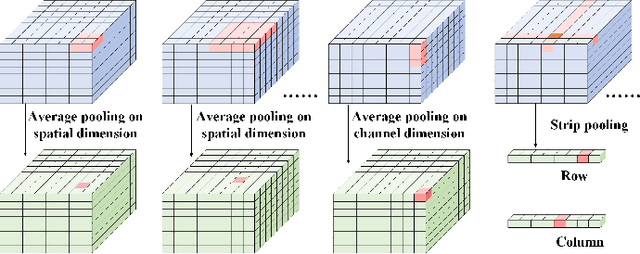
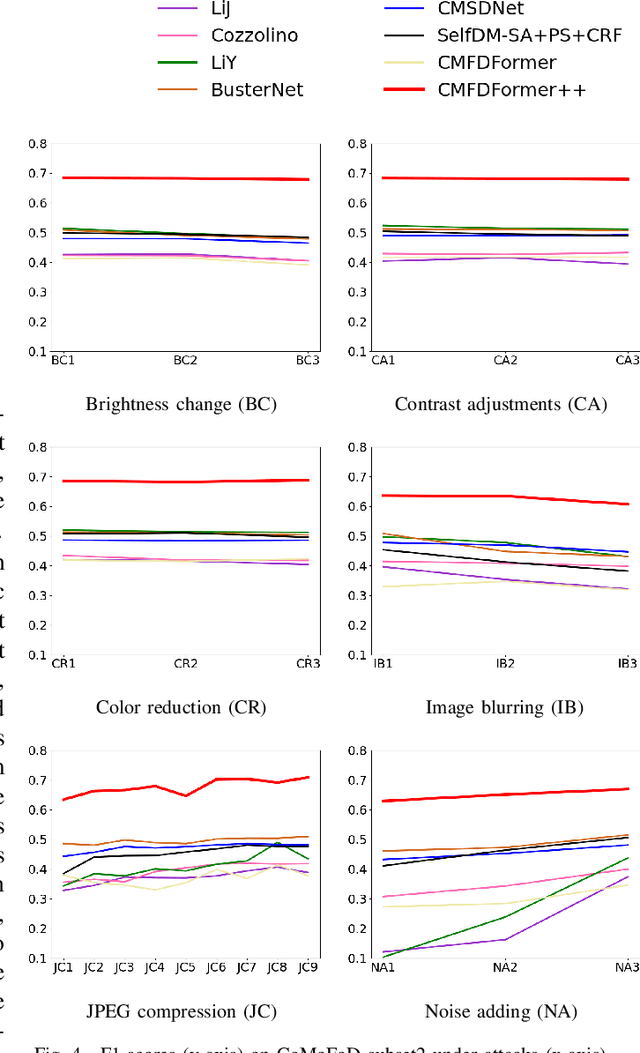
Copy-move forgery detection aims at detecting duplicated regions in a suspected forged image, and deep learning based copy-move forgery detection methods are in the ascendant. These deep learning based methods heavily rely on synthetic training data, and the performance will degrade when facing new tasks. In this paper, we propose a Transformer-style copy-move forgery detection network named as CMFDFormer, and provide a novel PCSD (Pooled Cube and Strip Distillation) continual learning framework to help CMFDFormer handle new tasks. CMFDFormer consists of a MiT (Mix Transformer) backbone network and a PHD (Pluggable Hybrid Decoder) mask prediction network. The MiT backbone network is a Transformer-style network which is adopted on the basis of comprehensive analyses with CNN-style and MLP-style backbones. The PHD network is constructed based on self-correlation computation, hierarchical feature integration, a multi-scale cycle fully-connected block and a mask reconstruction block. The PHD network is applicable to feature extractors of different styles for hierarchical multi-scale information extraction, achieving comparable performance. Last but not least, we propose a PCSD continual learning framework to improve the forgery detectability and avoid catastrophic forgetting when handling new tasks. Our continual learning framework restricts intermediate features from the PHD network, and takes advantage of both cube pooling and strip pooling. Extensive experiments on publicly available datasets demonstrate the good performance of CMFDFormer and the effectiveness of the PCSD continual learning framework.
Uncertainty Estimation on Sequential Labeling via Uncertainty Transmission
Nov 15, 2023Sequential labeling is a task predicting labels for each token in a sequence, such as Named Entity Recognition (NER). NER tasks aim to extract entities and predict their labels given a text, which is important in information extraction. Although previous works have shown great progress in improving NER performance, uncertainty estimation on NER (UE-NER) is still underexplored but essential. This work focuses on UE-NER, which aims to estimate uncertainty scores for the NER predictions. Previous uncertainty estimation models often overlook two unique characteristics of NER: the connection between entities (i.e., one entity embedding is learned based on the other ones) and wrong span cases in the entity extraction subtask. Therefore, we propose a Sequential Labeling Posterior Network (SLPN) to estimate uncertainty scores for the extracted entities, considering uncertainty transmitted from other tokens. Moreover, we have defined an evaluation strategy to address the specificity of wrong-span cases. Our SLPN has achieved significant improvements on two datasets, such as a 5.54-point improvement in AUPR on the MIT-Restaurant dataset.
CARE: Extracting Experimental Findings From Clinical Literature
Nov 16, 2023Extracting fine-grained experimental findings from literature can provide massive utility for scientific applications. Prior work has focused on developing annotation schemas and datasets for limited aspects of this problem, leading to simpler information extraction datasets which do not capture the real-world complexity and nuance required for this task. Focusing on biomedicine, this work presents CARE (Clinical Aggregation-oriented Result Extraction) -- a new IE dataset for the task of extracting clinical findings. We develop a new annotation schema capturing fine-grained findings as n-ary relations between entities and attributes, which includes phenomena challenging for current IE systems such as discontinuous entity spans, nested relations, and variable arity n-ary relations. Using this schema, we collect extensive annotations for 700 abstracts from two sources: clinical trials and case reports. We also benchmark the performance of various state-of-the-art IE systems on our dataset, including extractive models and generative LLMs in fully supervised and limited data settings. Our results demonstrate the difficulty of our dataset -- even SOTA models such as GPT4 struggle, particularly on relation extraction. We release our annotation schema and CARE to encourage further research on extracting and aggregating scientific findings from literature.
Chain of Thought with Explicit Evidence Reasoning for Few-shot Relation Extraction
Nov 15, 2023Few-shot relation extraction involves identifying the type of relationship between two specific entities within a text, using a limited number of annotated samples. A variety of solutions to this problem have emerged by applying meta-learning and neural graph techniques which typically necessitate a training process for adaptation. Recently, the strategy of in-context learning has been demonstrating notable results without the need of training. Few studies have already utilized in-context learning for zero-shot information extraction. Unfortunately, the evidence for inference is either not considered or implicitly modeled during the construction of chain-of-thought prompts. In this paper, we propose a novel approach for few-shot relation extraction using large language models, named CoT-ER, chain-of-thought with explicit evidence reasoning. In particular, CoT-ER first induces large language models to generate evidences using task-specific and concept-level knowledge. Then these evidences are explicitly incorporated into chain-of-thought prompting for relation extraction. Experimental results demonstrate that our CoT-ER approach (with 0% training data) achieves competitive performance compared to the fully-supervised (with 100% training data) state-of-the-art approach on the FewRel1.0 and FewRel2.0 datasets.
PIVOINE: Instruction Tuning for Open-world Information Extraction
May 24, 2023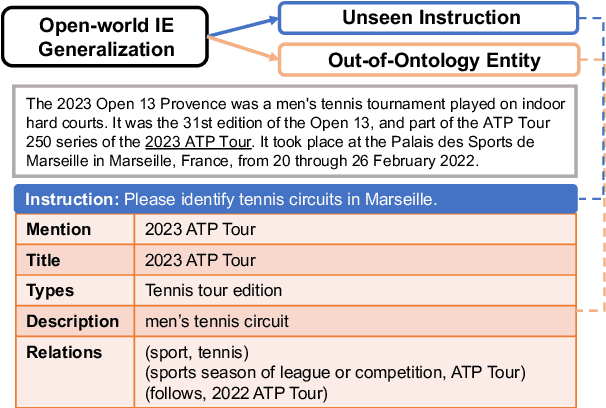

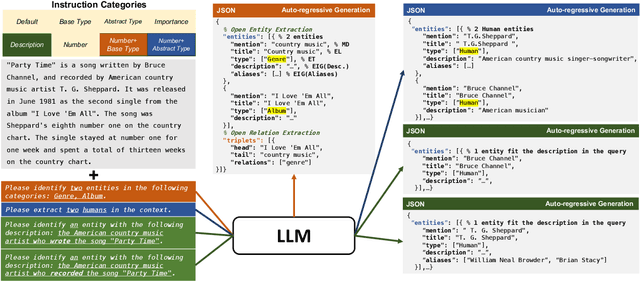
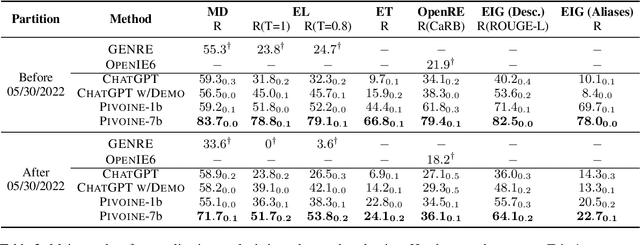
We consider the problem of Open-world Information Extraction (Open-world IE), which extracts comprehensive entity profiles from unstructured texts. Different from the conventional closed-world setting of Information Extraction (IE), Open-world IE considers a more general situation where entities and relations could be beyond a predefined ontology. More importantly, we seek to develop a large language model (LLM) that is able to perform Open-world IE to extract desirable entity profiles characterized by (possibly fine-grained) natural language instructions. We achieve this by finetuning LLMs using instruction tuning. In particular, we construct INSTRUCTOPENWIKI, a substantial instruction tuning dataset for Open-world IE enriched with a comprehensive corpus, extensive annotations, and diverse instructions. We finetune the pretrained BLOOM models on INSTRUCTOPENWIKI and obtain PIVOINE, an LLM for Open-world IE with strong instruction-following capabilities. Our experiments demonstrate that PIVOINE significantly outperforms traditional closed-world methods and other LLM baselines, displaying impressive generalization capabilities on both unseen instructions and out-of-ontology cases. Consequently, PIVOINE emerges as a promising solution to tackle the open-world challenge in IE effectively.
Comparison of pipeline, sequence-to-sequence, and GPT models for end-to-end relation extraction: experiments with the rare disease use-case
Nov 22, 2023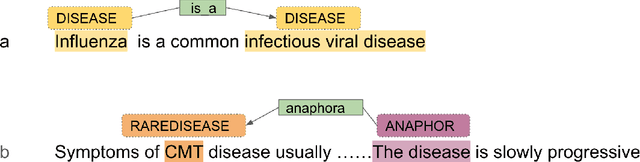
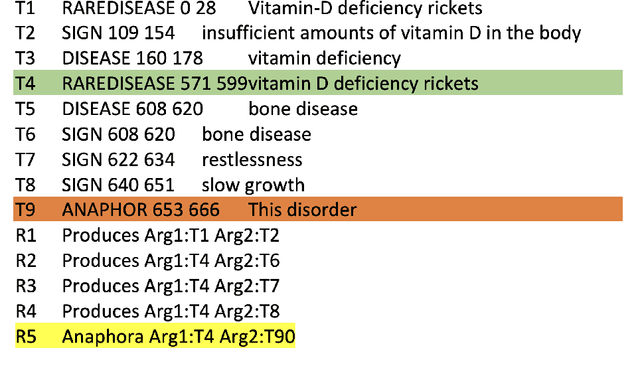
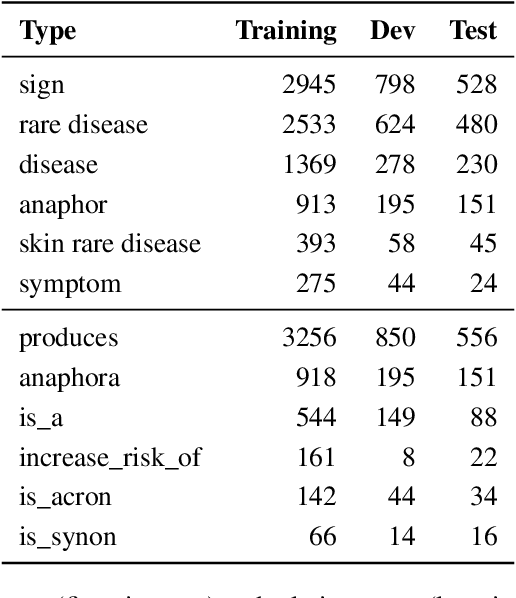


End-to-end relation extraction (E2ERE) is an important and realistic application of natural language processing (NLP) in biomedicine. In this paper, we aim to compare three prevailing paradigms for E2ERE using a complex dataset focused on rare diseases involving discontinuous and nested entities. We use the RareDis information extraction dataset to evaluate three competing approaches (for E2ERE): NER $\rightarrow$ RE pipelines, joint sequence to sequence models, and generative pre-trained transformer (GPT) models. We use comparable state-of-the-art models and best practices for each of these approaches and conduct error analyses to assess their failure modes. Our findings reveal that pipeline models are still the best, while sequence-to-sequence models are not far behind; GPT models with eight times as many parameters are worse than even sequence-to-sequence models and lose to pipeline models by over 10 F1 points. Partial matches and discontinuous entities caused many NER errors contributing to lower overall E2E performances. We also verify these findings on a second E2ERE dataset for chemical-protein interactions. Although generative LM-based methods are more suitable for zero-shot settings, when training data is available, our results show that it is better to work with more conventional models trained and tailored for E2ERE. More innovative methods are needed to marry the best of the both worlds from smaller encoder-decoder pipeline models and the larger GPT models to improve E2ERE. As of now, we see that well designed pipeline models offer substantial performance gains at a lower cost and carbon footprint for E2ERE. Our contribution is also the first to conduct E2ERE for the RareDis dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge