"Image": models, code, and papers
RFD-ECNet: Extreme Underwater Image Compression with Reference to Feature Dictionar
Aug 17, 2023
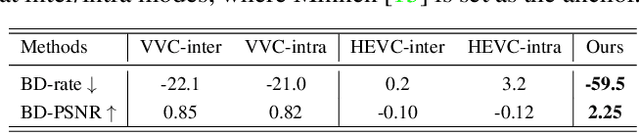
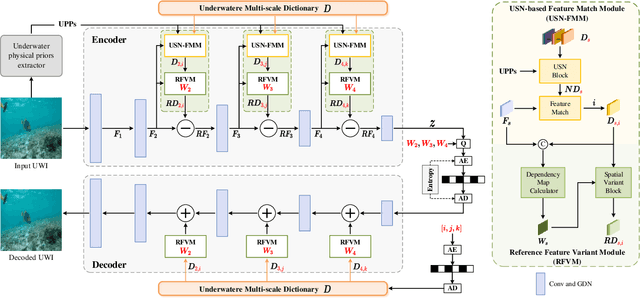

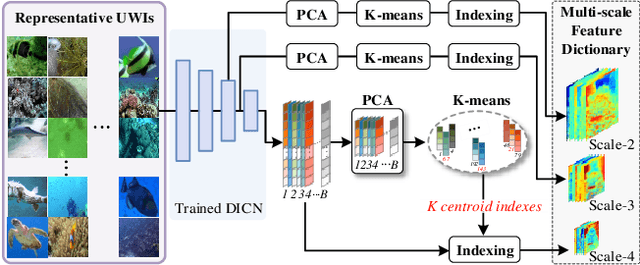
Thriving underwater applications demand efficient extreme compression technology to realize the transmission of underwater images (UWIs) in very narrow underwater bandwidth. However, existing image compression methods achieve inferior performance on UWIs because they do not consider the characteristics of UWIs: (1) Multifarious underwater styles of color shift and distance-dependent clarity, caused by the unique underwater physical imaging; (2) Massive redundancy between different UWIs, caused by the fact that different UWIs contain several common ocean objects, which have plenty of similarities in structures and semantics. To remove redundancy among UWIs, we first construct an exhaustive underwater multi-scale feature dictionary to provide coarse-to-fine reference features for UWI compression. Subsequently, an extreme UWI compression network with reference to the feature dictionary (RFD-ECNet) is creatively proposed, which utilizes feature match and reference feature variant to significantly remove redundancy among UWIs. To align the multifarious underwater styles and improve the accuracy of feature match, an underwater style normalized block (USNB) is proposed, which utilizes underwater physical priors extracted from the underwater physical imaging model to normalize the underwater styles of dictionary features toward the input. Moreover, a reference feature variant module (RFVM) is designed to adaptively morph the reference features, improving the similarity between the reference and input features. Experimental results on four UWI datasets show that our RFD-ECNet is the first work that achieves a significant BD-rate saving of 31% over the most advanced VVC.
Cooperative Colorization: Exploring Latent Cross-Domain Priors for NIR Image Spectrum Translation
Aug 07, 2023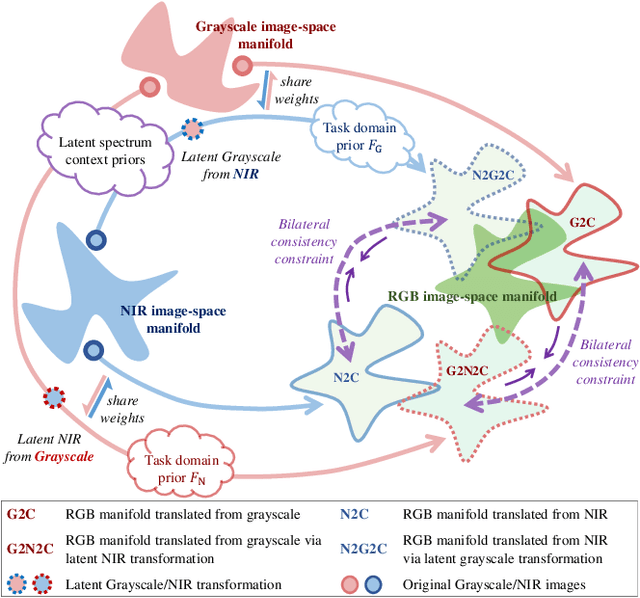
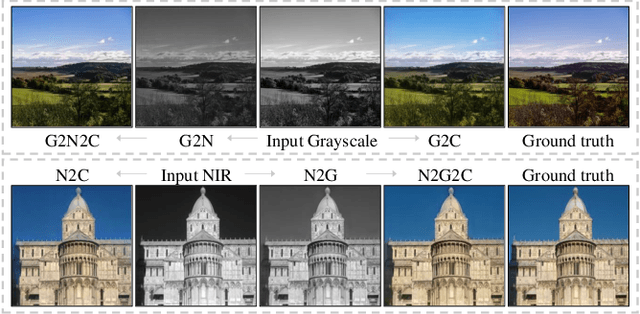
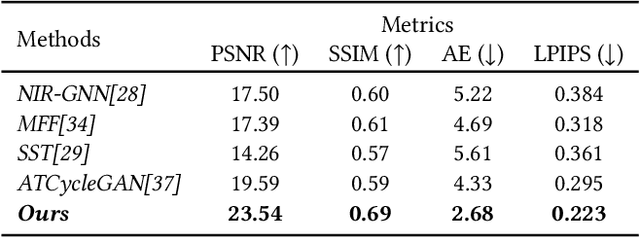

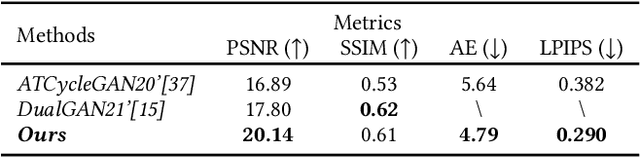
Near-infrared (NIR) image spectrum translation is a challenging problem with many promising applications. Existing methods struggle with the mapping ambiguity between the NIR and the RGB domains, and generalize poorly due to the limitations of models' learning capabilities and the unavailability of sufficient NIR-RGB image pairs for training. To address these challenges, we propose a cooperative learning paradigm that colorizes NIR images in parallel with another proxy grayscale colorization task by exploring latent cross-domain priors (i.e., latent spectrum context priors and task domain priors), dubbed CoColor. The complementary statistical and semantic spectrum information from these two task domains -- in the forms of pre-trained colorization networks -- are brought in as task domain priors. A bilateral domain translation module is subsequently designed, in which intermittent NIR images are generated from grayscale and colorized in parallel with authentic NIR images; and vice versa for the grayscale images. These intermittent transformations act as latent spectrum context priors for efficient domain knowledge exchange. We progressively fine-tune and fuse these modules with a series of pixel-level and feature-level consistency constraints. Experiments show that our proposed cooperative learning framework produces satisfactory spectrum translation outputs with diverse colors and rich textures, and outperforms state-of-the-art counterparts by 3.95dB and 4.66dB in terms of PNSR for the NIR and grayscale colorization tasks, respectively.
LiDAR Data Synthesis with Denoising Diffusion Probabilistic Models
Sep 17, 2023Generative modeling of 3D LiDAR data is an emerging task with promising applications for autonomous mobile robots, such as scalable simulation, scene manipulation, and sparse-to-dense completion of LiDAR point clouds. Existing approaches have shown the feasibility of image-based LiDAR data generation using deep generative models while still struggling with the fidelity of generated data and training instability. In this work, we present R2DM, a novel generative model for LiDAR data that can generate diverse and high-fidelity 3D scene point clouds based on the image representation of range and reflectance intensity. Our method is based on the denoising diffusion probabilistic models (DDPMs), which have demonstrated impressive results among generative model frameworks and have been significantly progressing in recent years. To effectively train DDPMs on the LiDAR domain, we first conduct an in-depth analysis regarding data representation, training objective, and spatial inductive bias. Based on our designed model R2DM, we also introduce a flexible LiDAR completion pipeline using the powerful properties of DDPMs. We demonstrate that our method outperforms the baselines on the generation task of KITTI-360 and KITTI-Raw datasets and the upsampling task of KITTI-360 datasets. Our code and pre-trained weights will be available at https://github.com/kazuto1011/r2dm.
MCPA: Multi-scale Cross Perceptron Attention Network for 2D Medical Image Segmentation
Jul 27, 2023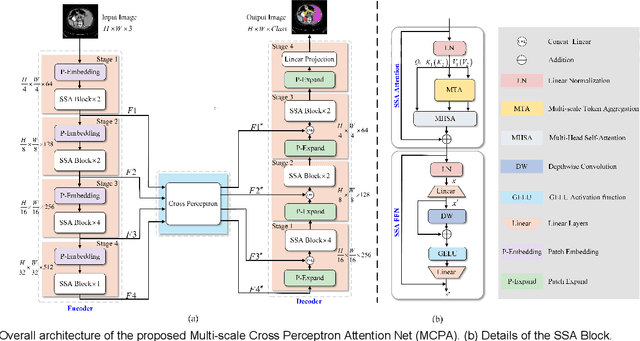
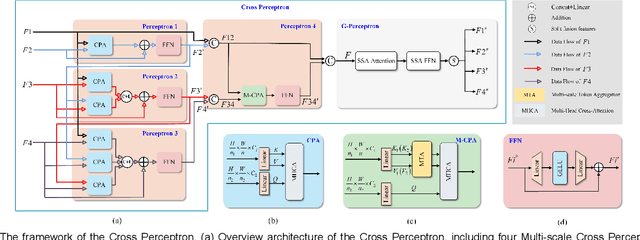
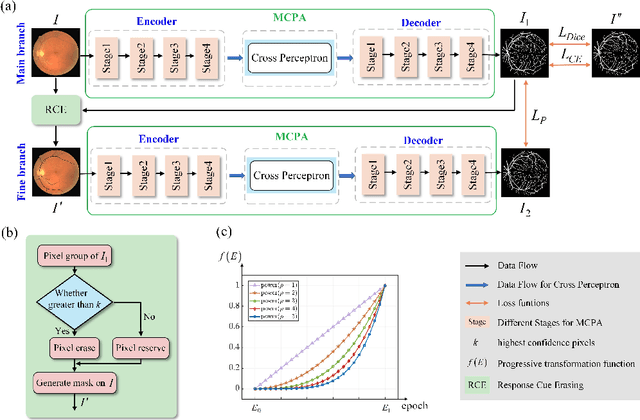
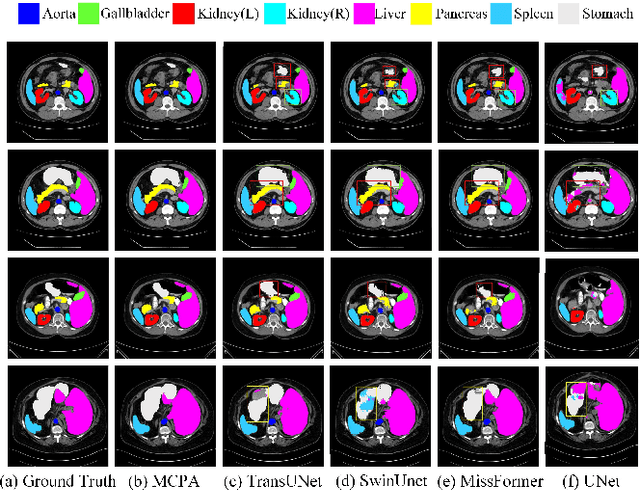
The UNet architecture, based on Convolutional Neural Networks (CNN), has demonstrated its remarkable performance in medical image analysis. However, it faces challenges in capturing long-range dependencies due to the limited receptive fields and inherent bias of convolutional operations. Recently, numerous transformer-based techniques have been incorporated into the UNet architecture to overcome this limitation by effectively capturing global feature correlations. However, the integration of the Transformer modules may result in the loss of local contextual information during the global feature fusion process. To overcome these challenges, we propose a 2D medical image segmentation model called Multi-scale Cross Perceptron Attention Network (MCPA). The MCPA consists of three main components: an encoder, a decoder, and a Cross Perceptron. The Cross Perceptron first captures the local correlations using multiple Multi-scale Cross Perceptron modules, facilitating the fusion of features across scales. The resulting multi-scale feature vectors are then spatially unfolded, concatenated, and fed through a Global Perceptron module to model global dependencies. Furthermore, we introduce a Progressive Dual-branch Structure to address the semantic segmentation of the image involving finer tissue structures. This structure gradually shifts the segmentation focus of MCPA network training from large-scale structural features to more sophisticated pixel-level features. We evaluate our proposed MCPA model on several publicly available medical image datasets from different tasks and devices, including the open large-scale dataset of CT (Synapse), MRI (ACDC), fundus camera (DRIVE, CHASE_DB1, HRF), and OCTA (ROSE). The experimental results show that our MCPA model achieves state-of-the-art performance. The code is available at https://github.com/simonustc/MCPA-for-2D-Medical-Image-Segmentation.
EP2P-Loc: End-to-End 3D Point to 2D Pixel Localization for Large-Scale Visual Localization
Sep 14, 2023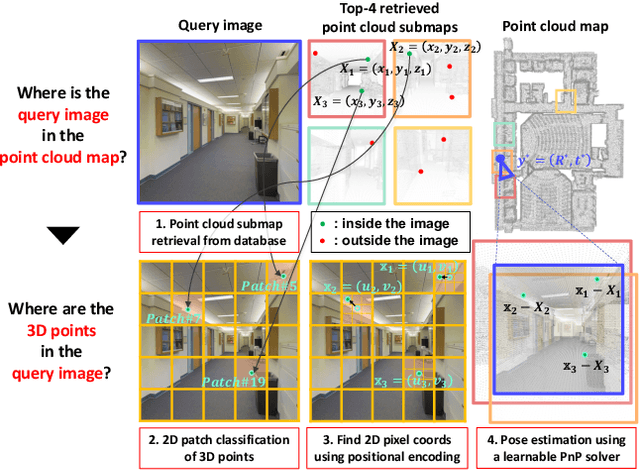
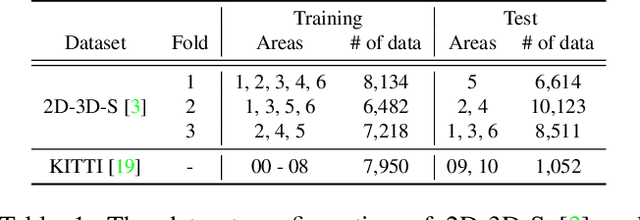
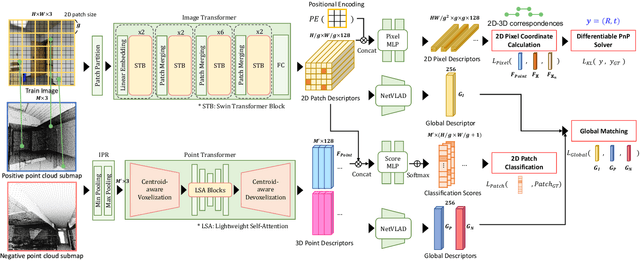
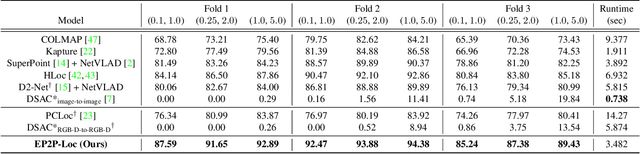
Visual localization is the task of estimating a 6-DoF camera pose of a query image within a provided 3D reference map. Thanks to recent advances in various 3D sensors, 3D point clouds are becoming a more accurate and affordable option for building the reference map, but research to match the points of 3D point clouds with pixels in 2D images for visual localization remains challenging. Existing approaches that jointly learn 2D-3D feature matching suffer from low inliers due to representational differences between the two modalities, and the methods that bypass this problem into classification have an issue of poor refinement. In this work, we propose EP2P-Loc, a novel large-scale visual localization method that mitigates such appearance discrepancy and enables end-to-end training for pose estimation. To increase the number of inliers, we propose a simple algorithm to remove invisible 3D points in the image, and find all 2D-3D correspondences without keypoint detection. To reduce memory usage and search complexity, we take a coarse-to-fine approach where we extract patch-level features from 2D images, then perform 2D patch classification on each 3D point, and obtain the exact corresponding 2D pixel coordinates through positional encoding. Finally, for the first time in this task, we employ a differentiable PnP for end-to-end training. In the experiments on newly curated large-scale indoor and outdoor benchmarks based on 2D-3D-S and KITTI, we show that our method achieves the state-of-the-art performance compared to existing visual localization and image-to-point cloud registration methods.
Neural scaling laws for phenotypic drug discovery
Sep 28, 2023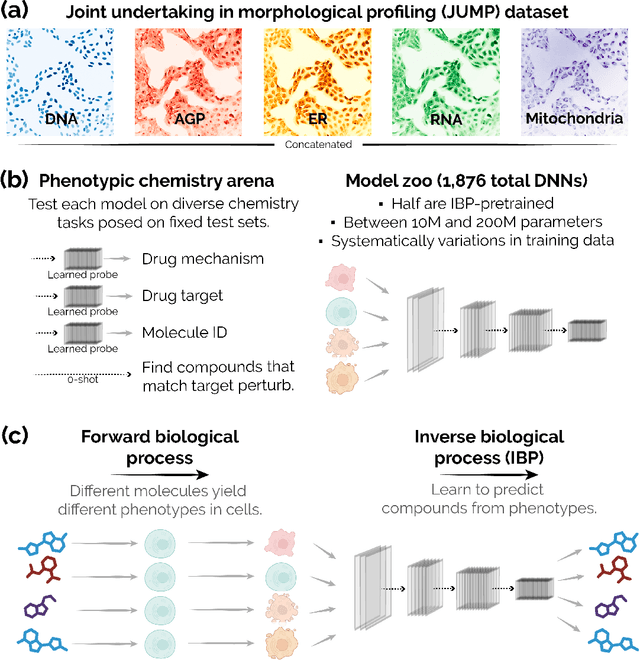
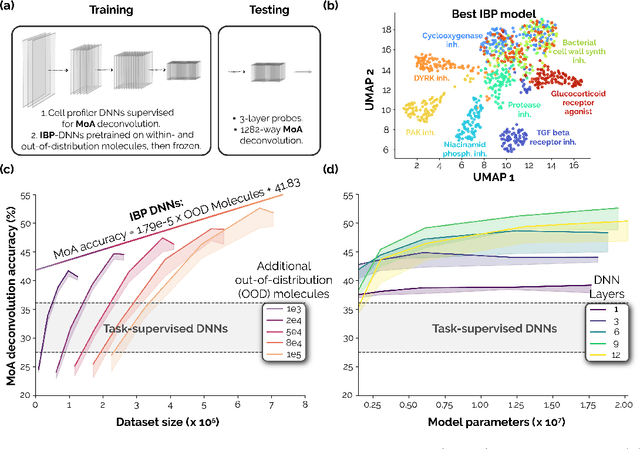
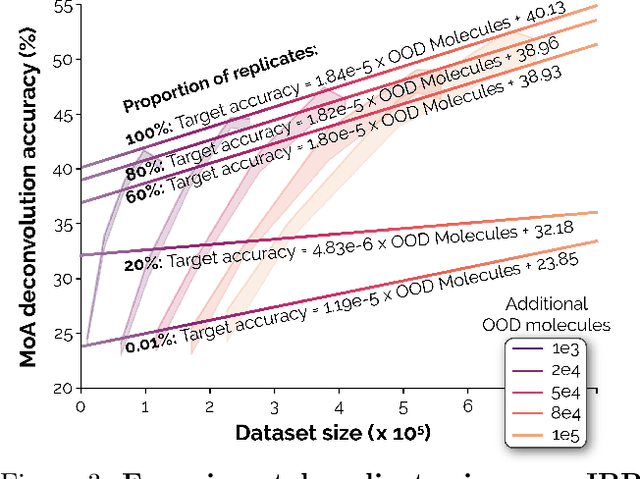
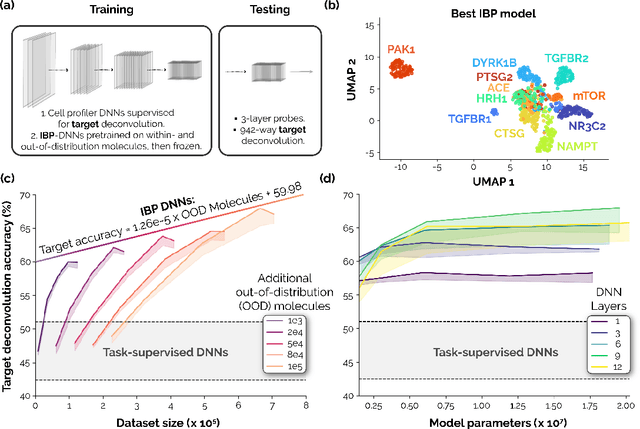
Recent breakthroughs by deep neural networks (DNNs) in natural language processing (NLP) and computer vision have been driven by a scale-up of models and data rather than the discovery of novel computing paradigms. Here, we investigate if scale can have a similar impact for models designed to aid small molecule drug discovery. We address this question through a large-scale and systematic analysis of how DNN size, data diet, and learning routines interact to impact accuracy on our Phenotypic Chemistry Arena (Pheno-CA) benchmark: a diverse set of drug development tasks posed on image-based high content screening data. Surprisingly, we find that DNNs explicitly supervised to solve tasks in the Pheno-CA do not continuously improve as their data and model size is scaled-up. To address this issue, we introduce a novel precursor task, the Inverse Biological Process (IBP), which is designed to resemble the causal objective functions that have proven successful for NLP. We indeed find that DNNs first trained with IBP then probed for performance on the Pheno-CA significantly outperform task-supervised DNNs. More importantly, the performance of these IBP-trained DNNs monotonically improves with data and model scale. Our findings reveal that the DNN ingredients needed to accurately solve small molecule drug development tasks are already in our hands, and project how much more experimental data is needed to achieve any desired level of improvement. We release our Pheno-CA benchmark and code to encourage further study of neural scaling laws for small molecule drug discovery.
A Survey on Deep Learning in Medical Image Registration: New Technologies, Uncertainty, Evaluation Metrics, and Beyond
Jul 28, 2023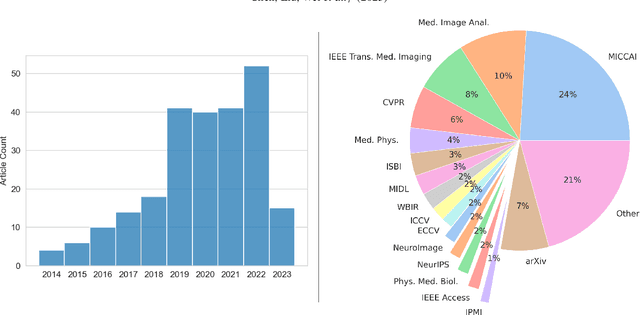
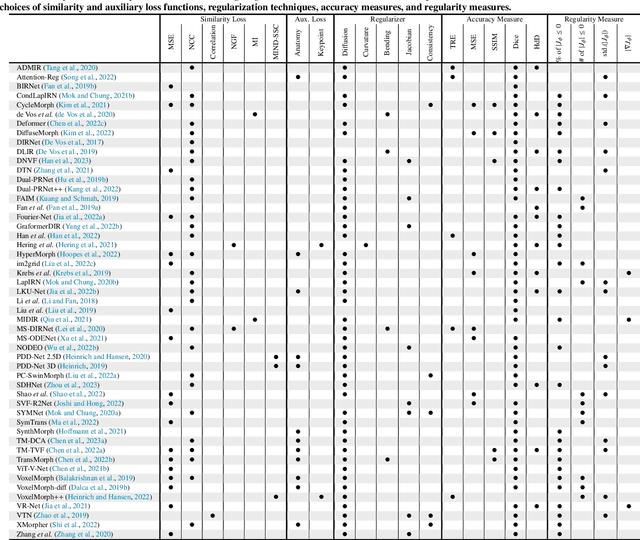
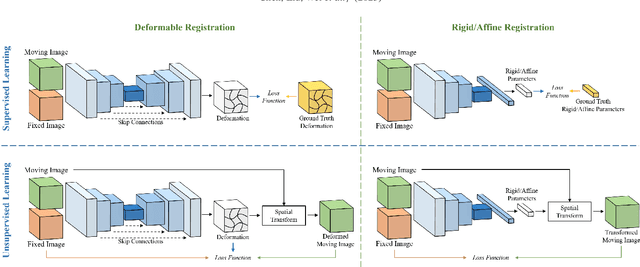

Over the past decade, deep learning technologies have greatly advanced the field of medical image registration. The initial developments, such as ResNet-based and U-Net-based networks, laid the groundwork for deep learning-driven image registration. Subsequent progress has been made in various aspects of deep learning-based registration, including similarity measures, deformation regularizations, and uncertainty estimation. These advancements have not only enriched the field of deformable image registration but have also facilitated its application in a wide range of tasks, including atlas construction, multi-atlas segmentation, motion estimation, and 2D-3D registration. In this paper, we present a comprehensive overview of the most recent advancements in deep learning-based image registration. We begin with a concise introduction to the core concepts of deep learning-based image registration. Then, we delve into innovative network architectures, loss functions specific to registration, and methods for estimating registration uncertainty. Additionally, this paper explores appropriate evaluation metrics for assessing the performance of deep learning models in registration tasks. Finally, we highlight the practical applications of these novel techniques in medical imaging and discuss the future prospects of deep learning-based image registration.
MIMIC: Masked Image Modeling with Image Correspondences
Jun 28, 2023
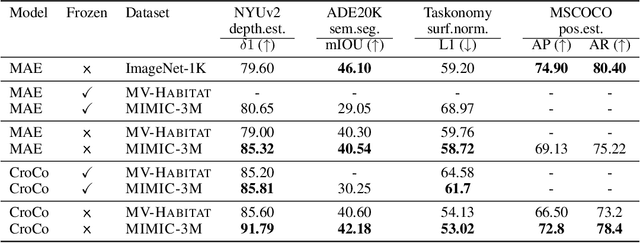

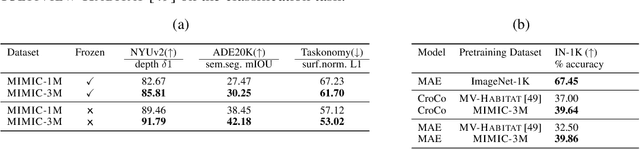
Many pixelwise dense prediction tasks-depth estimation and semantic segmentation in computer vision today rely on pretrained image representations. Therefore, curating effective pretraining datasets is vital. Unfortunately, the effective pretraining datasets are those with multi-view scenes and have only been curated using annotated 3D meshes, point clouds, and camera parameters from simulated environments. We propose a dataset-curation mechanism that does not require any annotations. We mine two datasets: MIMIC-1M with 1.3M and MIMIC-3M with 3.1M multi-view image pairs from open-sourced video datasets and from synthetic 3D environments. We train multiple self-supervised models with different masked image modeling objectives to showcase the following findings: Representations trained on MIMIC-3M outperform those mined using annotations on multiple downstream tasks, including depth estimation, semantic segmentation, surface normals, and pose estimation. They also outperform representations that are frozen and when downstream training data is limited to few-shot. Larger dataset (MIMIC-3M) significantly improves performance, which is promising since our curation method can arbitrarily scale to produce even larger datasets. MIMIC code, dataset, and pretrained models are open-sourced at https://github.com/RAIVNLab/MIMIC.
PromptMagician: Interactive Prompt Engineering for Text-to-Image Creation
Jul 18, 2023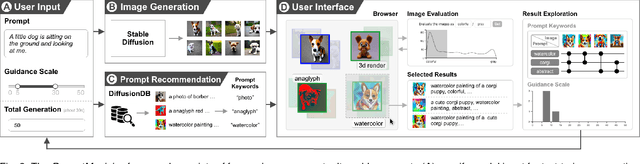
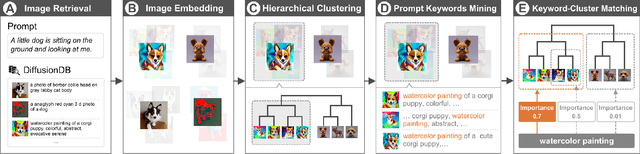
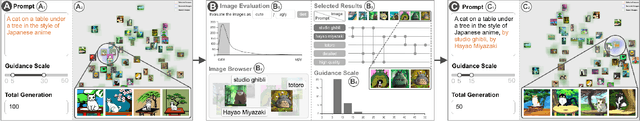

Generative text-to-image models have gained great popularity among the public for their powerful capability to generate high-quality images based on natural language prompts. However, developing effective prompts for desired images can be challenging due to the complexity and ambiguity of natural language. This research proposes PromptMagician, a visual analysis system that helps users explore the image results and refine the input prompts. The backbone of our system is a prompt recommendation model that takes user prompts as input, retrieves similar prompt-image pairs from DiffusionDB, and identifies special (important and relevant) prompt keywords. To facilitate interactive prompt refinement, PromptMagician introduces a multi-level visualization for the cross-modal embedding of the retrieved images and recommended keywords, and supports users in specifying multiple criteria for personalized exploration. Two usage scenarios, a user study, and expert interviews demonstrate the effectiveness and usability of our system, suggesting it facilitates prompt engineering and improves the creativity support of the generative text-to-image model.
Multiple Noises in Diffusion Model for Semi-Supervised Multi-Domain Translation
Sep 25, 2023Domain-to-domain translation involves generating a target domain sample given a condition in the source domain. Most existing methods focus on fixed input and output domains, i.e. they only work for specific configurations (i.e. for two domains, either $D_1\rightarrow{}D_2$ or $D_2\rightarrow{}D_1$). This paper proposes Multi-Domain Diffusion (MDD), a conditional diffusion framework for multi-domain translation in a semi-supervised context. Unlike previous methods, MDD does not require defining input and output domains, allowing translation between any partition of domains within a set (such as $(D_1, D_2)\rightarrow{}D_3$, $D_2\rightarrow{}(D_1, D_3)$, $D_3\rightarrow{}D_1$, etc. for 3 domains), without the need to train separate models for each domain configuration. The key idea behind MDD is to leverage the noise formulation of diffusion models by incorporating one noise level per domain, which allows missing domains to be modeled with noise in a natural way. This transforms the training task from a simple reconstruction task to a domain translation task, where the model relies on less noisy domains to reconstruct more noisy domains. We present results on a multi-domain (with more than two domains) synthetic image translation dataset with challenging semantic domain inversion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge