"Image": models, code, and papers
ADU-Depth: Attention-based Distillation with Uncertainty Modeling for Depth Estimation
Sep 26, 2023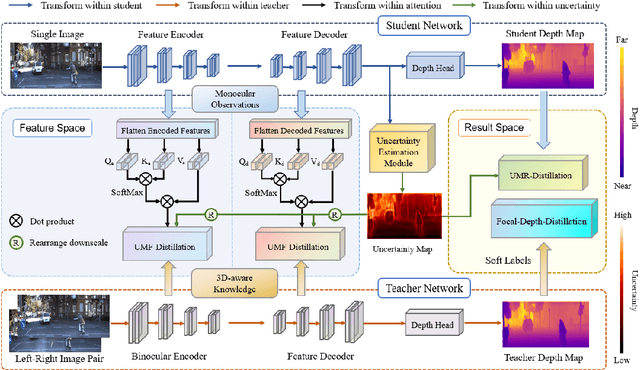
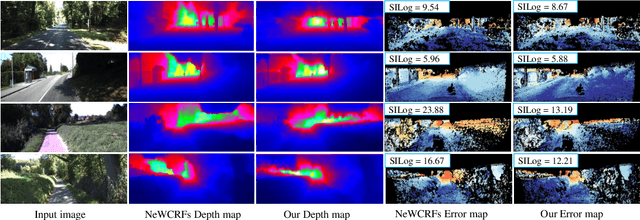
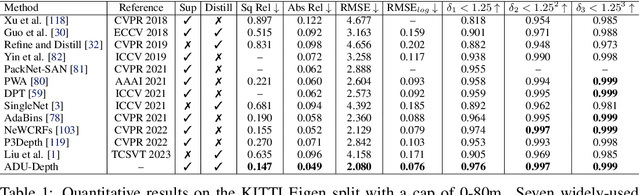
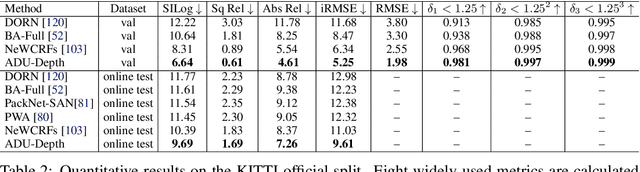

Monocular depth estimation is challenging due to its inherent ambiguity and ill-posed nature, yet it is quite important to many applications. While recent works achieve limited accuracy by designing increasingly complicated networks to extract features with limited spatial geometric cues from a single RGB image, we intend to introduce spatial cues by training a teacher network that leverages left-right image pairs as inputs and transferring the learned 3D geometry-aware knowledge to the monocular student network. Specifically, we present a novel knowledge distillation framework, named ADU-Depth, with the goal of leveraging the well-trained teacher network to guide the learning of the student network, thus boosting the precise depth estimation with the help of extra spatial scene information. To enable domain adaptation and ensure effective and smooth knowledge transfer from teacher to student, we apply both attention-adapted feature distillation and focal-depth-adapted response distillation in the training stage. In addition, we explicitly model the uncertainty of depth estimation to guide distillation in both feature space and result space to better produce 3D-aware knowledge from monocular observations and thus enhance the learning for hard-to-predict image regions. Our extensive experiments on the real depth estimation datasets KITTI and DrivingStereo demonstrate the effectiveness of the proposed method, which ranked 1st on the challenging KITTI online benchmark.
Improving Underwater Visual Tracking With a Large Scale Dataset and Image Enhancement
Aug 31, 2023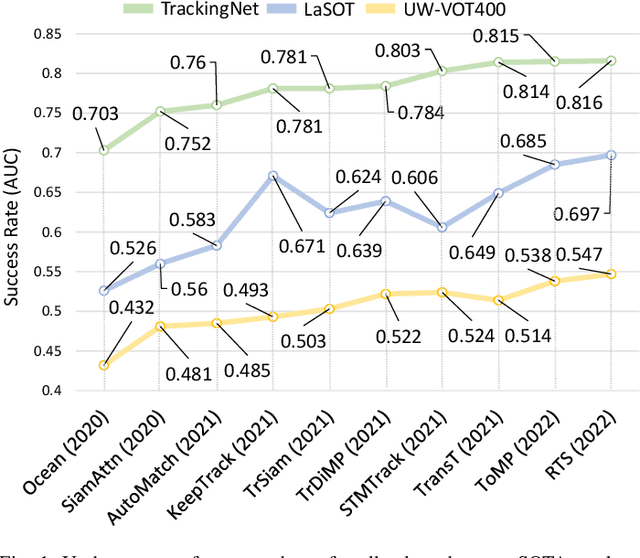
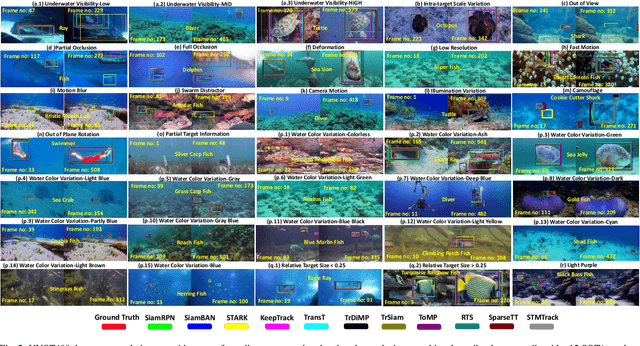
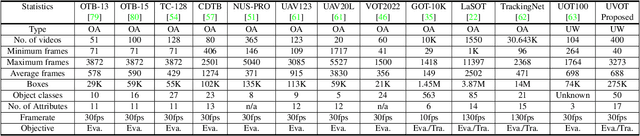

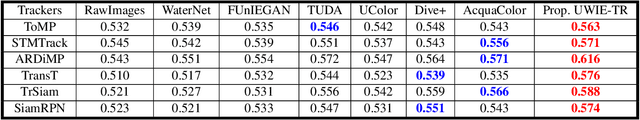
This paper presents a new dataset and general tracker enhancement method for Underwater Visual Object Tracking (UVOT). Despite its significance, underwater tracking has remained unexplored due to data inaccessibility. It poses distinct challenges; the underwater environment exhibits non-uniform lighting conditions, low visibility, lack of sharpness, low contrast, camouflage, and reflections from suspended particles. Performance of traditional tracking methods designed primarily for terrestrial or open-air scenarios drops in such conditions. We address the problem by proposing a novel underwater image enhancement algorithm designed specifically to boost tracking quality. The method has resulted in a significant performance improvement, of up to 5.0% AUC, of state-of-the-art (SOTA) visual trackers. To develop robust and accurate UVOT methods, large-scale datasets are required. To this end, we introduce a large-scale UVOT benchmark dataset consisting of 400 video segments and 275,000 manually annotated frames enabling underwater training and evaluation of deep trackers. The videos are labelled with several underwater-specific tracking attributes including watercolor variation, target distractors, camouflage, target relative size, and low visibility conditions. The UVOT400 dataset, tracking results, and the code are publicly available on: https://github.com/BasitAlawode/UWVOT400.
Consistency Regularization Improves Placenta Segmentation in Fetal EPI MRI Time Series
Oct 05, 2023The placenta plays a crucial role in fetal development. Automated 3D placenta segmentation from fetal EPI MRI holds promise for advancing prenatal care. This paper proposes an effective semi-supervised learning method for improving placenta segmentation in fetal EPI MRI time series. We employ consistency regularization loss that promotes consistency under spatial transformation of the same image and temporal consistency across nearby images in a time series. The experimental results show that the method improves the overall segmentation accuracy and provides better performance for outliers and hard samples. The evaluation also indicates that our method improves the temporal coherency of the prediction, which could lead to more accurate computation of temporal placental biomarkers. This work contributes to the study of the placenta and prenatal clinical decision-making. Code is available at https://github.com/firstmover/cr-seg.
Coloring Deep CNN Layers with Activation Hue Loss
Oct 05, 2023This paper proposes a novel hue-like angular parameter to model the structure of deep convolutional neural network (CNN) activation space, referred to as the {\em activation hue}, for the purpose of regularizing models for more effective learning. The activation hue generalizes the notion of color hue angle in standard 3-channel RGB intensity space to $N$-channel activation space. A series of observations based on nearest neighbor indexing of activation vectors with pre-trained networks indicate that class-informative activations are concentrated about an angle $\theta$ in both the $(x,y)$ image plane and in multi-channel activation space. A regularization term in the form of hue-like angular $\theta$ labels is proposed to complement standard one-hot loss. Training from scratch using combined one-hot + activation hue loss improves classification performance modestly for a wide variety of classification tasks, including ImageNet.
Watch Your Steps: Local Image and Scene Editing by Text Instructions
Aug 17, 2023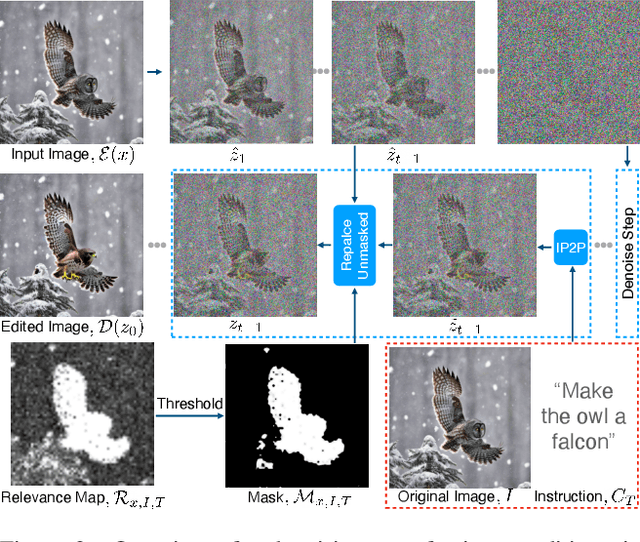
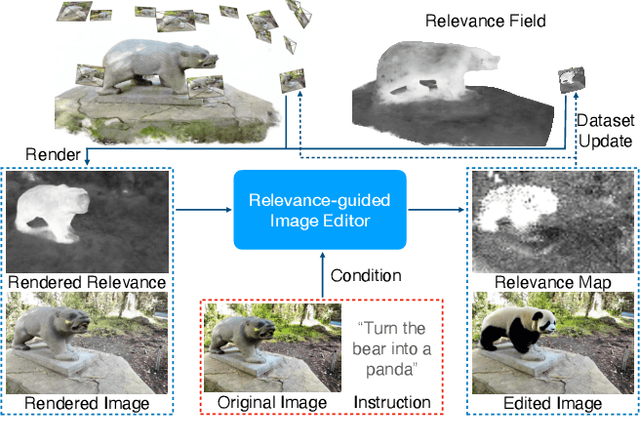
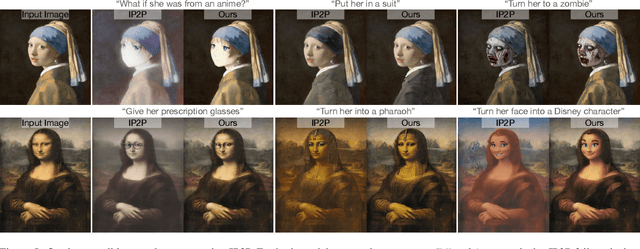

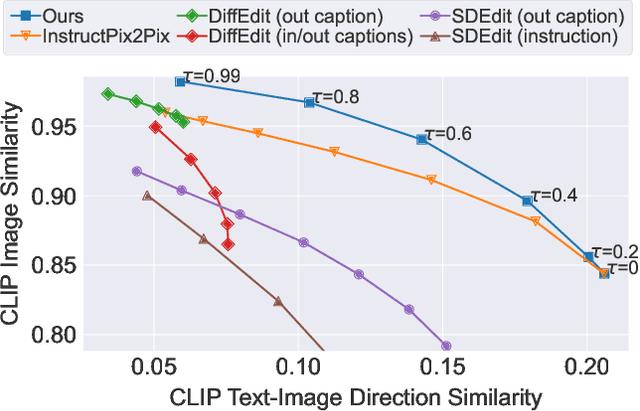

Denoising diffusion models have enabled high-quality image generation and editing. We present a method to localize the desired edit region implicit in a text instruction. We leverage InstructPix2Pix (IP2P) and identify the discrepancy between IP2P predictions with and without the instruction. This discrepancy is referred to as the relevance map. The relevance map conveys the importance of changing each pixel to achieve the edits, and is used to to guide the modifications. This guidance ensures that the irrelevant pixels remain unchanged. Relevance maps are further used to enhance the quality of text-guided editing of 3D scenes in the form of neural radiance fields. A field is trained on relevance maps of training views, denoted as the relevance field, defining the 3D region within which modifications should be made. We perform iterative updates on the training views guided by rendered relevance maps from the relevance field. Our method achieves state-of-the-art performance on both image and NeRF editing tasks. Project page: https://ashmrz.github.io/WatchYourSteps/
Exploring Transfer Learning in Medical Image Segmentation using Vision-Language Models
Aug 15, 2023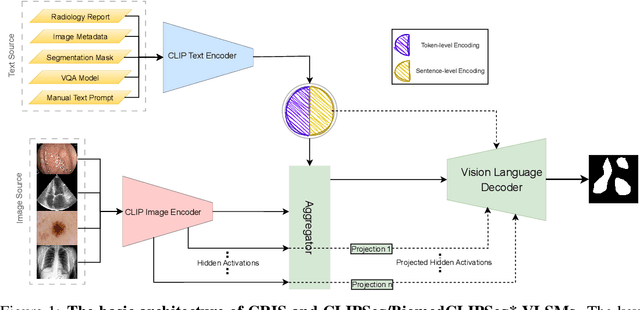
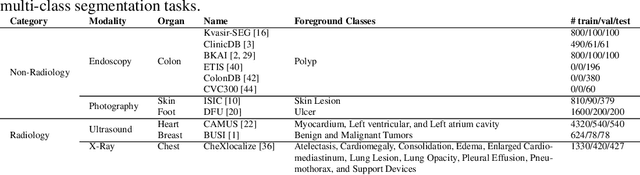
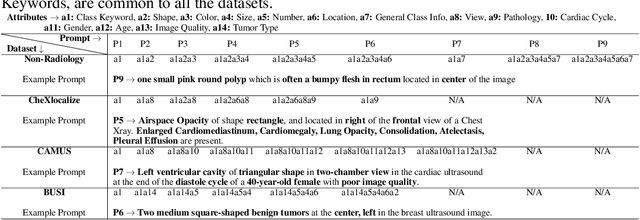


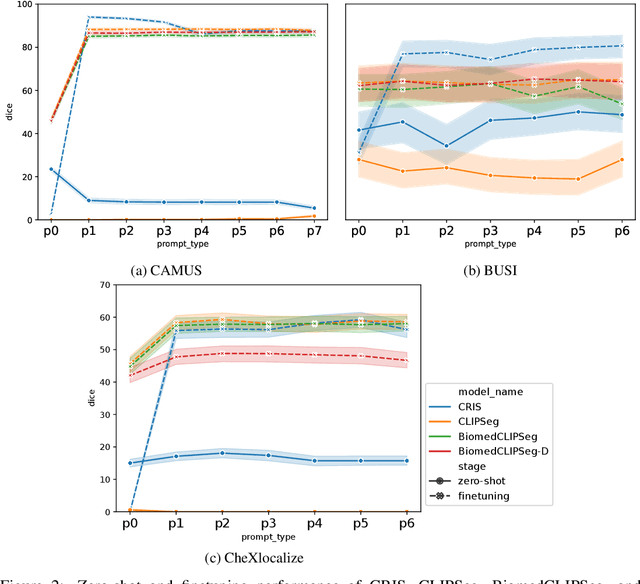
Medical Image Segmentation is crucial in various clinical applications within the medical domain. While state-of-the-art segmentation models have proven effective, integrating textual guidance to enhance visual features for this task remains an area with limited progress. Existing segmentation models that utilize textual guidance are primarily trained on open-domain images, raising concerns about their direct applicability in the medical domain without manual intervention or fine-tuning. To address these challenges, we propose using multimodal vision-language models for capturing semantic information from image descriptions and images, enabling the segmentation of diverse medical images. This study comprehensively evaluates existing vision language models across multiple datasets to assess their transferability from the open domain to the medical field. Furthermore, we introduce variations of image descriptions for previously unseen images in the dataset, revealing notable variations in model performance based on the generated prompts. Our findings highlight the distribution shift between the open-domain images and the medical domain and show that the segmentation models trained on open-domain images are not directly transferrable to the medical field. But their performance can be increased by finetuning them in the medical datasets. We report the zero-shot and finetuned segmentation performance of 4 Vision Language Models (VLMs) on 11 medical datasets using 9 types of prompts derived from 14 attributes.
IIHT: Medical Report Generation with Image-to-Indicator Hierarchical Transformer
Aug 10, 2023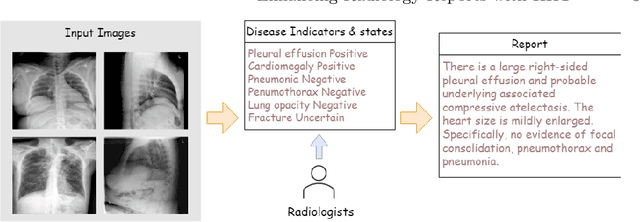
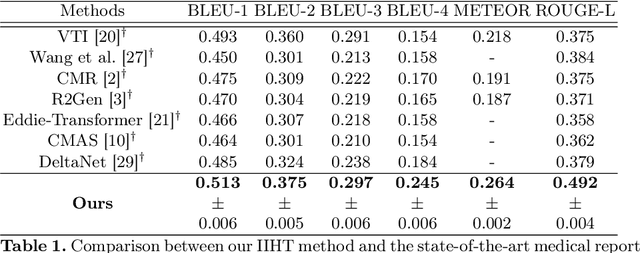
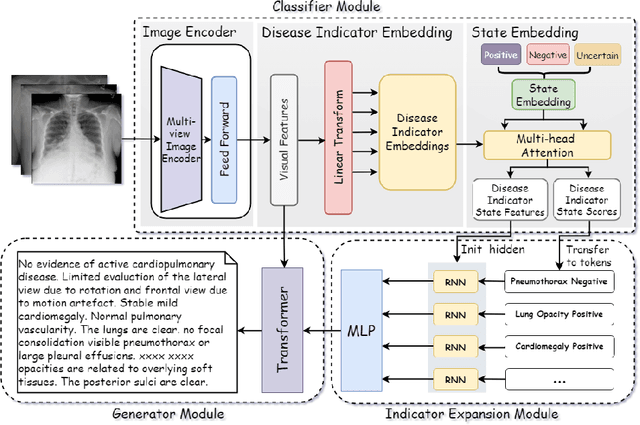
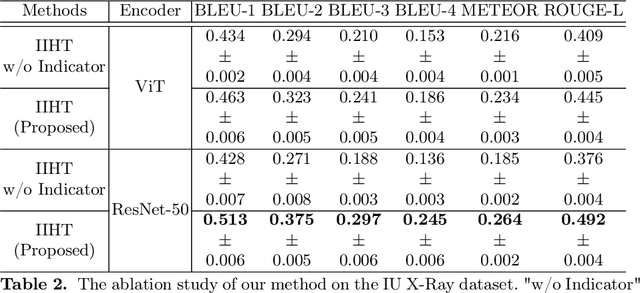
Automated medical report generation has become increasingly important in medical analysis. It can produce computer-aided diagnosis descriptions and thus significantly alleviate the doctors' work. Inspired by the huge success of neural machine translation and image captioning, various deep learning methods have been proposed for medical report generation. However, due to the inherent properties of medical data, including data imbalance and the length and correlation between report sequences, the generated reports by existing methods may exhibit linguistic fluency but lack adequate clinical accuracy. In this work, we propose an image-to-indicator hierarchical transformer (IIHT) framework for medical report generation. It consists of three modules, i.e., a classifier module, an indicator expansion module and a generator module. The classifier module first extracts image features from the input medical images and produces disease-related indicators with their corresponding states. The disease-related indicators are subsequently utilised as input for the indicator expansion module, incorporating the "data-text-data" strategy. The transformer-based generator then leverages these extracted features along with image features as auxiliary information to generate final reports. Furthermore, the proposed IIHT method is feasible for radiologists to modify disease indicators in real-world scenarios and integrate the operations into the indicator expansion module for fluent and accurate medical report generation. Extensive experiments and comparisons with state-of-the-art methods under various evaluation metrics demonstrate the great performance of the proposed method.
Impact of architecture on robustness and interpretability of multispectral deep neural networks
Sep 27, 2023Including information from additional spectral bands (e.g., near-infrared) can improve deep learning model performance for many vision-oriented tasks. There are many possible ways to incorporate this additional information into a deep learning model, but the optimal fusion strategy has not yet been determined and can vary between applications. At one extreme, known as "early fusion," additional bands are stacked as extra channels to obtain an input image with more than three channels. At the other extreme, known as "late fusion," RGB and non-RGB bands are passed through separate branches of a deep learning model and merged immediately before a final classification or segmentation layer. In this work, we characterize the performance of a suite of multispectral deep learning models with different fusion approaches, quantify their relative reliance on different input bands and evaluate their robustness to naturalistic image corruptions affecting one or more input channels.
Diagnosis of Helicobacter pylori using AutoEncoders for the Detection of Anomalous Staining Patterns in Immunohistochemistry Images
Sep 27, 2023This work addresses the detection of Helicobacter pylori a bacterium classified since 1994 as class 1 carcinogen to humans. By its highest specificity and sensitivity, the preferred diagnosis technique is the analysis of histological images with immunohistochemical staining, a process in which certain stained antibodies bind to antigens of the biological element of interest. This analysis is a time demanding task, which is currently done by an expert pathologist that visually inspects the digitized samples. We propose to use autoencoders to learn latent patterns of healthy tissue and detect H. pylori as an anomaly in image staining. Unlike existing classification approaches, an autoencoder is able to learn patterns in an unsupervised manner (without the need of image annotations) with high performance. In particular, our model has an overall 91% of accuracy with 86\% sensitivity, 96% specificity and 0.97 AUC in the detection of H. pylori.
Development of a Novel Quantum Pre-processing Filter to Improve Image Classification Accuracy of Neural Network Models
Aug 22, 2023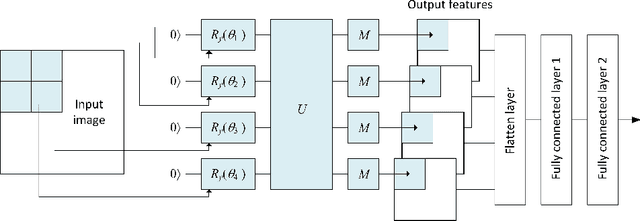
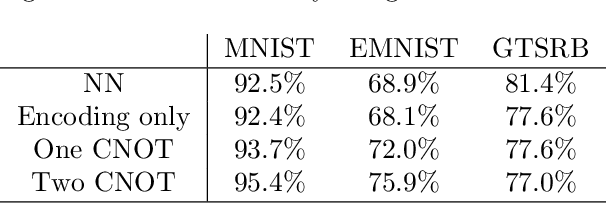
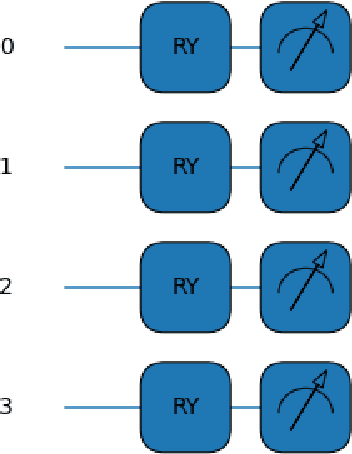
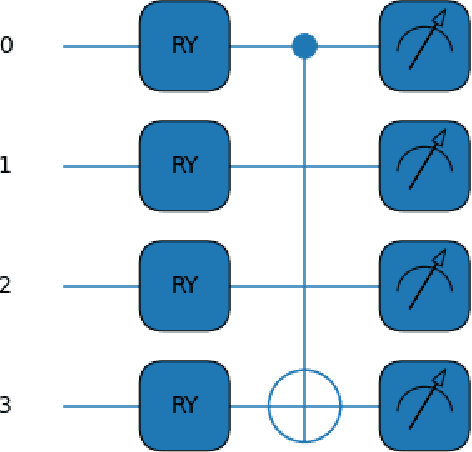
This paper proposes a novel quantum pre-processing filter (QPF) to improve the image classification accuracy of neural network (NN) models. A simple four qubit quantum circuit that uses Y rotation gates for encoding and two controlled NOT gates for creating correlation among the qubits is applied as a feature extraction filter prior to passing data into the fully connected NN architecture. By applying the QPF approach, the results show that the image classification accuracy based on the MNIST (handwritten 10 digits) and the EMNIST (handwritten 47 class digits and letters) datasets can be improved, from 92.5% to 95.4% and from 68.9% to 75.9%, respectively. These improvements were obtained without introducing extra model parameters or optimizations in the machine learning process. However, tests performed on the developed QPF approach against a relatively complex GTSRB dataset with 43 distinct class real-life traffic sign images showed a degradation in the classification accuracy. Considering this result, further research into the understanding and the design of a more suitable quantum circuit approach for image classification neural networks could be explored utilizing the baseline method proposed in this paper.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge