"Image": models, code, and papers
Improving mitosis detection on histopathology images using large vision-language models
Oct 11, 2023In certain types of cancerous tissue, mitotic count has been shown to be associated with tumor proliferation, poor prognosis, and therapeutic resistance. Due to the high inter-rater variability of mitotic counting by pathologists, convolutional neural networks (CNNs) have been employed to reduce the subjectivity of mitosis detection in hematoxylin and eosin (H&E)-stained whole slide images. However, most existing models have performance that lags behind expert panel review and only incorporate visual information. In this work, we demonstrate that pre-trained large-scale vision-language models that leverage both visual features and natural language improve mitosis detection accuracy. We formulate the mitosis detection task as an image captioning task and a visual question answering (VQA) task by including metadata such as tumor and scanner types as context. The effectiveness of our pipeline is demonstrated via comparison with various baseline models using 9,501 mitotic figures and 11,051 hard negatives (non-mitotic figures that are difficult to characterize) from the publicly available Mitosis Domain Generalization Challenge (MIDOG22) dataset.
Microscopy Image Segmentation via Point and Shape Regularized Data Synthesis
Aug 18, 2023Current deep learning-based approaches for the segmentation of microscopy images heavily rely on large amount of training data with dense annotation, which is highly costly and laborious in practice. Compared to full annotation where the complete contour of objects is depicted, point annotations, specifically object centroids, are much easier to acquire and still provide crucial information about the objects for subsequent segmentation. In this paper, we assume access to point annotations only during training and develop a unified pipeline for microscopy image segmentation using synthetically generated training data. Our framework includes three stages: (1) it takes point annotations and samples a pseudo dense segmentation mask constrained with shape priors; (2) with an image generative model trained in an unpaired manner, it translates the mask to a realistic microscopy image regularized by object level consistency; (3) the pseudo masks along with the synthetic images then constitute a pairwise dataset for training an ad-hoc segmentation model. On the public MoNuSeg dataset, our synthesis pipeline produces more diverse and realistic images than baseline models while maintaining high coherence between input masks and generated images. When using the identical segmentation backbones, the models trained on our synthetic dataset significantly outperform those trained with pseudo-labels or baseline-generated images. Moreover, our framework achieves comparable results to models trained on authentic microscopy images with dense labels, demonstrating its potential as a reliable and highly efficient alternative to labor-intensive manual pixel-wise annotations in microscopy image segmentation. The code is available.
CMISR: Circular Medical Image Super-Resolution
Aug 15, 2023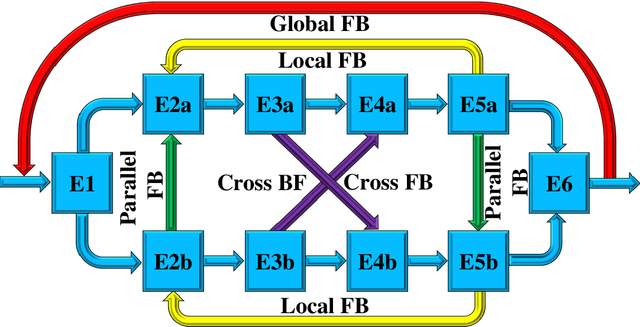
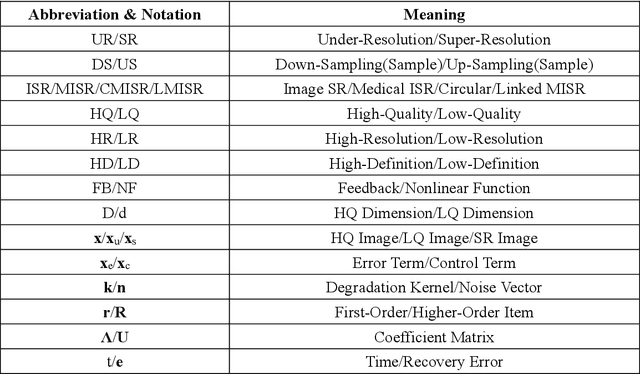
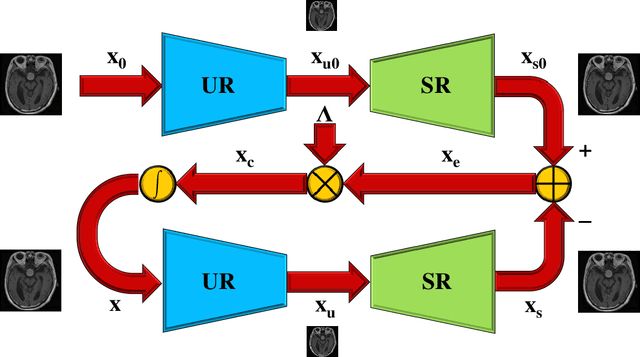
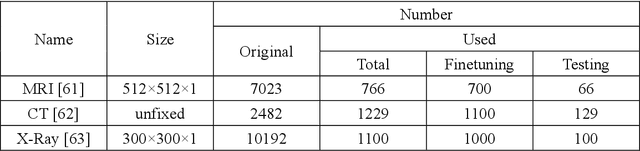
Classical methods of medical image super-resolution (MISR) utilize open-loop architecture with implicit under-resolution (UR) unit and explicit super-resolution (SR) unit. The UR unit can always be given, assumed, or estimated, while the SR unit is elaborately designed according to various SR algorithms. The closed-loop feedback mechanism is widely employed in current MISR approaches and can efficiently improve their performance. The feedback mechanism may be divided into two categories: local and global feedback. Therefore, this paper proposes a global feedback-based closed-cycle framework, circular MISR (CMISR), with unambiguous UR and SR elements. Mathematical model and closed-loop equation of CMISR are built. Mathematical proof with Taylor-series approximation indicates that CMISR has zero recovery error in steady-state. In addition, CMISR holds plug-and-play characteristic which can be established on any existing MISR algorithms. Five CMISR algorithms are respectively proposed based on the state-of-the-art open-loop MISR algorithms. Experimental results with three scale factors and on three open medical image datasets show that CMISR is superior to MISR in reconstruction performance and is particularly suited to medical images with strong edges or intense contrast.
Unveiling Fairness Biases in Deep Learning-Based Brain MRI Reconstruction
Sep 25, 2023Deep learning (DL) reconstruction particularly of MRI has led to improvements in image fidelity and reduction of acquisition time. In neuroimaging, DL methods can reconstruct high-quality images from undersampled data. However, it is essential to consider fairness in DL algorithms, particularly in terms of demographic characteristics. This study presents the first fairness analysis in a DL-based brain MRI reconstruction model. The model utilises the U-Net architecture for image reconstruction and explores the presence and sources of unfairness by implementing baseline Empirical Risk Minimisation (ERM) and rebalancing strategies. Model performance is evaluated using image reconstruction metrics. Our findings reveal statistically significant performance biases between the gender and age subgroups. Surprisingly, data imbalance and training discrimination are not the main sources of bias. This analysis provides insights of fairness in DL-based image reconstruction and aims to improve equity in medical AI applications.
Jurassic World Remake: Bringing Ancient Fossils Back to Life via Zero-Shot Long Image-to-Image Translation
Aug 14, 2023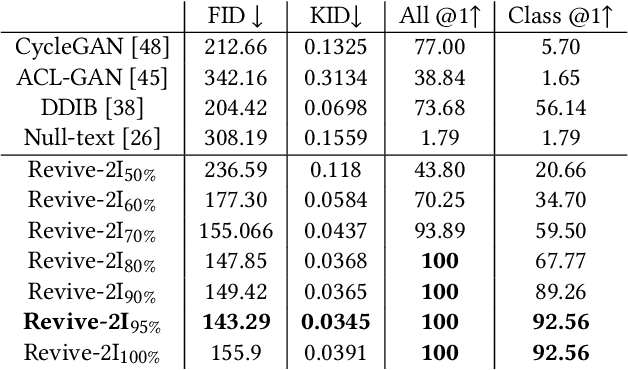
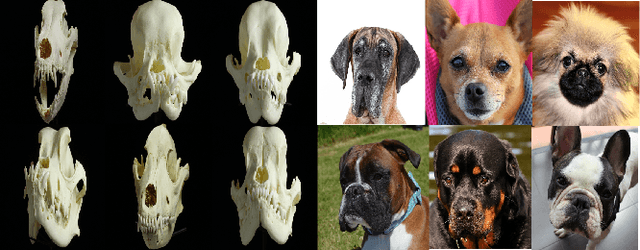
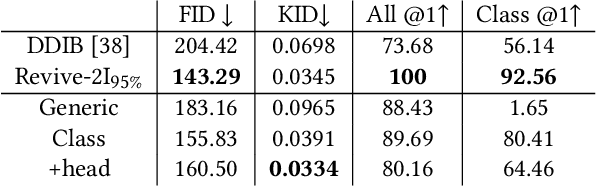
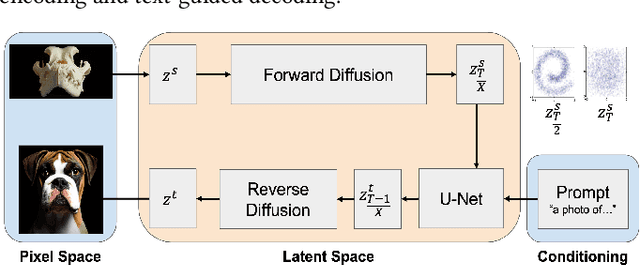
With a strong understanding of the target domain from natural language, we produce promising results in translating across large domain gaps and bringing skeletons back to life. In this work, we use text-guided latent diffusion models for zero-shot image-to-image translation (I2I) across large domain gaps (longI2I), where large amounts of new visual features and new geometry need to be generated to enter the target domain. Being able to perform translations across large domain gaps has a wide variety of real-world applications in criminology, astrology, environmental conservation, and paleontology. In this work, we introduce a new task Skull2Animal for translating between skulls and living animals. On this task, we find that unguided Generative Adversarial Networks (GANs) are not capable of translating across large domain gaps. Instead of these traditional I2I methods, we explore the use of guided diffusion and image editing models and provide a new benchmark model, Revive-2I, capable of performing zero-shot I2I via text-prompting latent diffusion models. We find that guidance is necessary for longI2I because, to bridge the large domain gap, prior knowledge about the target domain is needed. In addition, we find that prompting provides the best and most scalable information about the target domain as classifier-guided diffusion models require retraining for specific use cases and lack stronger constraints on the target domain because of the wide variety of images they are trained on.
Bi-Modality Medical Image Synthesis Using Semi-Supervised Sequential Generative Adversarial Networks
Aug 29, 2023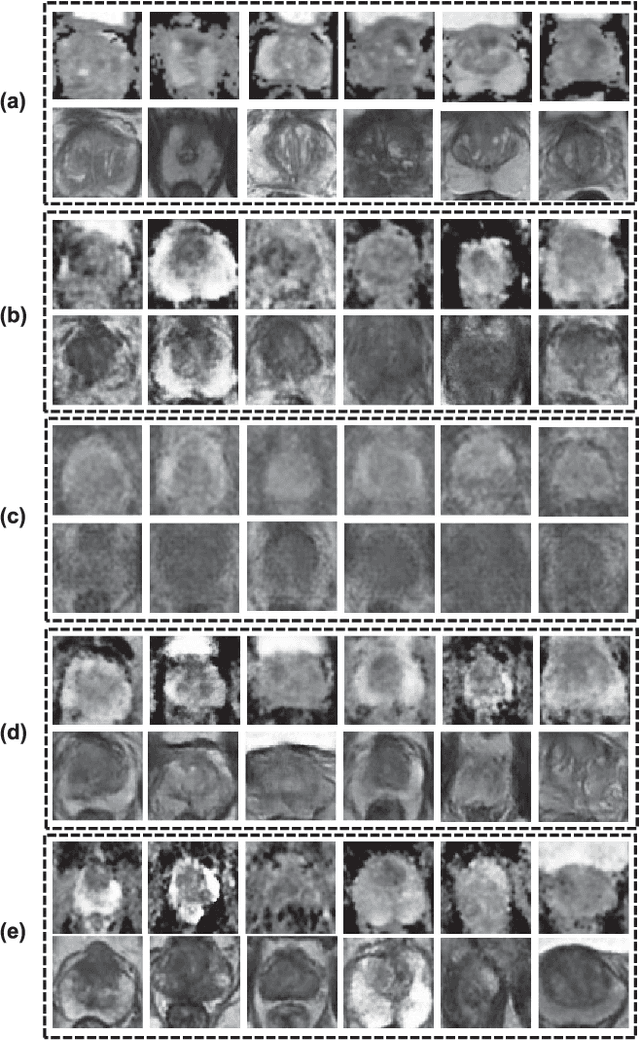
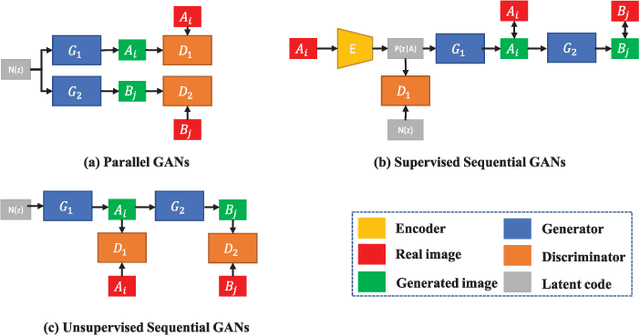
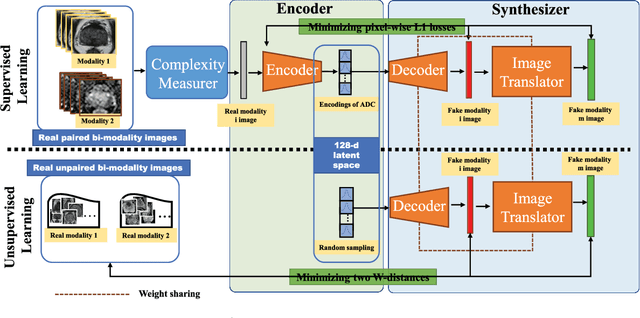
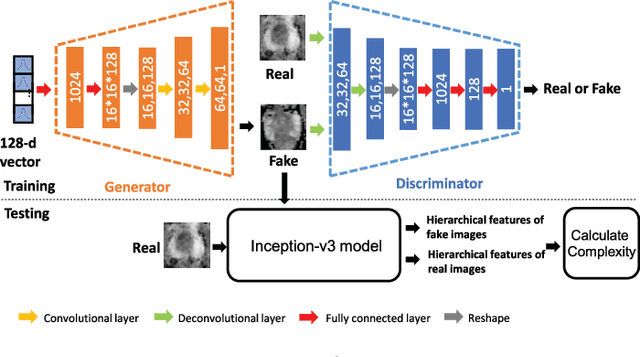
In this paper, we propose a bi-modality medical image synthesis approach based on sequential generative adversarial network (GAN) and semi-supervised learning. Our approach consists of two generative modules that synthesize images of the two modalities in a sequential order. A method for measuring the synthesis complexity is proposed to automatically determine the synthesis order in our sequential GAN. Images of the modality with a lower complexity are synthesized first, and the counterparts with a higher complexity are generated later. Our sequential GAN is trained end-to-end in a semi-supervised manner. In supervised training, the joint distribution of bi-modality images are learned from real paired images of the two modalities by explicitly minimizing the reconstruction losses between the real and synthetic images. To avoid overfitting limited training images, in unsupervised training, the marginal distribution of each modality is learned based on unpaired images by minimizing the Wasserstein distance between the distributions of real and fake images. We comprehensively evaluate the proposed model using two synthesis tasks based on three types of evaluate metrics and user studies. Visual and quantitative results demonstrate the superiority of our method to the state-of-the-art methods, and reasonable visual quality and clinical significance. Code is made publicly available at https://github.com/hustlinyi/Multimodal-Medical-Image-Synthesis.
I2SRM: Intra- and Inter-Sample Relationship Modeling for Multimodal Information Extraction
Oct 10, 2023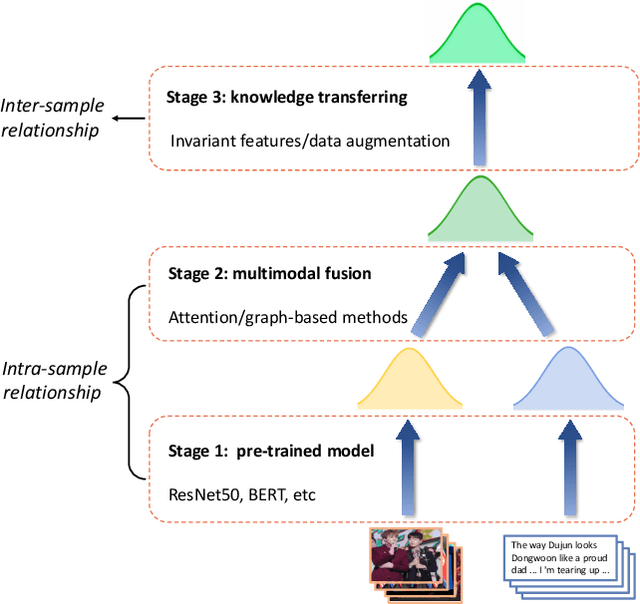
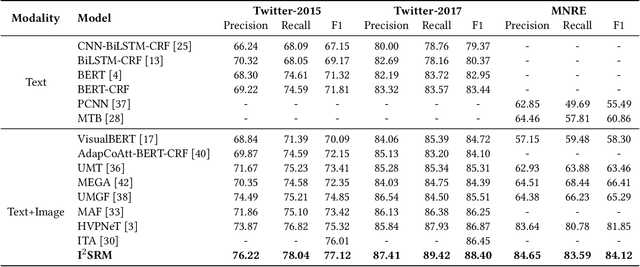
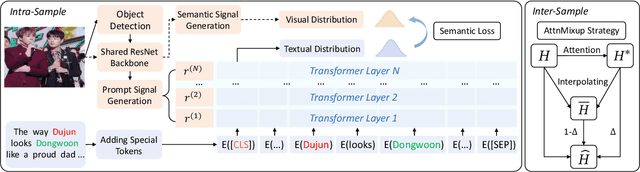
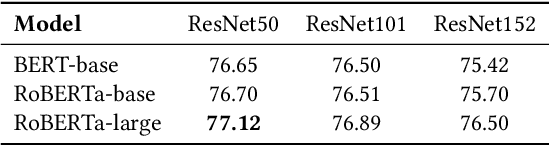
Multimodal information extraction is attracting research attention nowadays, which requires aggregating representations from different modalities. In this paper, we present the Intra- and Inter-Sample Relationship Modeling (I2SRM) method for this task, which contains two modules. Firstly, the intra-sample relationship modeling module operates on a single sample and aims to learn effective representations. Embeddings from textual and visual modalities are shifted to bridge the modality gap caused by distinct pre-trained language and image models. Secondly, the inter-sample relationship modeling module considers relationships among multiple samples and focuses on capturing the interactions. An AttnMixup strategy is proposed, which not only enables collaboration among samples but also augments data to improve generalization. We conduct extensive experiments on the multimodal named entity recognition datasets Twitter-2015 and Twitter-2017, and the multimodal relation extraction dataset MNRE. Our proposed method I2SRM achieves competitive results, 77.12% F1-score on Twitter-2015, 88.40% F1-score on Twitter-2017, and 84.12% F1-score on MNRE.
Topological RANSAC for instance verification and retrieval without fine-tuning
Oct 10, 2023This paper presents an innovative approach to enhancing explainable image retrieval, particularly in situations where a fine-tuning set is unavailable. The widely-used SPatial verification (SP) method, despite its efficacy, relies on a spatial model and the hypothesis-testing strategy for instance recognition, leading to inherent limitations, including the assumption of planar structures and neglect of topological relations among features. To address these shortcomings, we introduce a pioneering technique that replaces the spatial model with a topological one within the RANSAC process. We propose bio-inspired saccade and fovea functions to verify the topological consistency among features, effectively circumventing the issues associated with SP's spatial model. Our experimental results demonstrate that our method significantly outperforms SP, achieving state-of-the-art performance in non-fine-tuning retrieval. Furthermore, our approach can enhance performance when used in conjunction with fine-tuned features. Importantly, our method retains high explainability and is lightweight, offering a practical and adaptable solution for a variety of real-world applications.
Language-driven Open-Vocabulary Keypoint Detection for Animal Body and Face
Oct 10, 2023

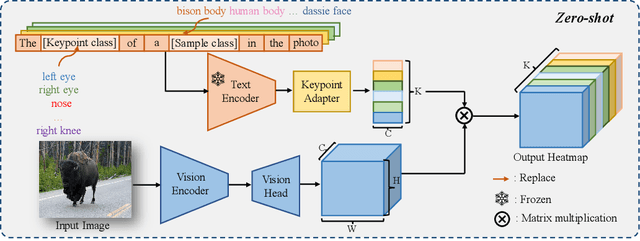
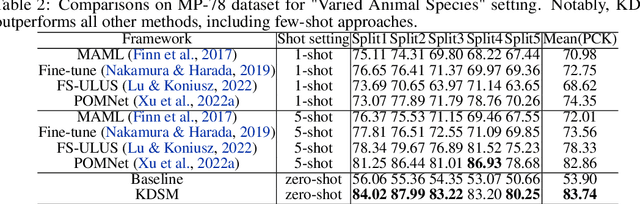
Current approaches for image-based keypoint detection on animal (including human) body and face are limited to specific keypoints and species. We address the limitation by proposing the Open-Vocabulary Keypoint Detection (OVKD) task. It aims to use text prompts to localize arbitrary keypoints of any species. To accomplish this objective, we propose Open-Vocabulary Keypoint Detection with Semantic-feature Matching (KDSM), which utilizes both vision and language models to harness the relationship between text and vision and thus achieve keypoint detection through associating text prompt with relevant keypoint features. Additionally, KDSM integrates domain distribution matrix matching and some special designs to reinforce the relationship between language and vision, thereby improving the model's generalizability and performance. Extensive experiments show that our proposed components bring significant performance improvements, and our overall method achieves impressive results in OVKD. Remarkably, our method outperforms the state-of-the-art few-shot keypoint detection methods using a zero-shot fashion. We will make the source code publicly accessible.
ObjectComposer: Consistent Generation of Multiple Objects Without Fine-tuning
Oct 10, 2023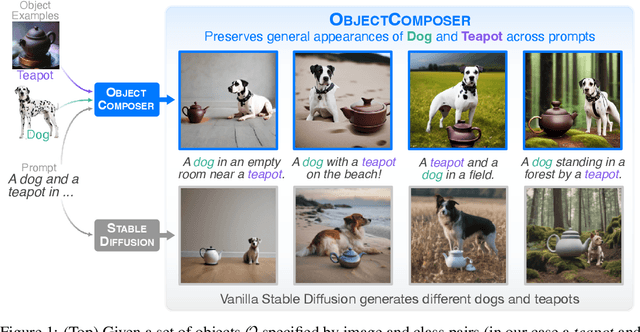
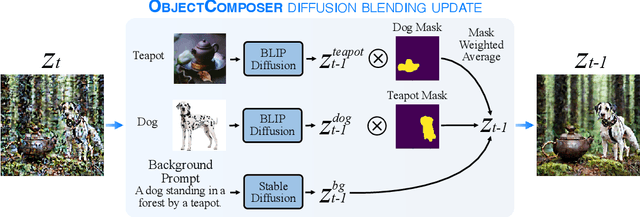

Recent text-to-image generative models can generate high-fidelity images from text prompts. However, these models struggle to consistently generate the same objects in different contexts with the same appearance. Consistent object generation is important to many downstream tasks like generating comic book illustrations with consistent characters and setting. Numerous approaches attempt to solve this problem by extending the vocabulary of diffusion models through fine-tuning. However, even lightweight fine-tuning approaches can be prohibitively expensive to run at scale and in real-time. We introduce a method called ObjectComposer for generating compositions of multiple objects that resemble user-specified images. Our approach is training-free, leveraging the abilities of preexisting models. We build upon the recent BLIP-Diffusion model, which can generate images of single objects specified by reference images. ObjectComposer enables the consistent generation of compositions containing multiple specific objects simultaneously, all without modifying the weights of the underlying models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge