"Image": models, code, and papers
Advanced Underwater Image Restoration in Complex Illumination Conditions
Sep 05, 2023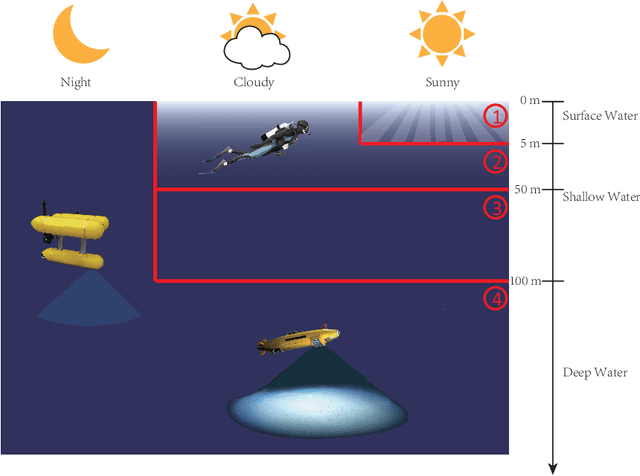
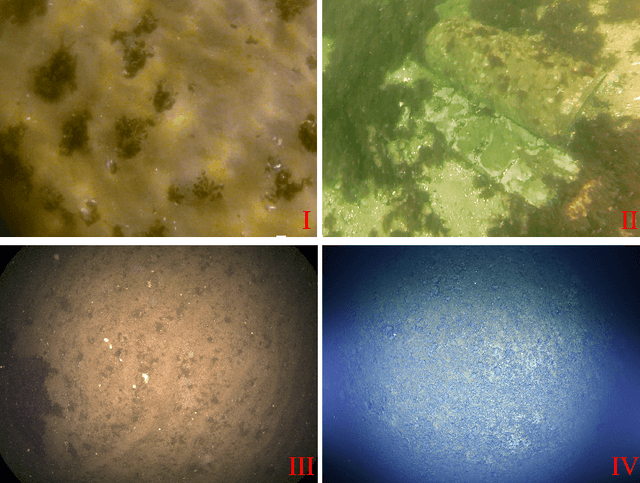

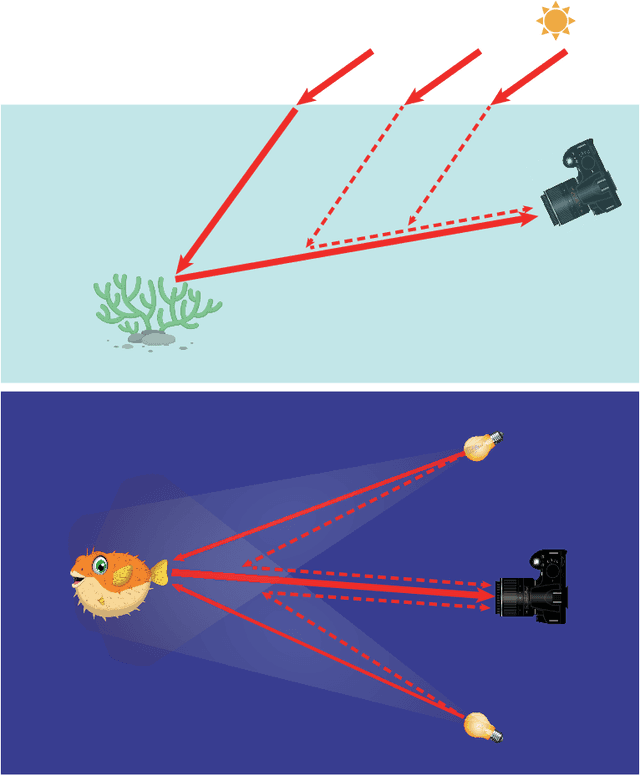
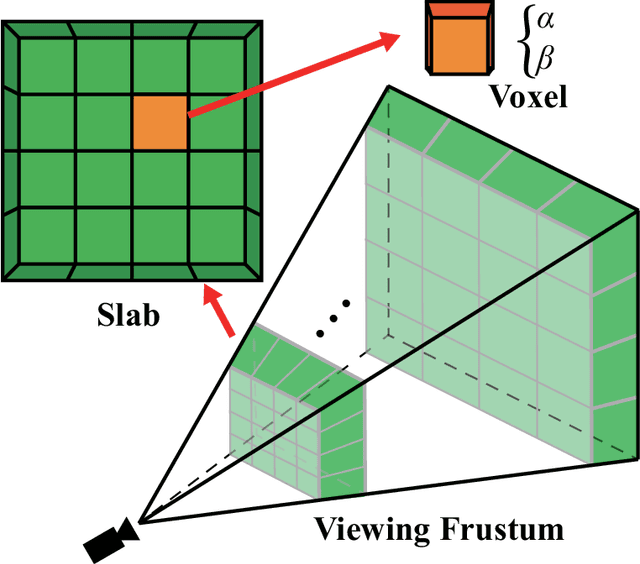
Underwater image restoration has been a challenging problem for decades since the advent of underwater photography. Most solutions focus on shallow water scenarios, where the scene is uniformly illuminated by the sunlight. However, the vast majority of uncharted underwater terrain is located beyond 200 meters depth where natural light is scarce and artificial illumination is needed. In such cases, light sources co-moving with the camera, dynamically change the scene appearance, which make shallow water restoration methods inadequate. In particular for multi-light source systems (composed of dozens of LEDs nowadays), calibrating each light is time-consuming, error-prone and tedious, and we observe that only the integrated illumination within the viewing volume of the camera is critical, rather than the individual light sources. The key idea of this paper is therefore to exploit the appearance changes of objects or the seafloor, when traversing the viewing frustum of the camera. Through new constraints assuming Lambertian surfaces, corresponding image pixels constrain the light field in front of the camera, and for each voxel a signal factor and a backscatter value are stored in a volumetric grid that can be used for very efficient image restoration of camera-light platforms, which facilitates consistently texturing large 3D models and maps that would otherwise be dominated by lighting and medium artifacts. To validate the effectiveness of our approach, we conducted extensive experiments on simulated and real-world datasets. The results of these experiments demonstrate the robustness of our approach in restoring the true albedo of objects, while mitigating the influence of lighting and medium effects. Furthermore, we demonstrate our approach can be readily extended to other scenarios, including in-air imaging with artificial illumination or other similar cases.
A Generic Fundus Image Enhancement Network Boosted by Frequency Self-supervised Representation Learning
Sep 02, 2023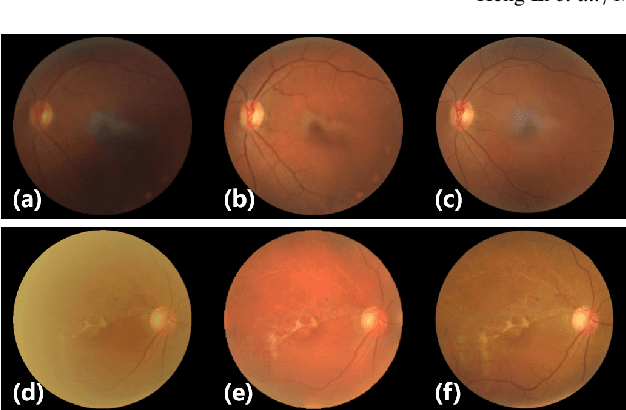
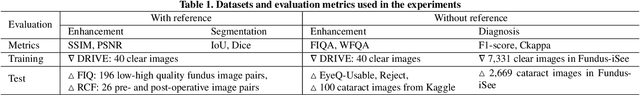


Fundus photography is prone to suffer from image quality degradation that impacts clinical examination performed by ophthalmologists or intelligent systems. Though enhancement algorithms have been developed to promote fundus observation on degraded images, high data demands and limited applicability hinder their clinical deployment. To circumvent this bottleneck, a generic fundus image enhancement network (GFE-Net) is developed in this study to robustly correct unknown fundus images without supervised or extra data. Levering image frequency information, self-supervised representation learning is conducted to learn robust structure-aware representations from degraded images. Then with a seamless architecture that couples representation learning and image enhancement, GFE-Net can accurately correct fundus images and meanwhile preserve retinal structures. Comprehensive experiments are implemented to demonstrate the effectiveness and advantages of GFE-Net. Compared with state-of-the-art algorithms, GFE-Net achieves superior performance in data dependency, enhancement performance, deployment efficiency, and scale generalizability. Follow-up fundus image analysis is also facilitated by GFE-Net, whose modules are respectively verified to be effective for image enhancement.
Bellybutton: Accessible and Customizable Deep-Learning Image Segmentation
Aug 31, 2023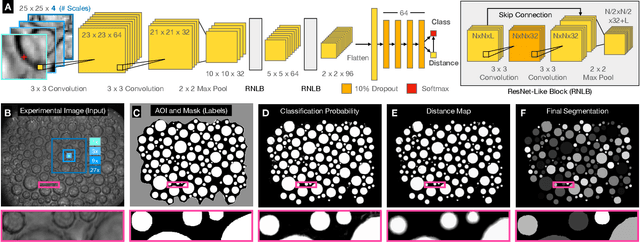
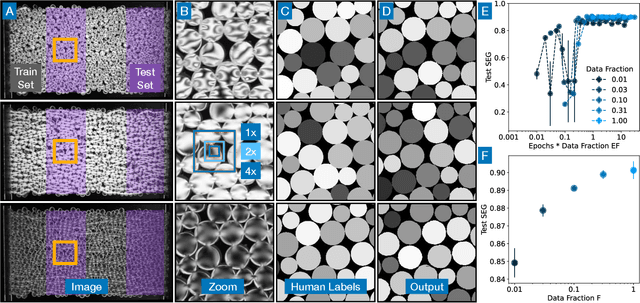
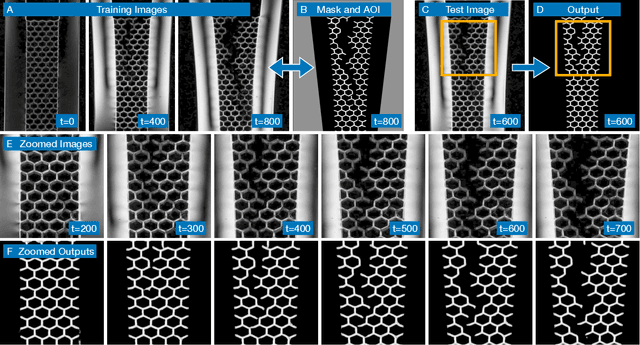


The conversion of raw images into quantifiable data can be a major hurdle in experimental research, and typically involves identifying region(s) of interest, a process known as segmentation. Machine learning tools for image segmentation are often specific to a set of tasks, such as tracking cells, or require substantial compute or coding knowledge to train and use. Here we introduce an easy-to-use (no coding required), image segmentation method, using a 15-layer convolutional neural network that can be trained on a laptop: Bellybutton. The algorithm trains on user-provided segmentation of example images, but, as we show, just one or even a portion of one training image can be sufficient in some cases. We detail the machine learning method and give three use cases where Bellybutton correctly segments images despite substantial lighting, shape, size, focus, and/or structure variation across the regions(s) of interest. Instructions for easy download and use, with further details and the datasets used in this paper are available at pypi.org/project/Bellybuttonseg.
Evaluating the Fairness of Discriminative Foundation Models in Computer Vision
Oct 18, 2023We propose a novel taxonomy for bias evaluation of discriminative foundation models, such as Contrastive Language-Pretraining (CLIP), that are used for labeling tasks. We then systematically evaluate existing methods for mitigating bias in these models with respect to our taxonomy. Specifically, we evaluate OpenAI's CLIP and OpenCLIP models for key applications, such as zero-shot classification, image retrieval and image captioning. We categorize desired behaviors based around three axes: (i) if the task concerns humans; (ii) how subjective the task is (i.e., how likely it is that people from a diverse range of backgrounds would agree on a labeling); and (iii) the intended purpose of the task and if fairness is better served by impartiality (i.e., making decisions independent of the protected attributes) or representation (i.e., making decisions to maximize diversity). Finally, we provide quantitative fairness evaluations for both binary-valued and multi-valued protected attributes over ten diverse datasets. We find that fair PCA, a post-processing method for fair representations, works very well for debiasing in most of the aforementioned tasks while incurring only minor loss of performance. However, different debiasing approaches vary in their effectiveness depending on the task. Hence, one should choose the debiasing approach depending on the specific use case.
Cloud-Magnetic Resonance Imaging System: In the Era of 6G and Artificial Intelligence
Oct 18, 2023Magnetic Resonance Imaging (MRI) plays an important role in medical diagnosis, generating petabytes of image data annually in large hospitals. This voluminous data stream requires a significant amount of network bandwidth and extensive storage infrastructure. Additionally, local data processing demands substantial manpower and hardware investments. Data isolation across different healthcare institutions hinders cross-institutional collaboration in clinics and research. In this work, we anticipate an innovative MRI system and its four generations that integrate emerging distributed cloud computing, 6G bandwidth, edge computing, federated learning, and blockchain technology. This system is called Cloud-MRI, aiming at solving the problems of MRI data storage security, transmission speed, AI algorithm maintenance, hardware upgrading, and collaborative work. The workflow commences with the transformation of k-space raw data into the standardized Imaging Society for Magnetic Resonance in Medicine Raw Data (ISMRMRD) format. Then, the data are uploaded to the cloud or edge nodes for fast image reconstruction, neural network training, and automatic analysis. Then, the outcomes are seamlessly transmitted to clinics or research institutes for diagnosis and other services. The Cloud-MRI system will save the raw imaging data, reduce the risk of data loss, facilitate inter-institutional medical collaboration, and finally improve diagnostic accuracy and work efficiency.
Auto-Prompting SAM for Mobile Friendly 3D Medical Image Segmentation
Aug 28, 2023The Segment Anything Model (SAM) has rapidly been adopted for segmenting a wide range of natural images. However, recent studies have indicated that SAM exhibits subpar performance on 3D medical image segmentation tasks. In addition to the domain gaps between natural and medical images, disparities in the spatial arrangement between 2D and 3D images, the substantial computational burden imposed by powerful GPU servers, and the time-consuming manual prompt generation impede the extension of SAM to a broader spectrum of medical image segmentation applications. To address these challenges, in this work, we introduce a novel method, AutoSAM Adapter, designed specifically for 3D multi-organ CT-based segmentation. We employ parameter-efficient adaptation techniques in developing an automatic prompt learning paradigm to facilitate the transformation of the SAM model's capabilities to 3D medical image segmentation, eliminating the need for manually generated prompts. Furthermore, we effectively transfer the acquired knowledge of the AutoSAM Adapter to other lightweight models specifically tailored for 3D medical image analysis, achieving state-of-the-art (SOTA) performance on medical image segmentation tasks. Through extensive experimental evaluation, we demonstrate the AutoSAM Adapter as a critical foundation for effectively leveraging the emerging ability of foundation models in 2D natural image segmentation for 3D medical image segmentation.
PAI-Diffusion: Constructing and Serving a Family of Open Chinese Diffusion Models for Text-to-image Synthesis on the Cloud
Sep 11, 2023
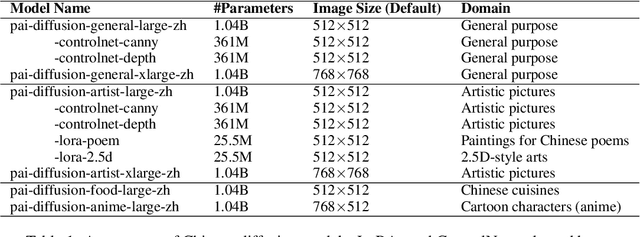
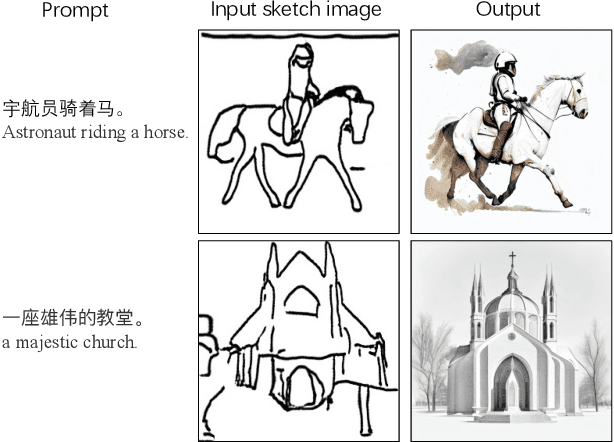
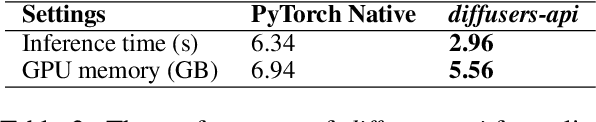
Text-to-image synthesis for the Chinese language poses unique challenges due to its large vocabulary size, and intricate character relationships. While existing diffusion models have shown promise in generating images from textual descriptions, they often neglect domain-specific contexts and lack robustness in handling the Chinese language. This paper introduces PAI-Diffusion, a comprehensive framework that addresses these limitations. PAI-Diffusion incorporates both general and domain-specific Chinese diffusion models, enabling the generation of contextually relevant images. It explores the potential of using LoRA and ControlNet for fine-grained image style transfer and image editing, empowering users with enhanced control over image generation. Moreover, PAI-Diffusion seamlessly integrates with Alibaba Cloud's Machine Learning Platform for AI, providing accessible and scalable solutions. All the Chinese diffusion model checkpoints, LoRAs, and ControlNets, including domain-specific ones, are publicly available. A user-friendly Chinese WebUI and the diffusers-api elastic inference toolkit, also open-sourced, further facilitate the easy deployment of PAI-Diffusion models in various environments, making it a valuable resource for Chinese text-to-image synthesis.
Woodpecker: Hallucination Correction for Multimodal Large Language Models
Oct 24, 2023
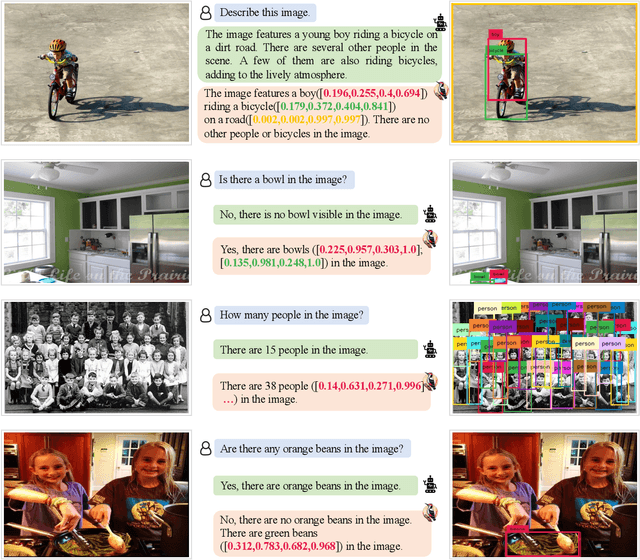
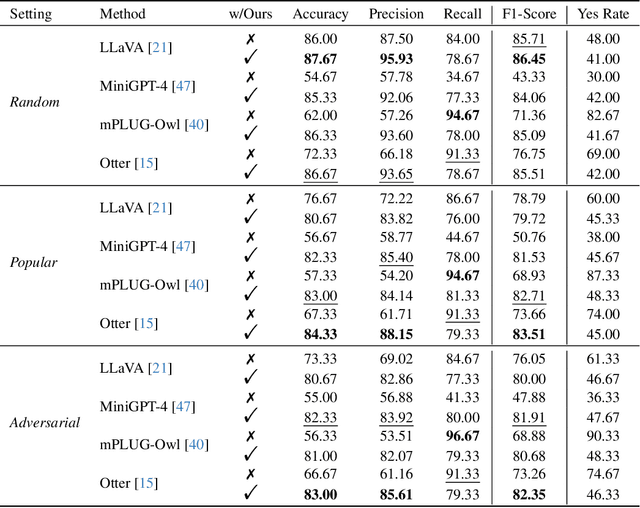

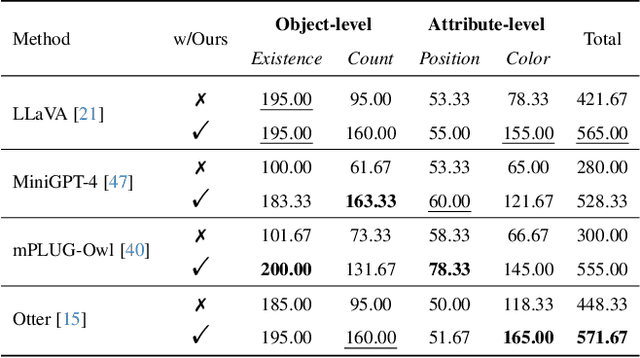
Hallucination is a big shadow hanging over the rapidly evolving Multimodal Large Language Models (MLLMs), referring to the phenomenon that the generated text is inconsistent with the image content. In order to mitigate hallucinations, existing studies mainly resort to an instruction-tuning manner that requires retraining the models with specific data. In this paper, we pave a different way, introducing a training-free method named Woodpecker. Like a woodpecker heals trees, it picks out and corrects hallucinations from the generated text. Concretely, Woodpecker consists of five stages: key concept extraction, question formulation, visual knowledge validation, visual claim generation, and hallucination correction. Implemented in a post-remedy manner, Woodpecker can easily serve different MLLMs, while being interpretable by accessing intermediate outputs of the five stages. We evaluate Woodpecker both quantitatively and qualitatively and show the huge potential of this new paradigm. On the POPE benchmark, our method obtains a 30.66%/24.33% improvement in accuracy over the baseline MiniGPT-4/mPLUG-Owl. The source code is released at https://github.com/BradyFU/Woodpecker.
Multimodal Representations for Teacher-Guided Compositional Visual Reasoning
Oct 24, 2023Neural Module Networks (NMN) are a compelling method for visual question answering, enabling the translation of a question into a program consisting of a series of reasoning sub-tasks that are sequentially executed on the image to produce an answer. NMNs provide enhanced explainability compared to integrated models, allowing for a better understanding of the underlying reasoning process. To improve the effectiveness of NMNs we propose to exploit features obtained by a large-scale cross-modal encoder. Also, the current training approach of NMNs relies on the propagation of module outputs to subsequent modules, leading to the accumulation of prediction errors and the generation of false answers. To mitigate this, we introduce an NMN learning strategy involving scheduled teacher guidance. Initially, the model is fully guided by the ground-truth intermediate outputs, but gradually transitions to an autonomous behavior as training progresses. This reduces error accumulation, thus improving training efficiency and final performance.We demonstrate that by incorporating cross-modal features and employing more effective training techniques for NMN, we achieve a favorable balance between performance and transparency in the reasoning process.
Unpaired MRI Super Resolution with Self-Supervised Contrastive Learning
Oct 24, 2023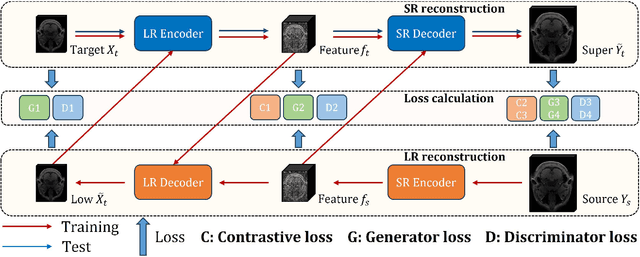
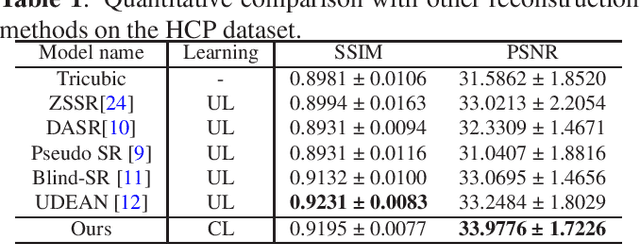
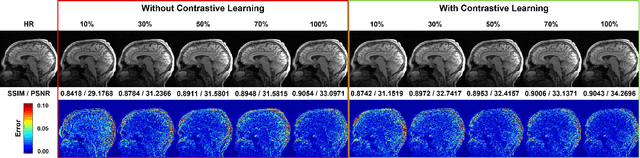
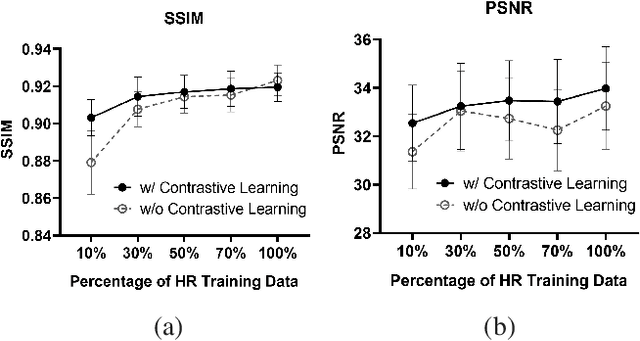
High-resolution (HR) magnetic resonance imaging (MRI) is crucial for enhancing diagnostic accuracy in clinical settings. Nonetheless, the inherent limitation of MRI resolution restricts its widespread applicability. Deep learning-based image super-resolution (SR) methods exhibit promise in improving MRI resolution without additional cost. However, these methods frequently require a substantial number of HR MRI images for training, which can be challenging to acquire. In this paper, we propose an unpaired MRI SR approach that employs self-supervised contrastive learning to enhance SR performance with limited training data. Our approach leverages both authentic HR images and synthetically generated SR images to construct positive and negative sample pairs, thus facilitating the learning of discriminative features. Empirical results presented in this study underscore significant enhancements in the peak signal-to-noise ratio and structural similarity index, even when a paucity of HR images is available. These findings accentuate the potential of our approach in addressing the challenge of limited training data, thereby contributing to the advancement of high-resolution MRI in clinical applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge