"Image": models, code, and papers
Egocentric Whole-Body Motion Capture with FisheyeViT and Diffusion-Based Motion Refinement
Dec 02, 2023In this work, we explore egocentric whole-body motion capture using a single fisheye camera, which simultaneously estimates human body and hand motion. This task presents significant challenges due to three factors: the lack of high-quality datasets, fisheye camera distortion, and human body self-occlusion. To address these challenges, we propose a novel approach that leverages FisheyeViT to extract fisheye image features, which are subsequently converted into pixel-aligned 3D heatmap representations for 3D human body pose prediction. For hand tracking, we incorporate dedicated hand detection and hand pose estimation networks for regressing 3D hand poses. Finally, we develop a diffusion-based whole-body motion prior model to refine the estimated whole-body motion while accounting for joint uncertainties. To train these networks, we collect a large synthetic dataset, EgoWholeBody, comprising 840,000 high-quality egocentric images captured across a diverse range of whole-body motion sequences. Quantitative and qualitative evaluations demonstrate the effectiveness of our method in producing high-quality whole-body motion estimates from a single egocentric camera.
BiomedJourney: Counterfactual Biomedical Image Generation by Instruction-Learning from Multimodal Patient Journeys
Oct 18, 2023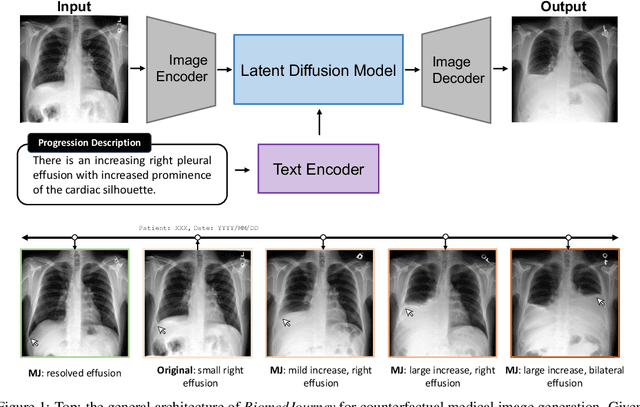
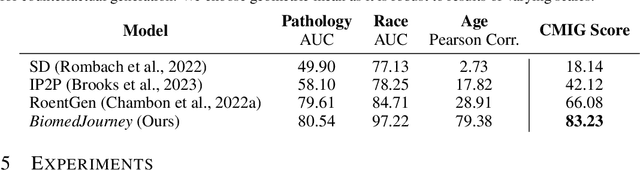
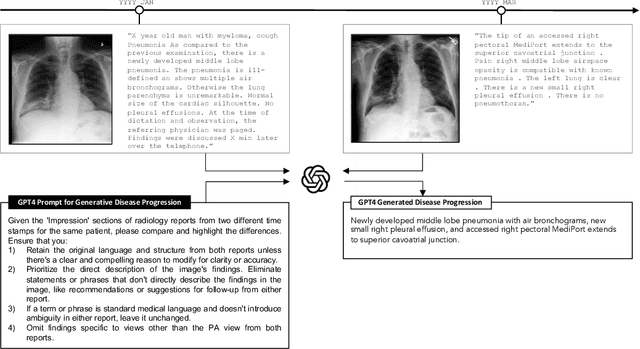
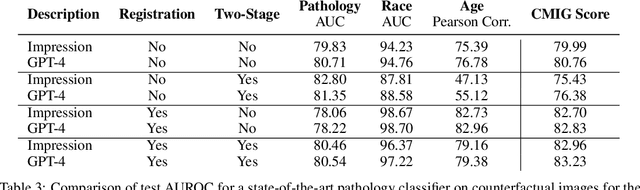
Rapid progress has been made in instruction-learning for image editing with natural-language instruction, as exemplified by InstructPix2Pix. In biomedicine, such methods can be applied to counterfactual image generation, which helps differentiate causal structure from spurious correlation and facilitate robust image interpretation for disease progression modeling. However, generic image-editing models are ill-suited for the biomedical domain, and counterfactual biomedical image generation is largely underexplored. In this paper, we present BiomedJourney, a novel method for counterfactual biomedical image generation by instruction-learning from multimodal patient journeys. Given a patient with two biomedical images taken at different time points, we use GPT-4 to process the corresponding imaging reports and generate a natural language description of disease progression. The resulting triples (prior image, progression description, new image) are then used to train a latent diffusion model for counterfactual biomedical image generation. Given the relative scarcity of image time series data, we introduce a two-stage curriculum that first pretrains the denoising network using the much more abundant single image-report pairs (with dummy prior image), and then continues training using the counterfactual triples. Experiments using the standard MIMIC-CXR dataset demonstrate the promise of our method. In a comprehensive battery of tests on counterfactual medical image generation, BiomedJourney substantially outperforms prior state-of-the-art methods in instruction image editing and medical image generation such as InstructPix2Pix and RoentGen. To facilitate future study in counterfactual medical generation, we plan to release our instruction-learning code and pretrained models.
CellMixer: Annotation-free Semantic Cell Segmentation of Heterogeneous Cell Populations
Dec 01, 2023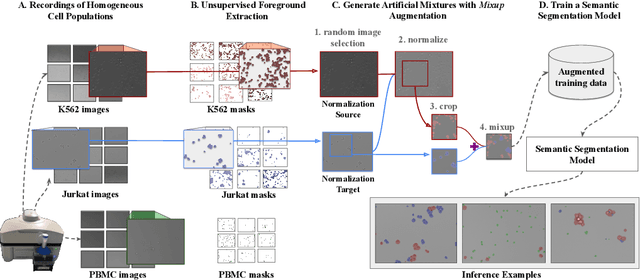
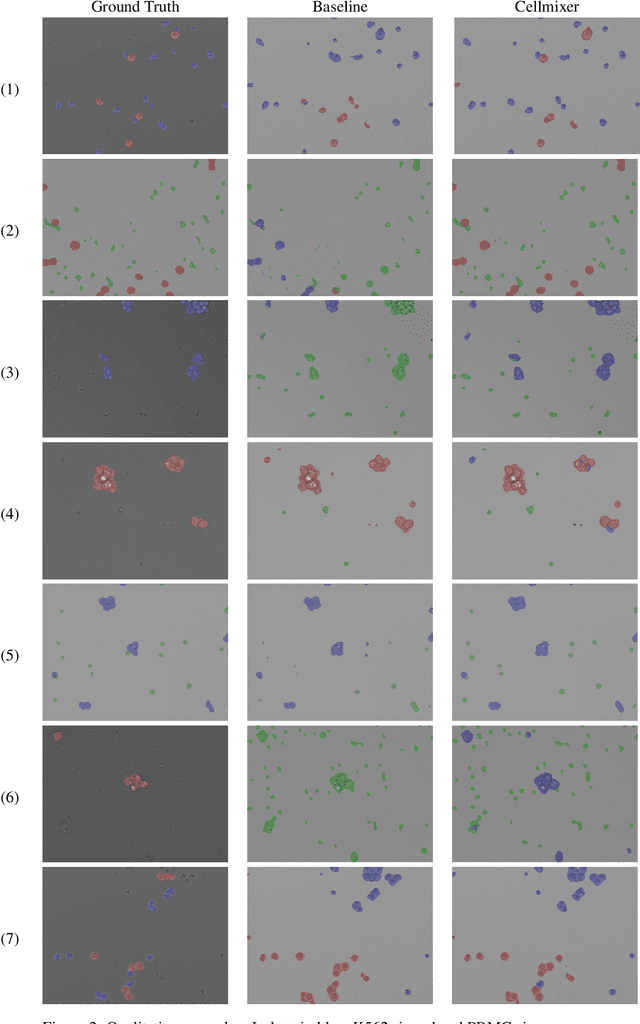
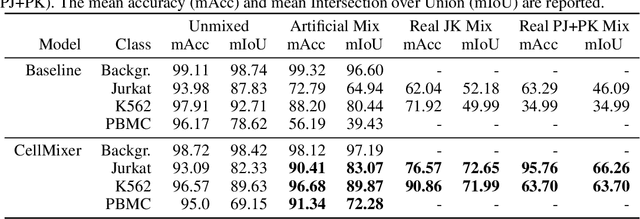

In recent years, several unsupervised cell segmentation methods have been presented, trying to omit the requirement of laborious pixel-level annotations for the training of a cell segmentation model. Most if not all of these methods handle the instance segmentation task by focusing on the detection of different cell instances ignoring their type. While such models prove adequate for certain tasks, like cell counting, other applications require the identification of each cell's type. In this paper, we present CellMixer, an innovative annotation-free approach for the semantic segmentation of heterogeneous cell populations. Our augmentation-based method enables the training of a segmentation model from image-level labels of homogeneous cell populations. Our results show that CellMixer can achieve competitive segmentation performance across multiple cell types and imaging modalities, demonstrating the method's scalability and potential for broader applications in medical imaging, cellular biology, and diagnostics.
DeepDR: Deep Structure-Aware RGB-D Inpainting for Diminished Reality
Dec 01, 2023Diminished reality (DR) refers to the removal of real objects from the environment by virtually replacing them with their background. Modern DR frameworks use inpainting to hallucinate unobserved regions. While recent deep learning-based inpainting is promising, the DR use case is complicated by the need to generate coherent structure and 3D geometry (i.e., depth), in particular for advanced applications, such as 3D scene editing. In this paper, we propose DeepDR, a first RGB-D inpainting framework fulfilling all requirements of DR: Plausible image and geometry inpainting with coherent structure, running at real-time frame rates, with minimal temporal artifacts. Our structure-aware generative network allows us to explicitly condition color and depth outputs on the scene semantics, overcoming the difficulty of reconstructing sharp and consistent boundaries in regions with complex backgrounds. Experimental results show that the proposed framework can outperform related work qualitatively and quantitatively.
REDUCR: Robust Data Downsampling Using Class Priority Reweighting
Dec 01, 2023Modern machine learning models are becoming increasingly expensive to train for real-world image and text classification tasks, where massive web-scale data is collected in a streaming fashion. To reduce the training cost, online batch selection techniques have been developed to choose the most informative datapoints. However, these techniques can suffer from poor worst-class generalization performance due to class imbalance and distributional shifts. This work introduces REDUCR, a robust and efficient data downsampling method that uses class priority reweighting. REDUCR reduces the training data while preserving worst-class generalization performance. REDUCR assigns priority weights to datapoints in a class-aware manner using an online learning algorithm. We demonstrate the data efficiency and robust performance of REDUCR on vision and text classification tasks. On web-scraped datasets with imbalanced class distributions, REDUCR significantly improves worst-class test accuracy (and average accuracy), surpassing state-of-the-art methods by around 15%.
Impact of Data Augmentation on QCNNs
Dec 01, 2023In recent years, Classical Convolutional Neural Networks (CNNs) have been applied for image recognition successfully. Quantum Convolutional Neural Networks (QCNNs) are proposed as a novel generalization to CNNs by using quantum mechanisms. The quantum mechanisms lead to an efficient training process in QCNNs by reducing the size of input from $N$ to $log_2N$. This paper implements and compares both CNNs and QCNNs by testing losses and prediction accuracy on three commonly used datasets. The datasets include the MNIST hand-written digits, Fashion MNIST and cat/dog face images. Additionally, data augmentation (DA), a technique commonly used in CNNs to improve the performance of classification by generating similar images based on original inputs, is also implemented in QCNNs. Surprisingly, the results showed that data augmentation didn't improve QCNNs performance. The reasons and logic behind this result are discussed, hoping to expand our understanding of Quantum machine learning theory.
Localizing and Editing Knowledge in Text-to-Image Generative Models
Oct 20, 2023Text-to-Image Diffusion Models such as Stable-Diffusion and Imagen have achieved unprecedented quality of photorealism with state-of-the-art FID scores on MS-COCO and other generation benchmarks. Given a caption, image generation requires fine-grained knowledge about attributes such as object structure, style, and viewpoint amongst others. Where does this information reside in text-to-image generative models? In our paper, we tackle this question and understand how knowledge corresponding to distinct visual attributes is stored in large-scale text-to-image diffusion models. We adapt Causal Mediation Analysis for text-to-image models and trace knowledge about distinct visual attributes to various (causal) components in the (i) UNet and (ii) text-encoder of the diffusion model. In particular, we show that unlike generative large-language models, knowledge about different attributes is not localized in isolated components, but is instead distributed amongst a set of components in the conditional UNet. These sets of components are often distinct for different visual attributes. Remarkably, we find that the CLIP text-encoder in public text-to-image models such as Stable-Diffusion contains only one causal state across different visual attributes, and this is the first self-attention layer corresponding to the last subject token of the attribute in the caption. This is in stark contrast to the causal states in other language models which are often the mid-MLP layers. Based on this observation of only one causal state in the text-encoder, we introduce a fast, data-free model editing method Diff-QuickFix which can effectively edit concepts in text-to-image models. DiffQuickFix can edit (ablate) concepts in under a second with a closed-form update, providing a significant 1000x speedup and comparable editing performance to existing fine-tuning based editing methods.
Contrastive Learning-Based Spectral Knowledge Distillation for Multi-Modality and Missing Modality Scenarios in Semantic Segmentation
Dec 04, 2023Improving the performance of semantic segmentation models using multispectral information is crucial, especially for environments with low-light and adverse conditions. Multi-modal fusion techniques pursue either the learning of cross-modality features to generate a fused image or engage in knowledge distillation but address multimodal and missing modality scenarios as distinct issues, which is not an optimal approach for multi-sensor models. To address this, a novel multi-modal fusion approach called CSK-Net is proposed, which uses a contrastive learning-based spectral knowledge distillation technique along with an automatic mixed feature exchange mechanism for semantic segmentation in optical (EO) and infrared (IR) images. The distillation scheme extracts detailed textures from the optical images and distills them into the optical branch of CSK-Net. The model encoder consists of shared convolution weights with separate batch norm (BN) layers for both modalities, to capture the multi-spectral information from different modalities of the same objects. A Novel Gated Spectral Unit (GSU) and mixed feature exchange strategy are proposed to increase the correlation of modality-shared information and decrease the modality-specific information during the distillation process. Comprehensive experiments show that CSK-Net surpasses state-of-the-art models in multi-modal tasks and for missing modalities when exclusively utilizing IR data for inference across three public benchmarking datasets. For missing modality scenarios, the performance increase is achieved without additional computational costs compared to the baseline segmentation models.
High-resolution Image-based Malware Classification using Multiple Instance Learning
Nov 21, 2023This paper proposes a novel method of classifying malware into families using high-resolution greyscale images and multiple instance learning to overcome adversarial binary enlargement. Current methods of visualisation-based malware classification largely rely on lossy transformations of inputs such as resizing to handle the large, variable-sized images. Through empirical analysis and experimentation, it is shown that these approaches cause crucial information loss that can be exploited. The proposed solution divides the images into patches and uses embedding-based multiple instance learning with a convolutional neural network and an attention aggregation function for classification. The implementation is evaluated on the Microsoft Malware Classification dataset and achieves accuracies of up to $96.6\%$ on adversarially enlarged samples compared to the baseline of $22.8\%$. The Python code is available online at https://github.com/timppeters/MIL-Malware-Images .
3DGAUnet: 3D generative adversarial networks with a 3D U-Net based generator to achieve the accurate and effective synthesis of clinical tumor image data for pancreatic cancer
Nov 09, 2023Pancreatic ductal adenocarcinoma (PDAC) presents a critical global health challenge, and early detection is crucial for improving the 5-year survival rate. Recent medical imaging and computational algorithm advances offer potential solutions for early diagnosis. Deep learning, particularly in the form of convolutional neural networks (CNNs), has demonstrated success in medical image analysis tasks, including classification and segmentation. However, the limited availability of clinical data for training purposes continues to provide a significant obstacle. Data augmentation, generative adversarial networks (GANs), and cross-validation are potential techniques to address this limitation and improve model performance, but effective solutions are still rare for 3D PDAC, where contrast is especially poor owing to the high heterogeneity in both tumor and background tissues. In this study, we developed a new GAN-based model, named 3DGAUnet, for generating realistic 3D CT images of PDAC tumors and pancreatic tissue, which can generate the interslice connection data that the existing 2D CT image synthesis models lack. Our innovation is to develop a 3D U-Net architecture for the generator to improve shape and texture learning for PDAC tumors and pancreatic tissue. Our approach offers a promising path to tackle the urgent requirement for creative and synergistic methods to combat PDAC. The development of this GAN-based model has the potential to alleviate data scarcity issues, elevate the quality of synthesized data, and thereby facilitate the progression of deep learning models to enhance the accuracy and early detection of PDAC tumors, which could profoundly impact patient outcomes. Furthermore, this model has the potential to be adapted to other types of solid tumors, hence making significant contributions to the field of medical imaging in terms of image processing models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge