"Image": models, code, and papers
FoodTracker: A Real-time Food Detection Mobile Application by Deep Convolutional Neural Networks
Sep 16, 2019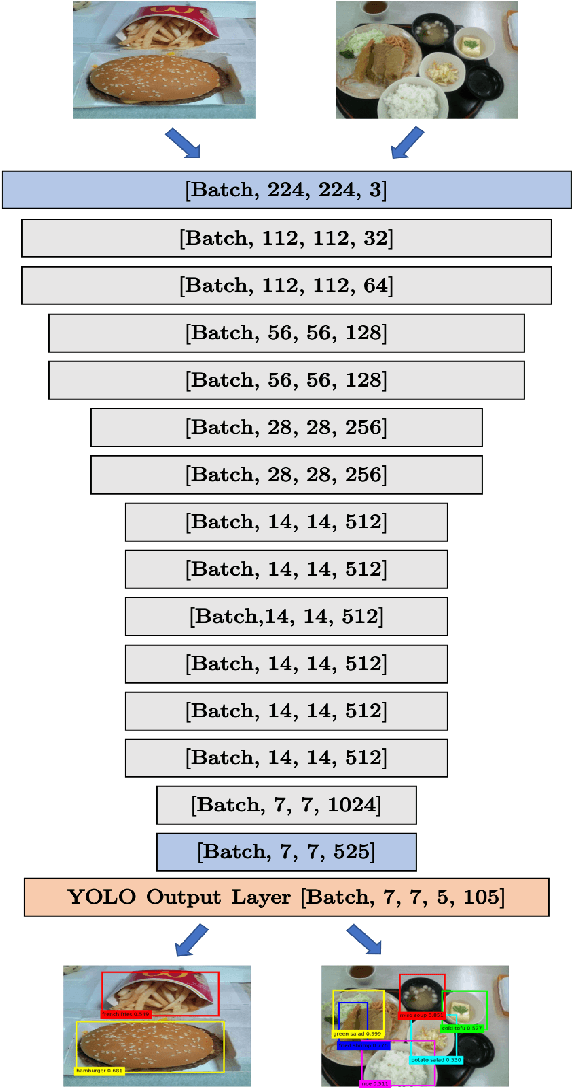
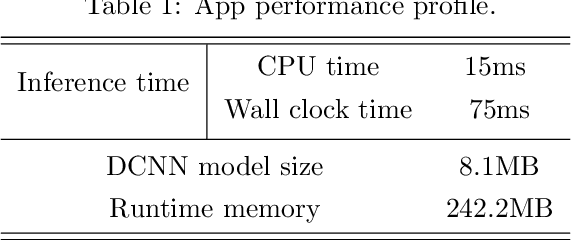
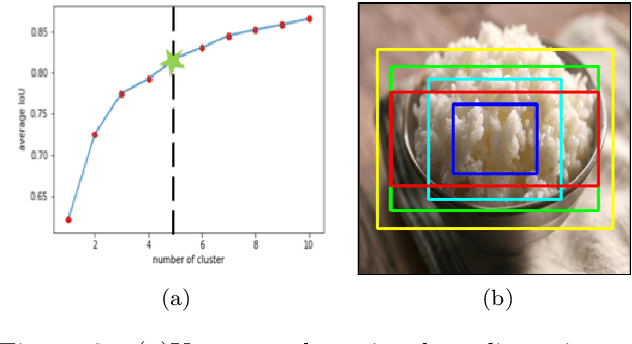
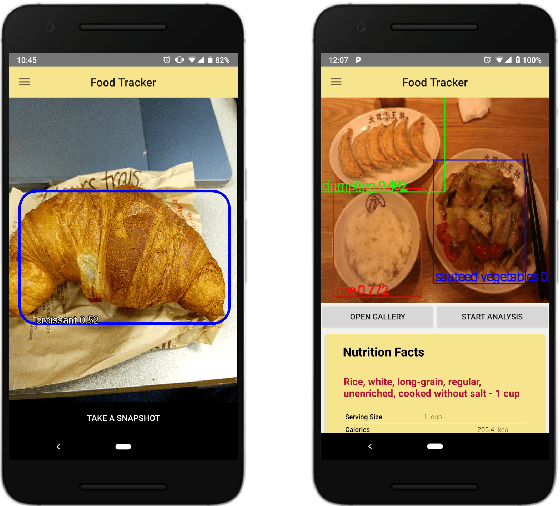
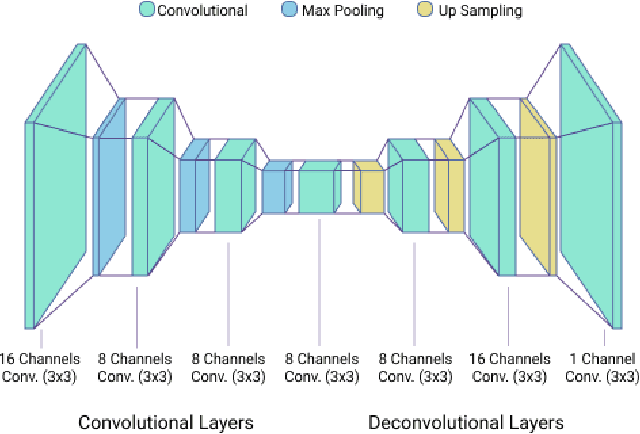
We present a mobile application made to recognize food items of multi-object meal from a single image in real-time, and then return the nutrition facts with components and approximate amounts. Our work is organized in two parts. First, we build a deep convolutional neural network merging with YOLO, a state-of-the-art detection strategy, to achieve simultaneous multi-object recognition and localization with nearly 80% mean average precision. Second, we adapt our model into a mobile application with extending function for nutrition analysis. After inferring and decoding the model output in the app side, we present detection results that include bounding box position and class label in either real-time or local mode. Our model is well-suited for mobile devices with negligible inference time and small memory requirements with a deep learning algorithm.
Identifying Candidate Spaces for Advert Implantation
Oct 08, 2019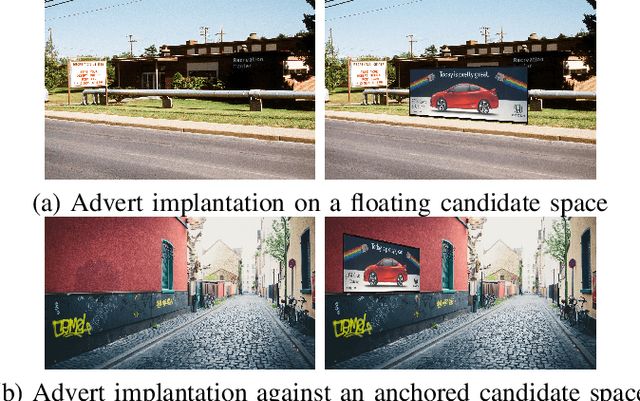

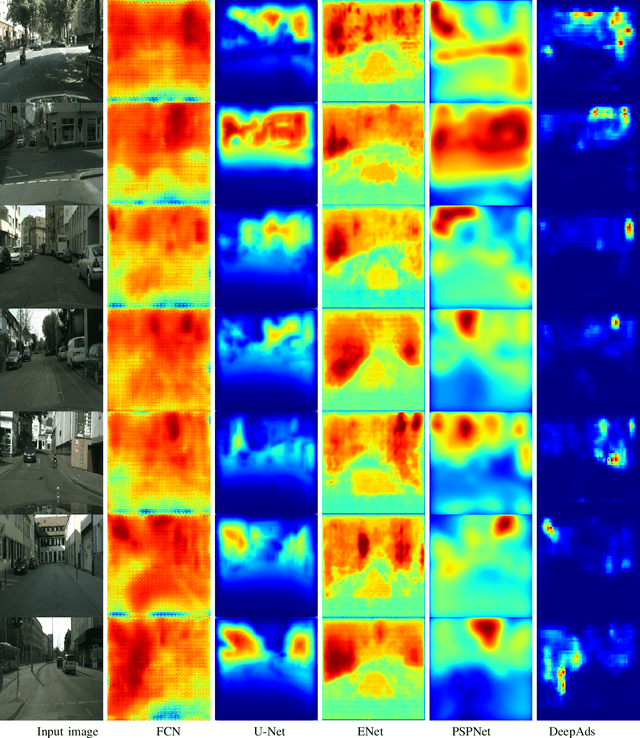
Virtual advertising is an important and promising feature in the area of online advertising. It involves integrating adverts onto live or recorded videos for product placements and targeted advertisements. Such integration of adverts is primarily done by video editors in the post-production stage, which is cumbersome and time-consuming. Therefore, it is important to automatically identify candidate spaces in a video frame, wherein new adverts can be implanted. The candidate space should match the scene perspective, and also have a high quality of experience according to human subjective judgment. In this paper, we propose the use of a bespoke neural net that can assist the video editors in identifying candidate spaces. We benchmark our approach against several deep-learning architectures on a large-scale image dataset of candidate spaces of outdoor scenes. Our work is the first of its kind in this area of multimedia and augmented reality applications, and achieves the best results.
Non-Causal Tracking by Deblatting
Sep 15, 2019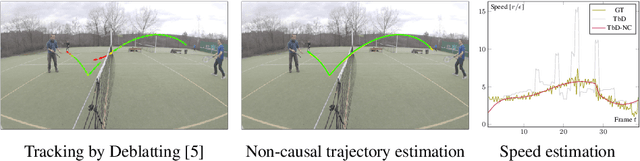
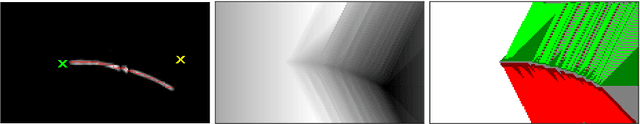
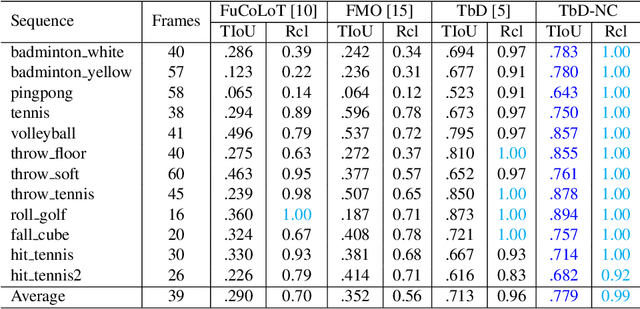


Tracking by Deblatting stands for solving an inverse problem of deblurring and image matting for tracking motion-blurred objects. We propose non-causal Tracking by Deblatting which estimates continuous, complete and accurate object trajectories. Energy minimization by dynamic programming is used to detect abrupt changes of motion, called bounces. High-order polynomials are fitted to segments, which are parts of the trajectory separated by bounces. The output is a continuous trajectory function which assigns location for every real-valued time stamp from zero to the number of frames. Additionally, we show that from the trajectory function precise physical calculations are possible, such as radius, gravity or sub-frame object velocity. Velocity estimation is compared to the high-speed camera measurements and radars. Results show high performance of the proposed method in terms of Trajectory-IoU, recall and velocity estimation.
Interactive Classification for Deep Learning Interpretation
Jun 14, 2018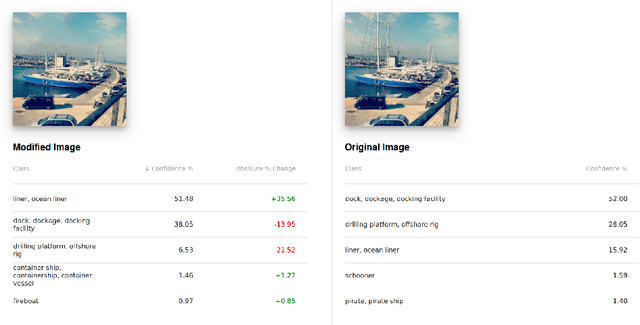
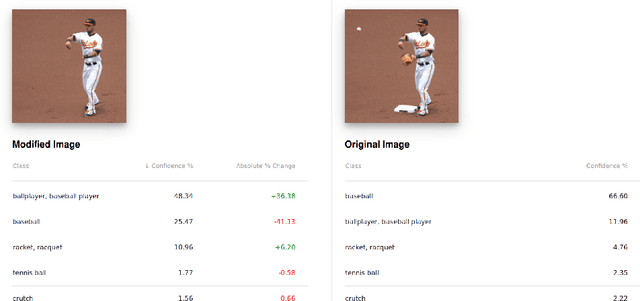
We present an interactive system enabling users to manipulate images to explore the robustness and sensitivity of deep learning image classifiers. Using modern web technologies to run in-browser inference, users can remove image features using inpainting algorithms and obtain new classifications in real time, which allows them to ask a variety of "what if" questions by experimentally modifying images and seeing how the model reacts. Our system allows users to compare and contrast what image regions humans and machine learning models use for classification, revealing a wide range of surprising results ranging from spectacular failures (e.g., a "water bottle" image becomes a "concert" when removing a person) to impressive resilience (e.g., a "baseball player" image remains correctly classified even without a glove or base). We demonstrate our system at The 2018 Conference on Computer Vision and Pattern Recognition (CVPR) for the audience to try it live. Our system is open-sourced at https://github.com/poloclub/interactive-classification. A video demo is available at https://youtu.be/llub5GcOF6w.
Brain-wise Tumor Segmentation and Patient Overall Survival Prediction
Sep 15, 2019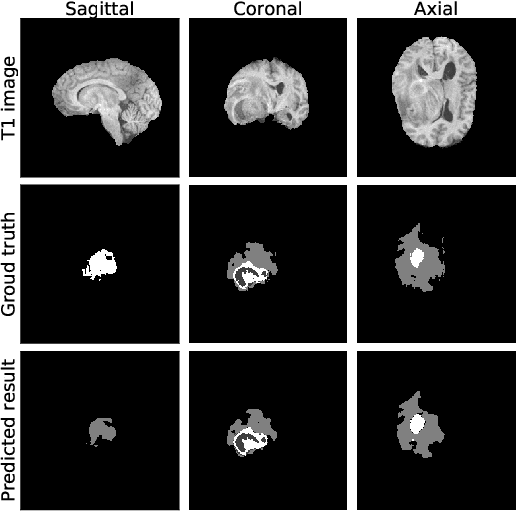
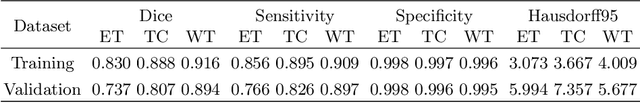

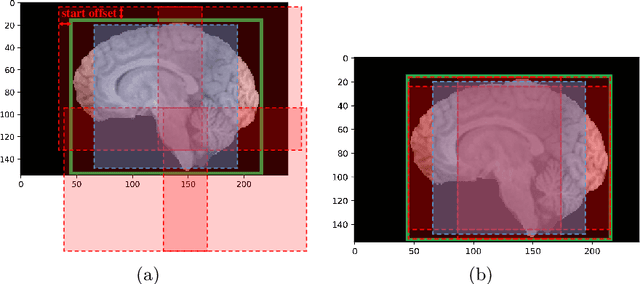
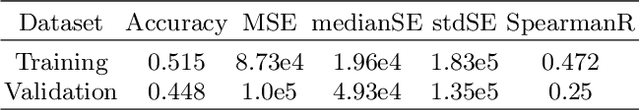
Past few years have witnessed the prevalence of deep learning in many application scenarios, among which is medical image processing. Diagnosis and treatment of brain tumors require a delicate segmentation of brain tumors as a prerequisite. However, such kind of work conventionally costs cerebral surgeons a lot of precious time. Computer vision techniques could provide surgeons a relief from the tedious marking procedure. In this paper, a 3D U-net based deep learning model has been trained with the help of brain-wise normalization and patching strategies for the brain tumor segmentation task in BraTS 2019 competition. Dice coefficients for enhancing tumor, tumor core, and the whole tumor are 0.737, 0.807 and 0.894 respectively on validation dataset. Furthermore, numerical features extracted from predicted tumor labels have been used for the overall survival days prediction task. The prediction accuracy on validation dataset is 0.448.
Likelihood Assignment for Out-of-Distribution Inputs in Deep Generative Models is Sensitive to Prior Distribution Choice
Nov 15, 2019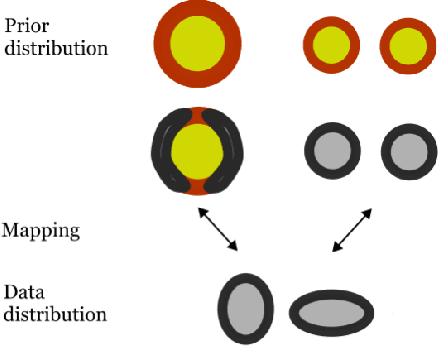
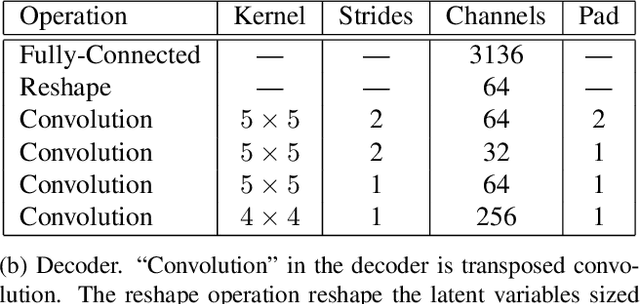
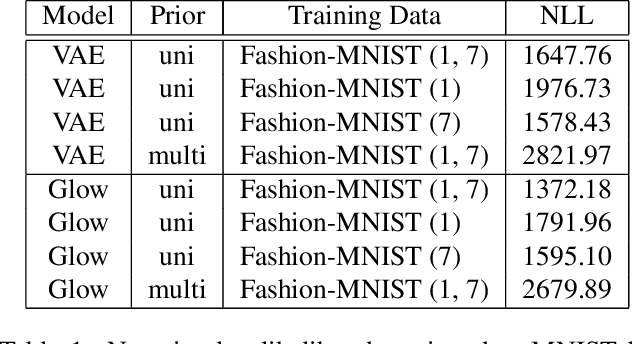
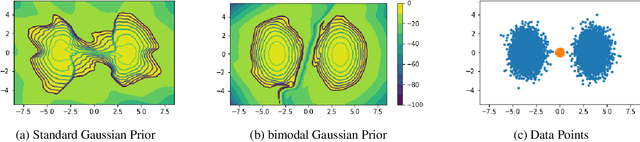

Recent work has shown that deep generative models assign higher likelihood to out-of-distribution inputs than to training data. We show that a factor underlying this phenomenon is a mismatch between the nature of the prior distribution and that of the data distribution, a problem found in widely used deep generative models such as VAEs and Glow. While a typical choice for a prior distribution is a standard Gaussian distribution, properties of distributions of real data sets may not be consistent with a unimodal prior distribution. This paper focuses on the relationship between the choice of a prior distribution and the likelihoods assigned to out-of-distribution inputs. We propose the use of a mixture distribution as a prior to make likelihoods assigned by deep generative models sensitive to out-of-distribution inputs. Furthermore, we explain the theoretical advantages of adopting a mixture distribution as the prior, and we present experimental results to support our claims. Finally, we demonstrate that a mixture prior lowers the out-of-distribution likelihood with respect to two pairs of real image data sets: Fashion-MNIST vs. MNIST and CIFAR10 vs. SVHN.
QC-Automator: Deep Learning-based Automated Quality Control for Diffusion MR Images
Nov 15, 2019
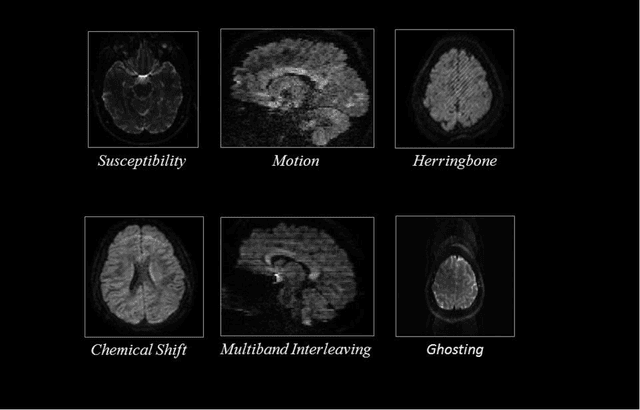

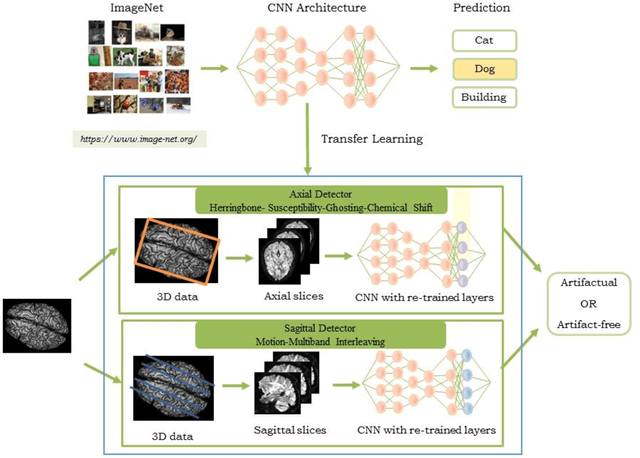
Quality assessment of diffusion MRI (dMRI) data is essential prior to any analysis, so that appropriate pre-processing can be used to improve data quality and ensure that the presence of MRI artifacts do not affect the results of subsequent image analysis. Manual quality assessment of the data is subjective, possibly error-prone, and infeasible, especially considering the growing number of consortium-like studies, underlining the need for automation of the process. In this paper, we have developed a deep-learning-based automated quality control (QC) tool, QC-Automator, for dMRI data, that can handle a variety of artifacts such as motion, multiband interleaving, ghosting, susceptibility, herringbone and chemical shifts. QC-Automator uses convolutional neural networks along with transfer learning to train the automated artifact detection on a labeled dataset of ~332000 slices of dMRI data, from 155 unique subjects and 5 scanners with different dMRI acquisitions, achieving a 98% accuracy in detecting artifacts. The method is fast and paves the way for efficient and effective artifact detection in large datasets. It is also demonstrated to be replicable on other datasets with different acquisition parameters.
Merging and Shifting of Images with Prominence Coefficient for Predictive Analysis using Combined Image
Jul 30, 2014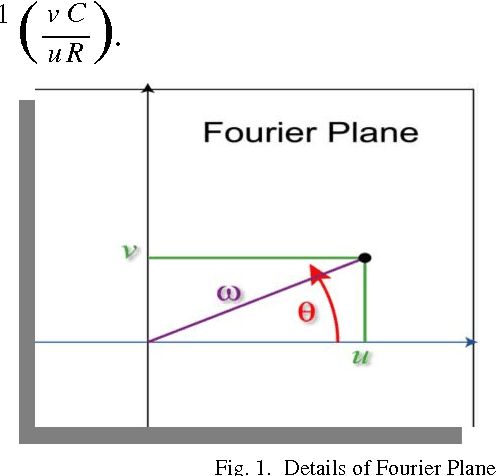
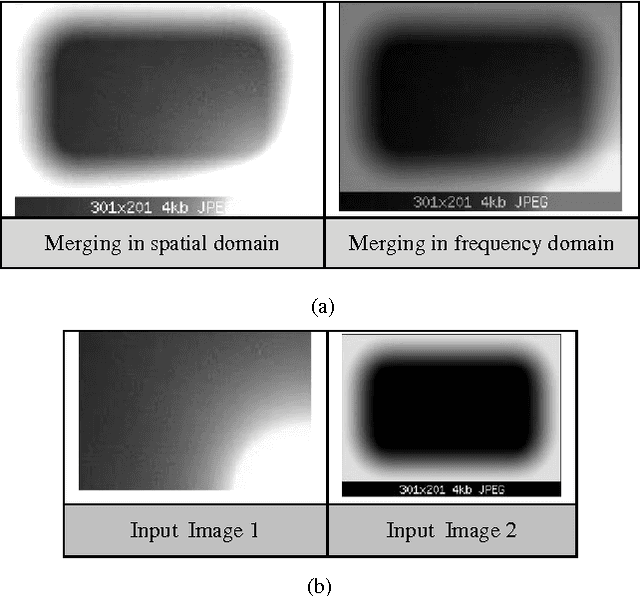
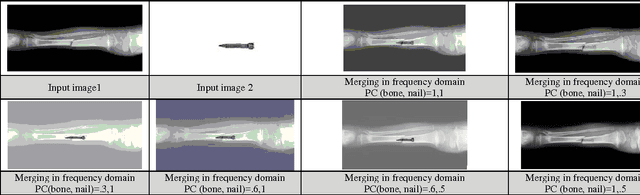
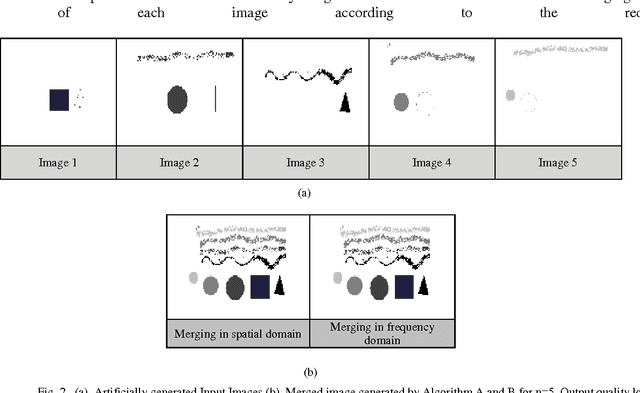


Shifting of objects in an image and merging many images after appropriate shifting is being used in several engineering and scientific applications which require complex perception development. A method has been presented here which could be used in precision engineering and biological applications where more precise prediction is required of a combined phenomenon with varying prominence of each phenomenon. Accurate merging of intended pixels can be achieved in high quality using frequency domain techniques even though initial properties of the original pixels are lost in this process. This paper introduces a technique to shift and merge various images with varying prominence of each image. A coefficient named prominence coefficient has been introduced which is capable of making some of the images transparent and highlighting the rest as per requirement of merging process which can be used as a simple but effective technique for overlapped view of a set of images.
Computed Tomography Image Enhancement using 3D Convolutional Neural Network
Jul 18, 2018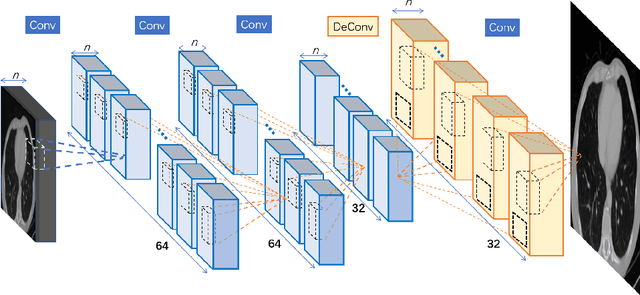
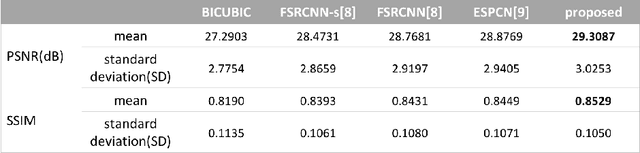
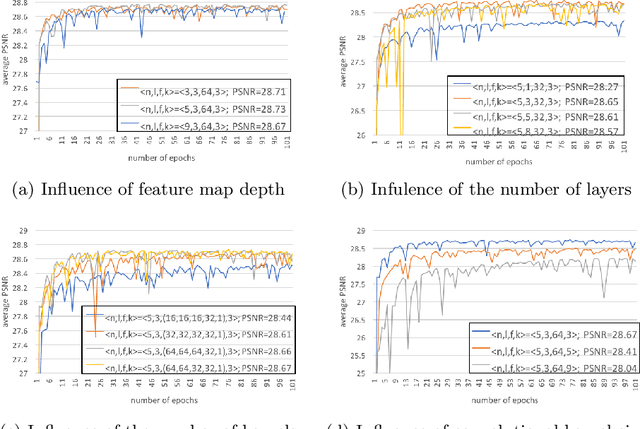
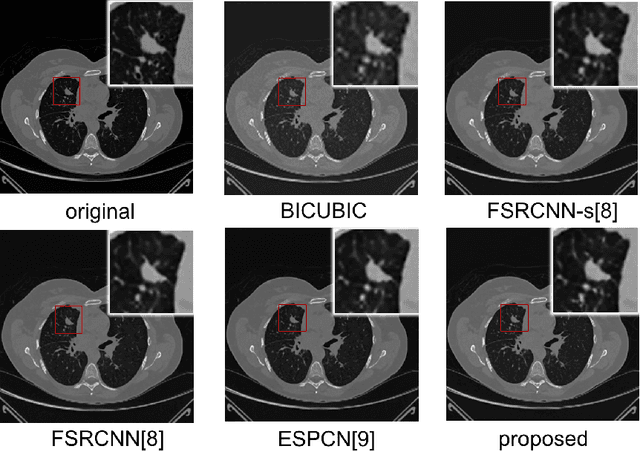
Computed tomography (CT) is increasingly being used for cancer screening, such as early detection of lung cancer. However, CT studies have varying pixel spacing due to differences in acquisition parameters. Thick slice CTs have lower resolution, hindering tasks such as nodule characterization during computer-aided detection due to partial volume effect. In this study, we propose a novel 3D enhancement convolutional neural network (3DECNN) to improve the spatial resolution of CT studies that were acquired using lower resolution/slice thicknesses to higher resolutions. Using a subset of the LIDC dataset consisting of 20,672 CT slices from 100 scans, we simulated lower resolution/thick section scans then attempted to reconstruct the original images using our 3DECNN network. A significant improvement in PSNR (29.3087dB vs. 28.8769dB, p-value < 2.2e-16) and SSIM (0.8529dB vs. 0.8449dB, p-value < 2.2e-16) compared to other state-of-art deep learning methods is observed.
AttentionBoost: Learning What to Attend by Boosting Fully Convolutional Networks
Aug 06, 2019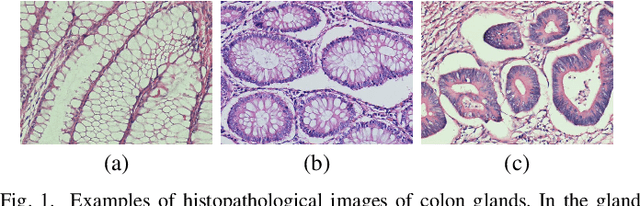
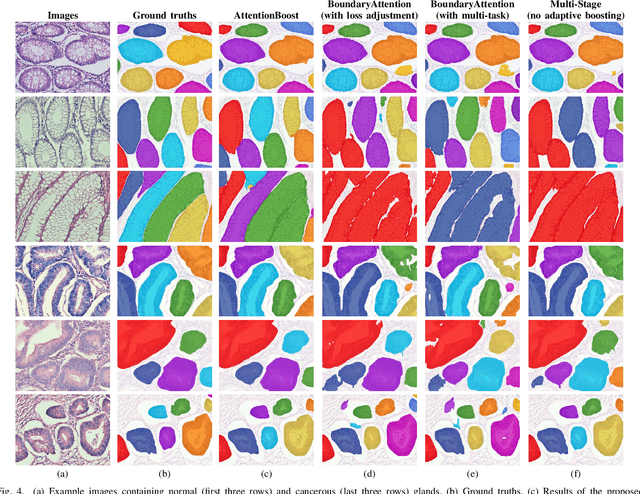
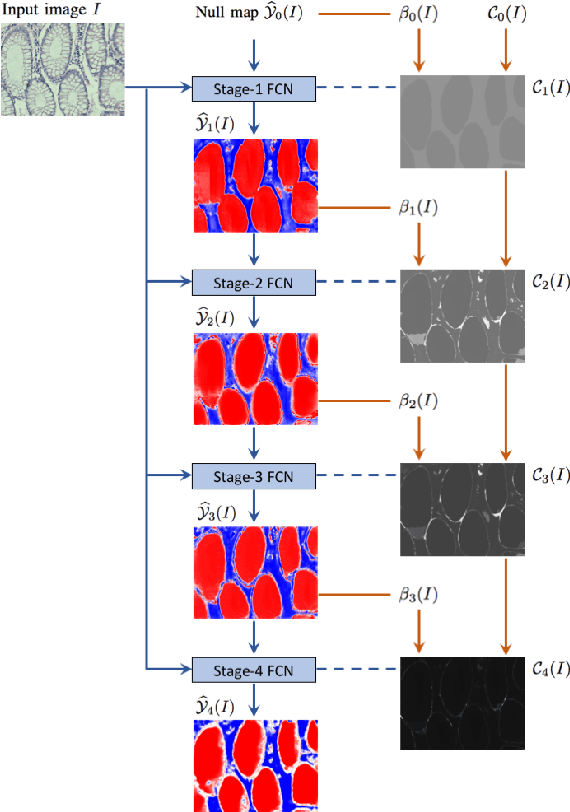
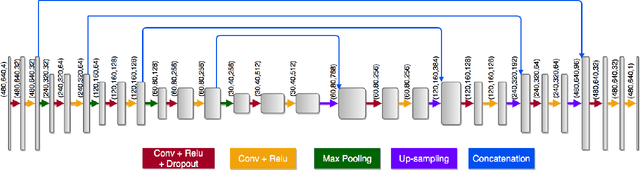

Dense prediction models are widely used for image segmentation. One important challenge is to sufficiently train these models to yield good generalizations for hard-to-learn pixels. A typical group of such hard-to-learn pixels are boundaries between instances. Many studies have proposed to give specific attention to learning the boundary pixels. They include designing multi-task networks with an additional task of boundary prediction and increasing the weights of boundary pixels' predictions in the loss function. Such strategies require defining what to attend beforehand and incorporating this defined attention to the learning model. However, there may exist other groups of hard-to-learn pixels and manually defining and incorporating the appropriate attention for each group may not be feasible. In order to provide a more attainable and scalable solution, this paper proposes AttentionBoost, which is a new multi-attention learning model based on adaptive boosting. AttentionBoost designs a multi-stage network and introduces a new loss adjustment mechanism for a dense prediction model to adaptively learn what to attend at each stage directly on image data without necessitating any prior definition about what to attend. This mechanism modulates the attention of each stage to correct the mistakes of previous stages, by adjusting the loss weight of each pixel prediction separately with respect to how accurate the previous stages are on this pixel. This mechanism enables AttentionBoost to learn different attentions for different pixels at the same stage, according to difficulty of learning these pixels, as well as multiple attentions for the same pixel at different stages, according to confidence of these stages on their predictions for this pixel. Using gland segmentation as a showcase application, our experiments demonstrate that AttentionBoost improves the results of its counterparts.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge