"Image": models, code, and papers
Examining the Capability of GANs to Replace Real Biomedical Images in Classification Models Training
Apr 18, 2019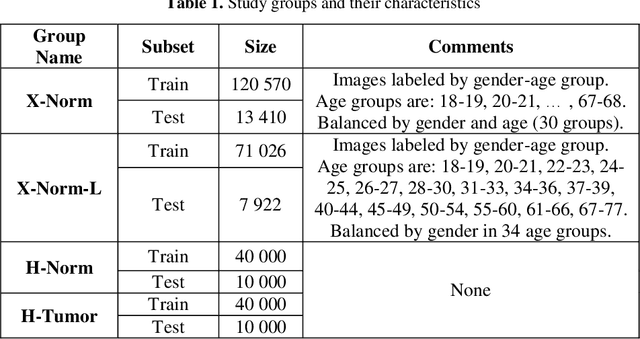
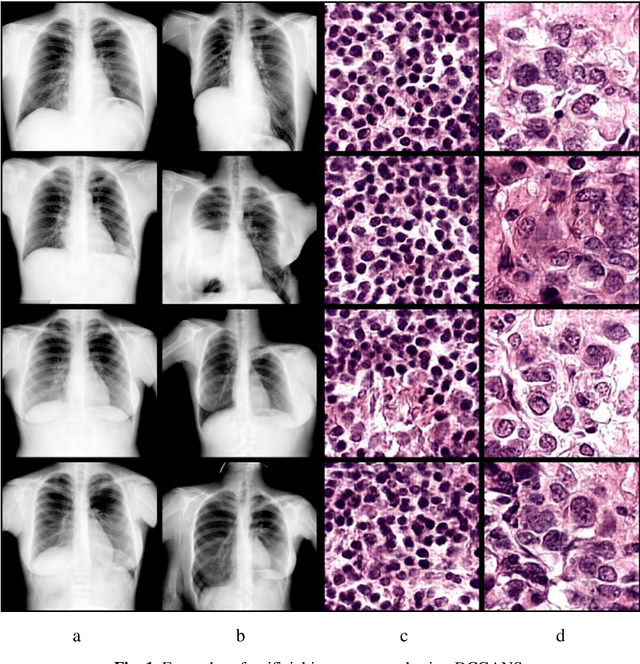
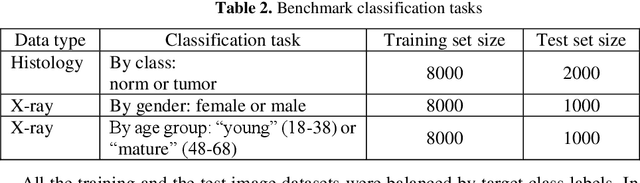
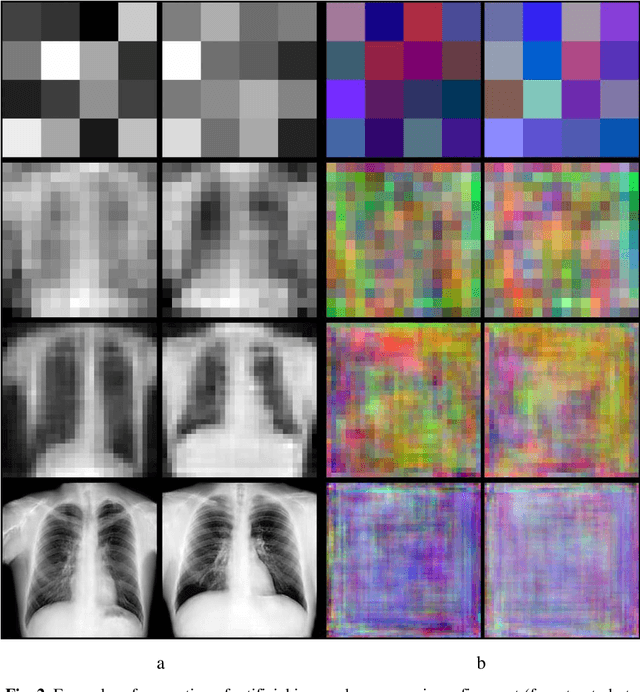
In this paper, we explore the possibility of generating artificial biomedical images that can be used as a substitute for real image datasets in applied machine learning tasks. We are focusing on generation of realistic chest X-ray images as well as on the lymph node histology images using the two recent GAN architectures including DCGAN and PGGAN. The possibility of the use of artificial images instead of real ones for training machine learning models was examined by benchmark classification tasks being solved using conventional and deep learning methods. In particular, a comparison was made by replacing real images with synthetic ones at the model training stage and comparing the prediction results with the ones obtained while training on the real image data. It was found that the drop of classification accuracy caused by such training data substitution ranged between 2.2% and 3.5% for deep learning models and between 5.5% and 13.25% for conventional methods such as LBP + Random Forests.
Dragonfly Algorithm and its Applications in Applied Science -- Survey
Nov 25, 2019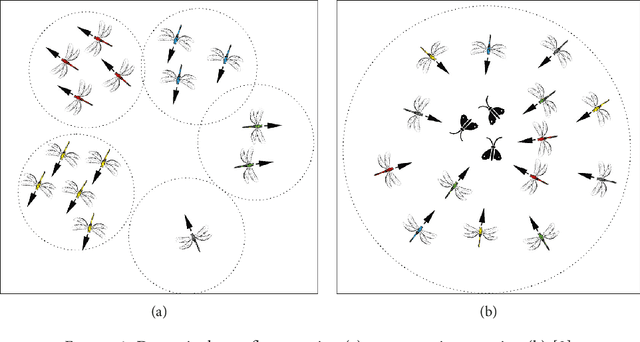
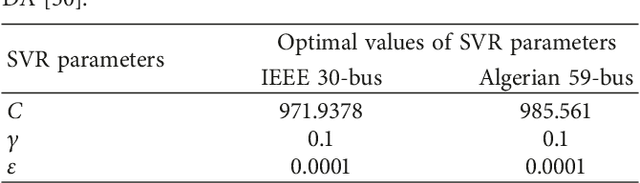
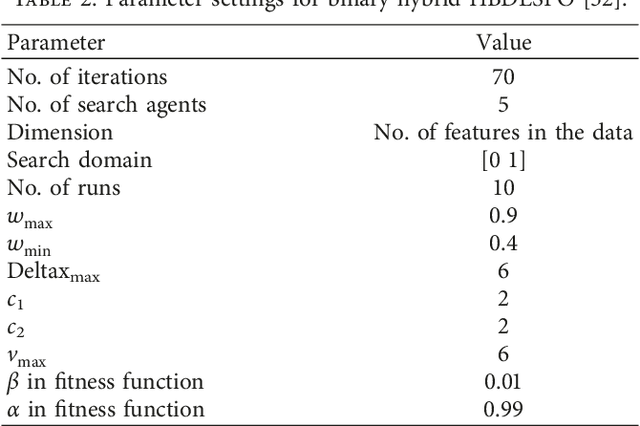
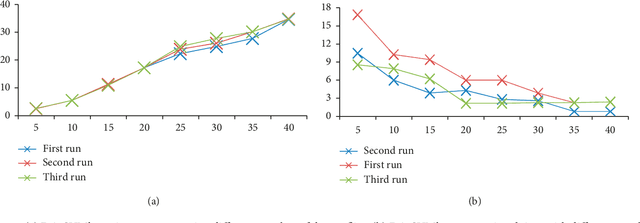
One of the most recently developed heuristic optimization algorithms is dragonfly by Mirjalili. Dragonfly algorithm has shown its ability to optimizing different real world problems. It has three variants. In this work, an overview of the algorithm and its variants is presented. Moreover, the hybridization versions of the algorithm are discussed. Furthermore, the results of the applications that utilized dragonfly algorithm in applied science are offered in the following area: Machine Learning, Image Processing, Wireless, and Networking. It is then compared with some other metaheuristic algorithms. In addition, the algorithm is tested on the CEC-C06 2019 benchmark functions. The results prove that the algorithm has great exploration ability and its convergence rate is better than other algorithms in the literature, such as PSO and GA. In general, in this survey the strong and weak points of the algorithm are discussed. Furthermore, some future works that will help in improving the algorithm's weak points are recommended. This study is conducted with the hope of offering beneficial information about dragonfly algorithm to the researchers who want to study the algorithm.
Improving Super-Resolution Methods via Incremental Residual Learning
Aug 21, 2018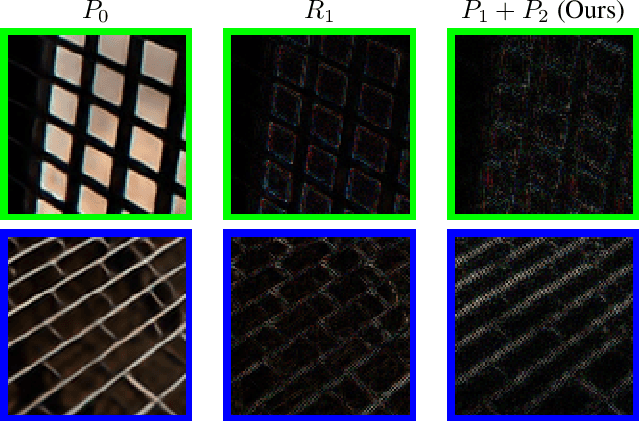
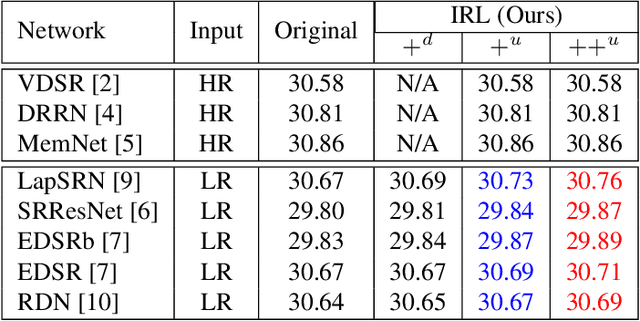
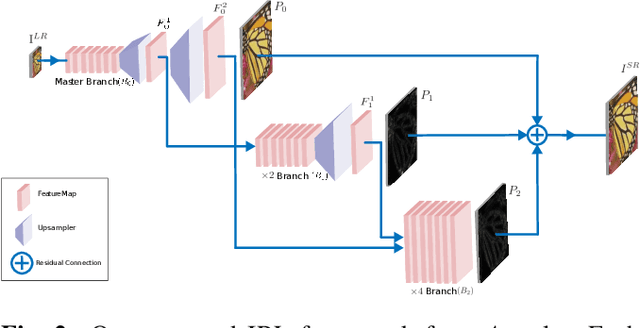
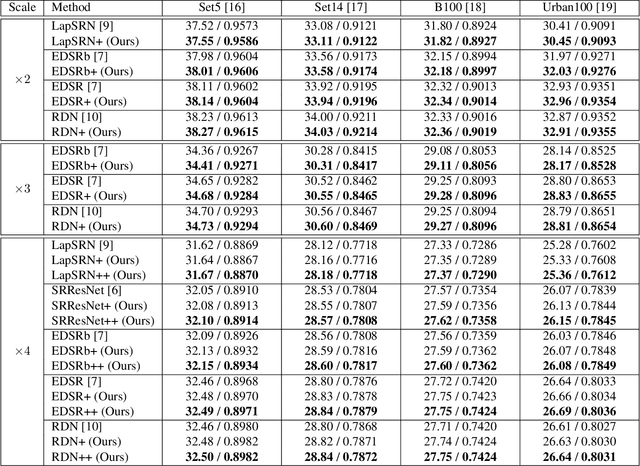
Recently, deep Convolutional Neural Networks (CNNs) have shown promising performance in accurate reconstruction of high resolution (HR) image, given its low resolution (LR) counter-part. However, recent state-of-the-art methods operate primarily on LR image for memory efficiency, but we show that it comes at the cost of performance. Furthermore, because spatial dimensions of input and output of such networks do not match, it's not possible to learn residuals in image space; we show that learning residuals in image space leads to performance enhancement. To this end, we propose a novel Incremental Residual Learning (IRL) framework to solve the above mentioned issues. In IRL, a set of branches i.e arbitrary image-to-image networks are trained sequentially where each branch takes spatially upsampled higher dimensional feature maps as input and predicts the residuals of all previous branches combined. We plug recent state of the art methods as base networks in IRL framework and demonstrate the consistent performance enhancement through extensive experiments on public benchmark datasets to set a new state of the art for super-resolution. Compared to the base networks our method incurs no extra memory overhead as only one branch is trained at a time. Furthermore, as our method is trained to learned residuals, complete set of branches are trained in only 20% of time relative to base network.
Deep Decomposition Learning for Inverse Imaging Problems
Nov 25, 2019
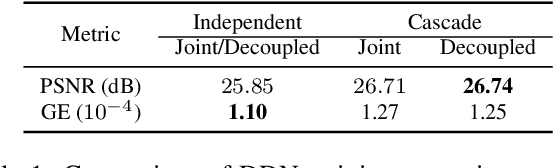

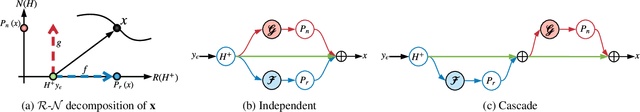

Deep learning is emerging as a new paradigm for solving inverse imaging problems. However, the deep learning methods often lack the assurance of traditional physics-based methods due to the lack of physical information considerations in neural network training and deploying. The appropriate supervision and explicit calibration by the information of the physic model can enhance the neural network learning and its practical performance. In this paper, inspired by the geometry that data can be decomposed by two components from the null-space of the forward operator and the range space of its pseudo-inverse, we train neural networks to learn the two components and therefore learn the decomposition, i.e. we explicitly reformulate the neural network layers as learning range-nullspace decomposition functions with reference to the layer inputs, instead of learning unreferenced functions. We show that the decomposition networks not only produce superior results, but also enjoy good interpretability and generalization. We demonstrate the advantages of decomposition learning on different inverse problems including compressive sensing and image super-resolution as examples.
Deep Probabilistic Modeling of Glioma Growth
Jul 09, 2019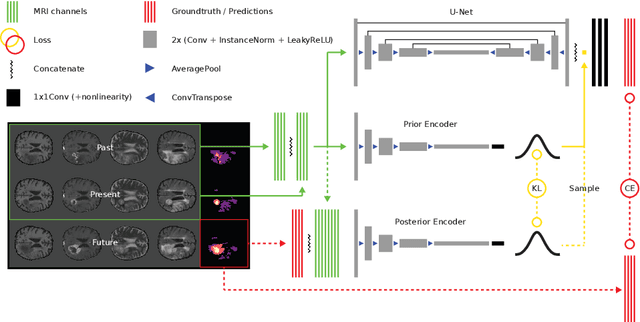
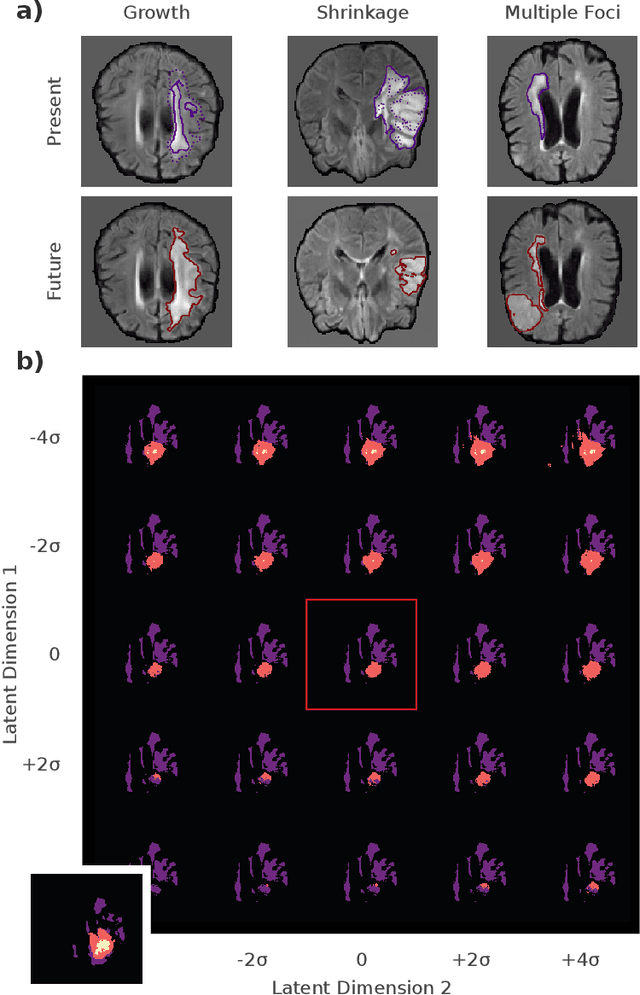
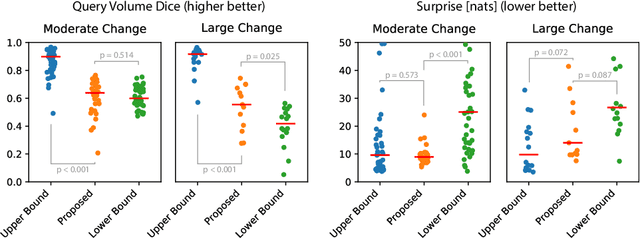
Existing approaches to modeling the dynamics of brain tumor growth, specifically glioma, employ biologically inspired models of cell diffusion, using image data to estimate the associated parameters. In this work, we propose an alternative approach based on recent advances in probabilistic segmentation and representation learning that implicitly learns growth dynamics directly from data without an underlying explicit model. We present evidence that our approach is able to learn a distribution of plausible future tumor appearances conditioned on past observations of the same tumor.
ColorFool: Semantic Adversarial Colorization
Nov 25, 2019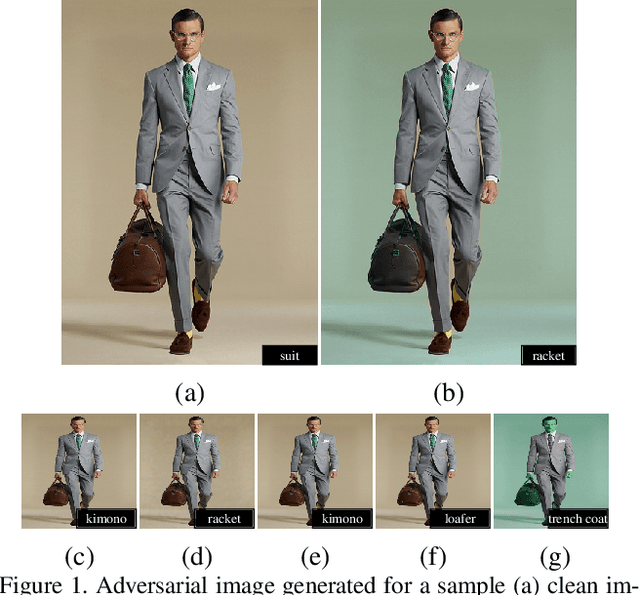
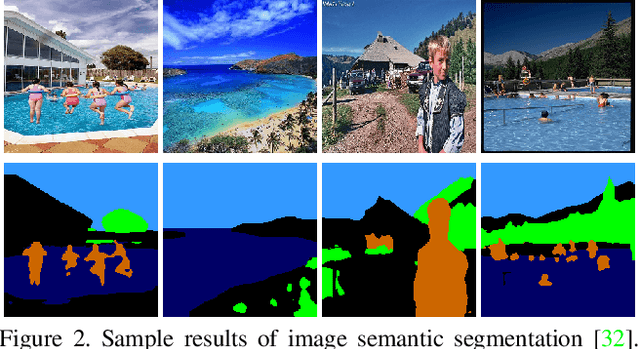



Adversarial attacks that generate small L_p-norm perturbations to mislead classifiers have limited success in black-box settings and with unseen classifiers. These attacks are also fragile with defenses that use denoising filters and to adversarial training procedures. Instead, adversarial attacks that generate unrestricted perturbations are more robust to defenses, are generally more successful in black-box settings and are more transferable to unseen classifiers. However, unrestricted perturbations may be noticeable to humans. In this paper, we propose a content-based black-box adversarial attack that generates unrestricted perturbations by exploiting image semantics to selectively modify colors within chosen ranges that are perceived as natural by humans. We show that the proposed approach, ColorFool, outperforms in terms of success rate, robustness to defense frameworks and transferability five state-of-the-art adversarial attacks on two different tasks, scene and object classification, when attacking three state-of-the-art deep neural networks using three standard datasets. We will make the code of the proposed approach and the whole evaluation framework publicly available.
Deep Cytometry
Apr 09, 2019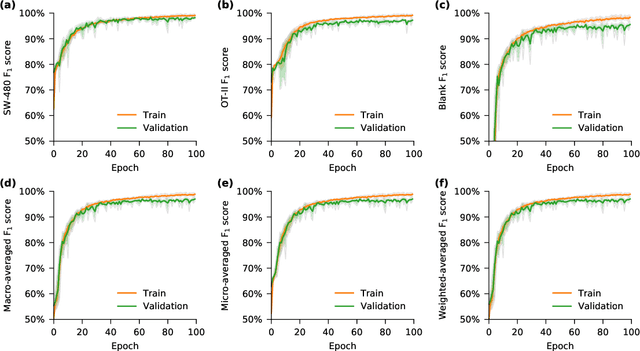
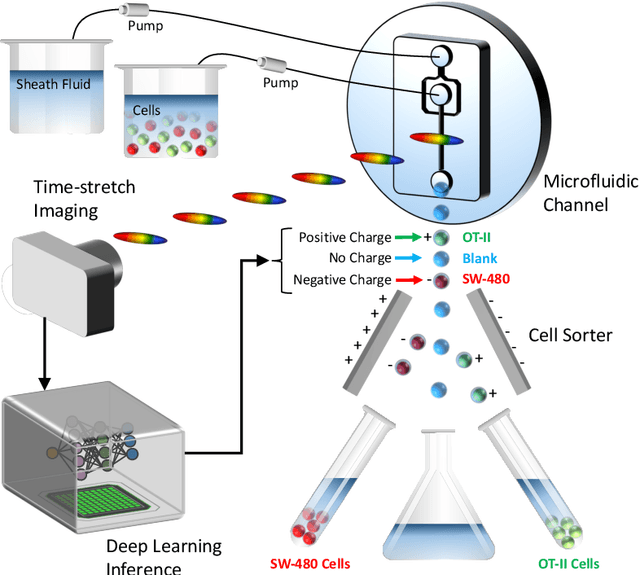
Deep learning has achieved spectacular performance in image and speech recognition and synthesis. It outperforms other machine learning algorithms in problems where large amounts of data are available. In the area of measurement technology, instruments based on the Photonic Time Stretch have established record real-time measurement throughput in spectroscopy, optical coherence tomography, and imaging flow cytometry. These extreme-throughput instruments generate approximately 1 Tbit/s of continuous measurement data and have led to the discovery of rare phenomena in nonlinear and complex systems as well as new types of biomedical instruments. Owing to the abundance of data they generate, time stretch instruments are a natural fit to deep learning classification. Previously we had shown that high-throughput label-free cell classification with high accuracy can be achieved through a combination of time stretch microscopy, image processing and feature extraction, followed by deep learning for finding cancer cells in the blood. Such a technology holds promise for early detection of primary cancer or metastasis. Here we describe a new implementation of deep learning which entirely avoids the computationally costly image processing and feature extraction pipeline. The improvement in computational efficiency makes this new technology suitable for cell sorting via deep learning. Our neural network takes less than a millisecond to classify the cells, fast enough to provide a decision to a cell sorter. We demonstrate the applicability of our new method in the classification of OT-II white blood cells and SW-480 epithelial cancer cells with more than 95\% accuracy in a label-free fashion.
Visuallly Grounded Generation of Entailments from Premises
Sep 21, 2019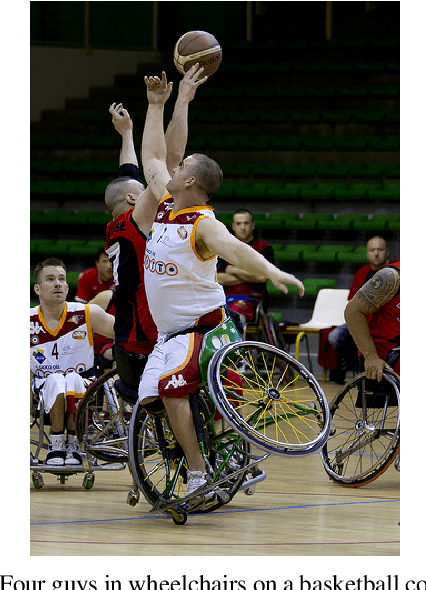
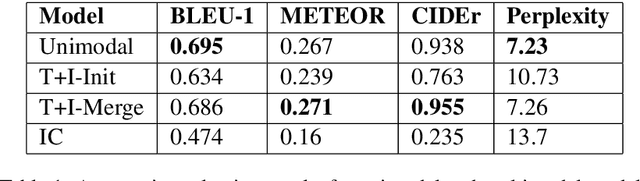
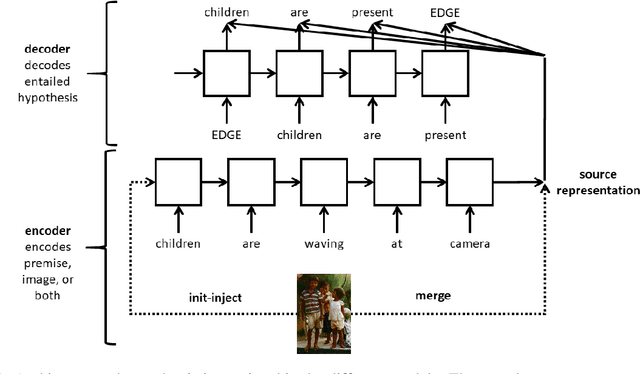

Natural Language Inference (NLI) is the task of determining the semantic relationship between a premise and a hypothesis. In this paper, we focus on the {\em generation} of hypotheses from premises in a multimodal setting, to generate a sentence (hypothesis) given an image and/or its description (premise) as the input. The main goals of this paper are (a) to investigate whether it is reasonable to frame NLI as a generation task; and (b) to consider the degree to which grounding textual premises in visual information is beneficial to generation. We compare different neural architectures, showing through automatic and human evaluation that entailments can indeed be generated successfully. We also show that multimodal models outperform unimodal models in this task, albeit marginally.
Automatic Whole-body Bone Age Assessment Using Deep Hierarchical Features
Jan 29, 2019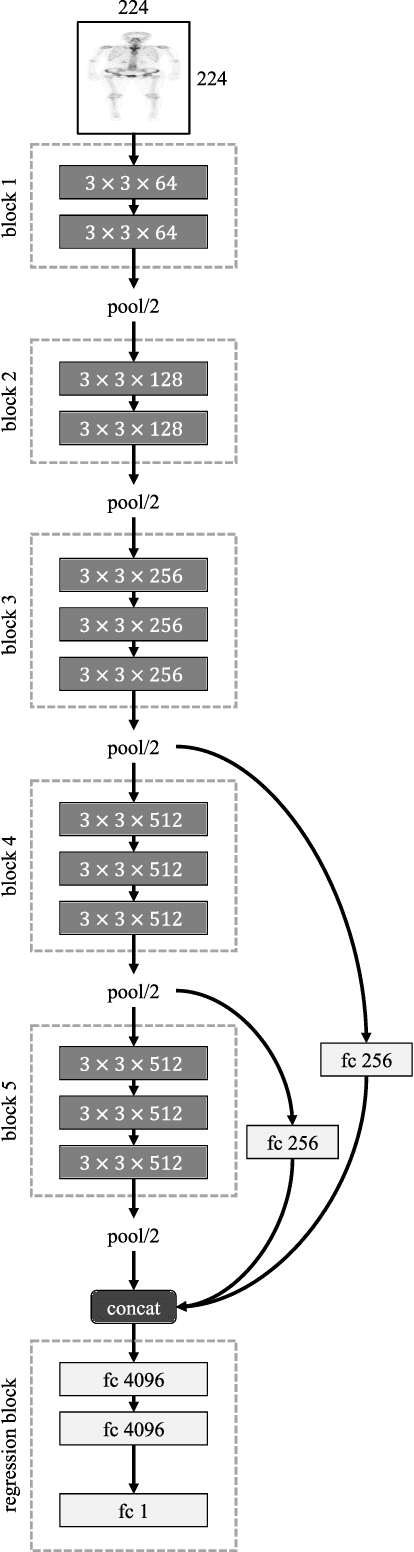
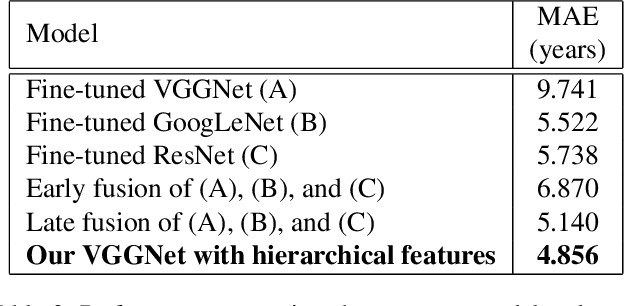


Bone age assessment gives us evidence to analyze the children growth status and the rejuvenation involved chronological and biological ages. All the previous works consider left-hand X-ray image of a child in their works. In this paper, we carry out a study on estimating human age using whole-body bone CT images and a novel convolutional neural network. Our model with additional connections shows an effective way to generate a massive number of vital features while reducing overfitting influence on small training data in the medical image analysis research area. A dataset and a comparison with common deep architectures will be provided for future research in this field.
A Synchronized Multi-Modal Attention-Caption Dataset and Analysis
Mar 06, 2019
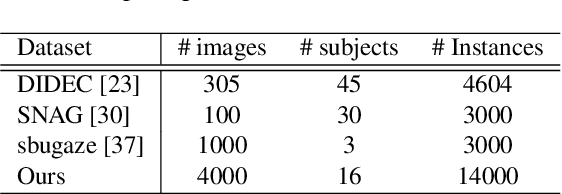
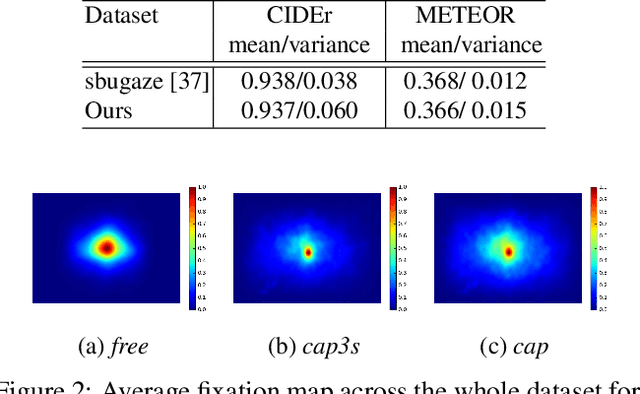

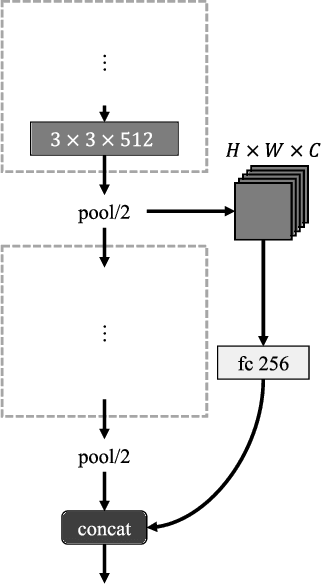
In this work, we present a novel multi-modal dataset consisting of eye movements and verbal descriptions recorded synchronously over images. Using this data, we study the differences between human attention in free-viewing and image captioning tasks. We look into the relationship between human attention and language constructs during perception and sentence articulation. We also compare human and machine attention, in particular the top-down soft attention approach that is argued to mimick human attention, in captioning tasks. Our study reveals that, (1) human attention behaviour in free-viewing is different than image description as humans tend to fixate on a greater variety of regions under the latter task; (2) there is a strong relationship between the described objects and the objects attended by subjects ($97\%$ of described objects are being attended); (3) a convolutional neural network as feature encoder captures regions that human attend under image captioning to a great extent (around $78\%$); (4) the soft-attention as the top-down mechanism does not agree with human attention behaviour neither spatially nor temporally; and (5) soft-attention does not add strong beneficial human-like attention behaviour for the task of captioning as it has low correlation between caption scores and attention consistency scores, indicating a large gap between human and machine in regard to top-down attention.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge