"Image": models, code, and papers
Learning beamforming in ultrasound imaging
Dec 19, 2018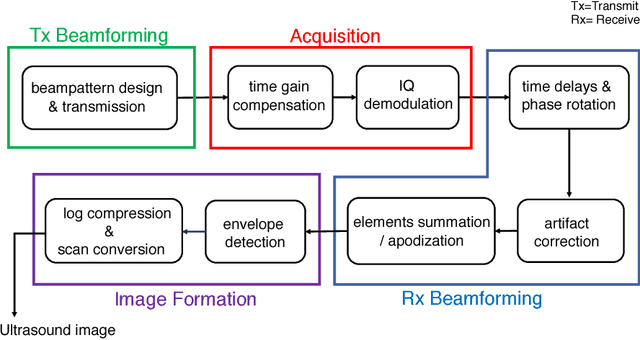
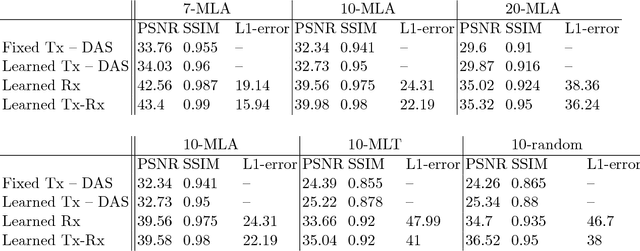
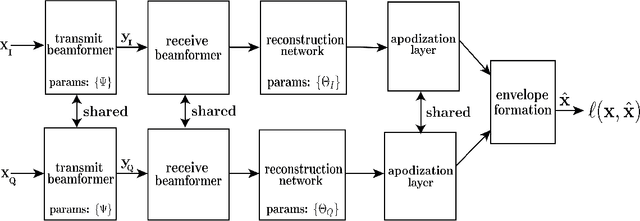
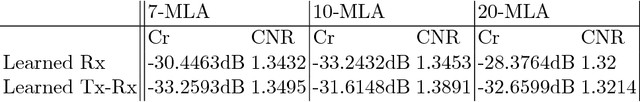
Medical ultrasound (US) is a widespread imaging modality owing its popularity to cost efficiency, portability, speed, and lack of harmful ionizing radiation. In this paper, we demonstrate that replacing the traditional ultrasound processing pipeline with a data-driven, learnable counterpart leads to significant improvement in image quality. Moreover, we demonstrate that greater improvement can be achieved through a learning-based design of the transmitted beam patterns simultaneously with learning an image reconstruction pipeline. We evaluate our method on an in-vivo first-harmonic cardiac ultrasound dataset acquired from volunteers and demonstrate the significance of the learned pipeline and transmit beam patterns on the image quality when compared to standard transmit and receive beamformers used in high frame-rate US imaging. We believe that the presented methodology provides a fundamentally different perspective on the classical problem of ultrasound beam pattern design.
CEREALS - Cost-Effective REgion-based Active Learning for Semantic Segmentation
Oct 23, 2018
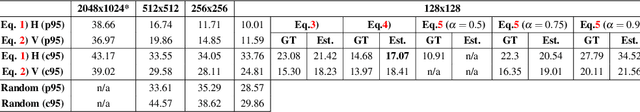
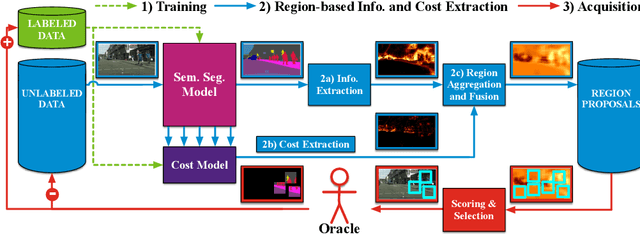
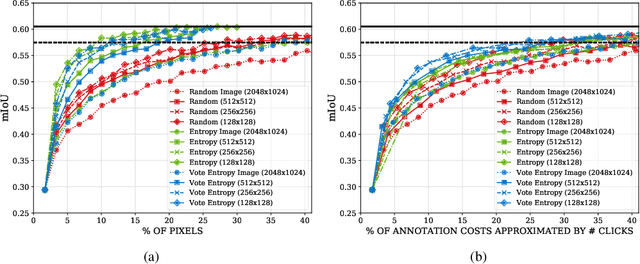
State of the art methods for semantic image segmentation are trained in a supervised fashion using a large corpus of fully labeled training images. However, gathering such a corpus is expensive, due to human annotation effort, in contrast to gathering unlabeled data. We propose an active learning-based strategy, called CEREALS, in which a human only has to hand-label a few, automatically selected, regions within an unlabeled image corpus. This minimizes human annotation effort while maximizing the performance of a semantic image segmentation method. The automatic selection procedure is achieved by: a) using a suitable information measure combined with an estimate about human annotation effort, which is inferred from a learned cost model, and b) exploiting the spatial coherency of an image. The performance of CEREALS is demonstrated on Cityscapes, where we are able to reduce the annotation effort to 17%, while keeping 95% of the mean Intersection over Union (mIoU) of a model that was trained with the fully annotated training set of Cityscapes.
Single-Path NAS: Device-Aware Efficient ConvNet Design
May 10, 2019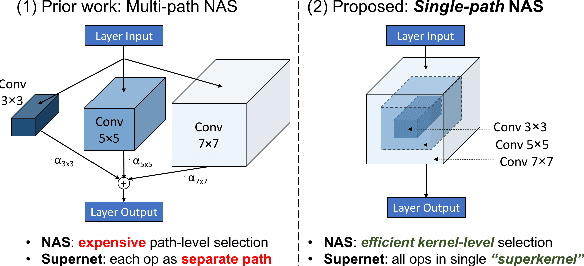
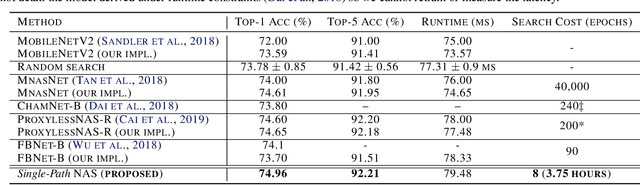
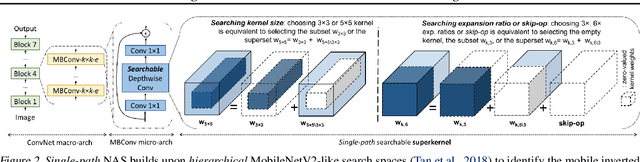
Can we automatically design a Convolutional Network (ConvNet) with the highest image classification accuracy under the latency constraint of a mobile device? Neural Architecture Search (NAS) for ConvNet design is a challenging problem due to the combinatorially large design space and search time (at least 200 GPU-hours). To alleviate this complexity, we propose Single-Path NAS, a novel differentiable NAS method for designing device-efficient ConvNets in less than 4 hours. 1. Novel NAS formulation: our method introduces a single-path, over-parameterized ConvNet to encode all architectural decisions with shared convolutional kernel parameters. 2. NAS efficiency: Our method decreases the NAS search cost down to 8 epochs (30 TPU-hours), i.e., up to 5,000x faster compared to prior work. 3. On-device image classification: Single-Path NAS achieves 74.96% top-1 accuracy on ImageNet with 79ms inference latency on a Pixel 1 phone, which is state-of-the-art accuracy compared to NAS methods with similar latency (<80ms).
Neural Ordinary Differential Equations for Semantic Segmentation of Individual Colon Glands
Oct 23, 2019

Automated medical image segmentation plays a key role in quantitative research and diagnostics. Convolutional neural networks based on the U-Net architecture are the state-of-the-art. A key disadvantage is the hard-coding of the receptive field size, which requires architecture optimization for each segmentation task. Furthermore, increasing the receptive field results in an increasing number of weights. Recently, Neural Ordinary Differential Equations (NODE) have been proposed, a new type of continuous depth deep neural network. This framework allows for a dynamic receptive field at a fixed memory cost and a smaller amount of parameters. We show on a colon gland segmentation dataset (GlaS) that these NODEs can be used within the U-Net framework to improve segmentation results while reducing memory load and parameter counts.
Revisiting Landscape Analysis in Deep Neural Networks: Eliminating Decreasing Paths to Infinity
Dec 31, 2019Traditional landscape analysis of deep neural networks aims to show that no sub-optimal local minima exist in some appropriate sense. From this, one may be tempted to conclude that descent algorithms which escape saddle points will reach a good local minimum. However, basic optimization theory tell us that it is also possible for a descent algorithm to diverge to infinity if there are paths leading to infinity, along which the loss function decreases. It is not clear whether for non-linear neural networks there exists one setting that no bad local-min and no decreasing paths to infinity can be simultaneously achieved. In this paper, we give the first positive answer to this question. More specifically, for a large class of over-parameterized deep neural networks with appropriate regularizers, the loss function has no bad local minima and no decreasing paths to infinity. The key mathematical trick is to show that the set of regularizers which may be undesirable can be viewed as the image of a Lipschitz continuous mapping from a lower-dimensional Euclidean space to a higher-dimensional Euclidean space, and thus has zero measure.
Fast Compressive Sensing Recovery Using Generative Models with Structured Latent Variables
Feb 22, 2019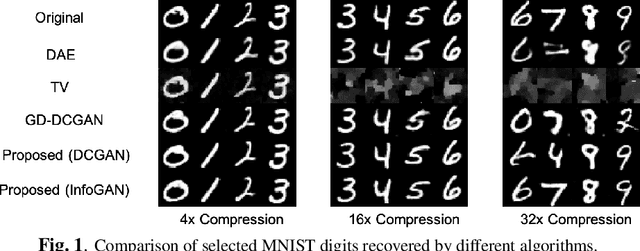
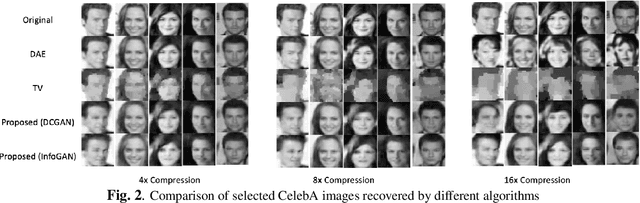
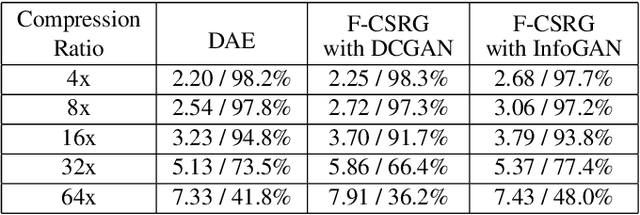

Deep learning models have significantly improved the visual quality and accuracy on compressive sensing recovery. In this paper, we propose an algorithm for signal reconstruction from compressed measurements with image priors captured by a generative model. We search and constrain on latent variable space to make the method stable when the number of compressed measurements is extremely limited. We show that, by exploiting certain structures of the latent variables, the proposed method produces improved reconstruction accuracy and preserves realistic and non-smooth features in the image. Our algorithm achieves high computation speed by projecting between the original signal space and the latent variable space in an alternating fashion.
A Non-Local Structure Tensor Based Approach for Multicomponent Image Recovery Problems
Oct 14, 2014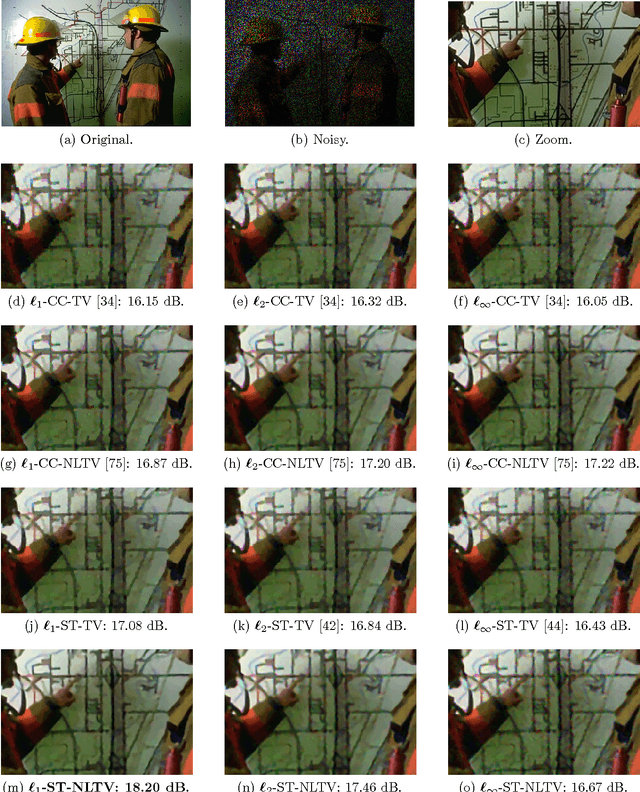
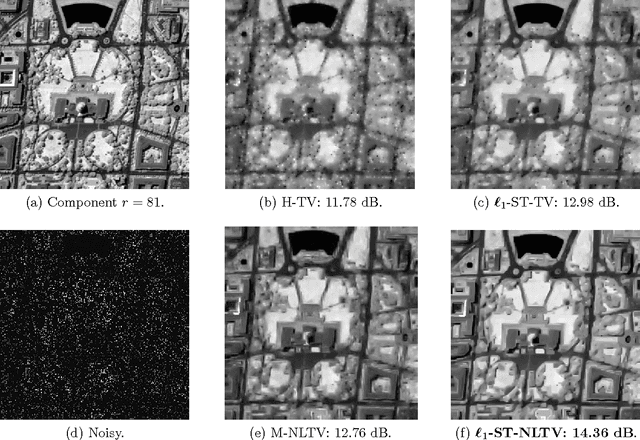
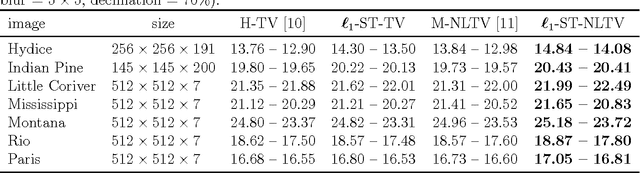
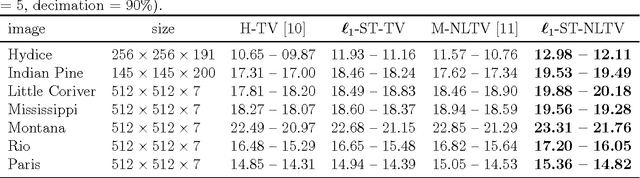


Non-Local Total Variation (NLTV) has emerged as a useful tool in variational methods for image recovery problems. In this paper, we extend the NLTV-based regularization to multicomponent images by taking advantage of the Structure Tensor (ST) resulting from the gradient of a multicomponent image. The proposed approach allows us to penalize the non-local variations, jointly for the different components, through various $\ell_{1,p}$ matrix norms with $p \ge 1$. To facilitate the choice of the hyper-parameters, we adopt a constrained convex optimization approach in which we minimize the data fidelity term subject to a constraint involving the ST-NLTV regularization. The resulting convex optimization problem is solved with a novel epigraphical projection method. This formulation can be efficiently implemented thanks to the flexibility offered by recent primal-dual proximal algorithms. Experiments are carried out for multispectral and hyperspectral images. The results demonstrate the interest of introducing a non-local structure tensor regularization and show that the proposed approach leads to significant improvements in terms of convergence speed over current state-of-the-art methods.
Deep weakly-supervised learning methods for classification and localization in histology images: a survey
Sep 26, 2019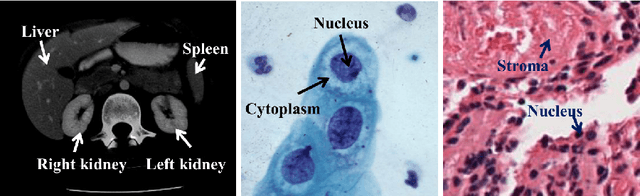
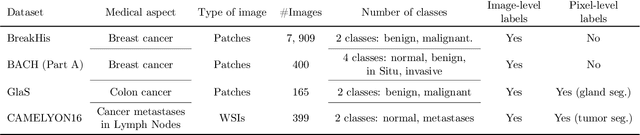

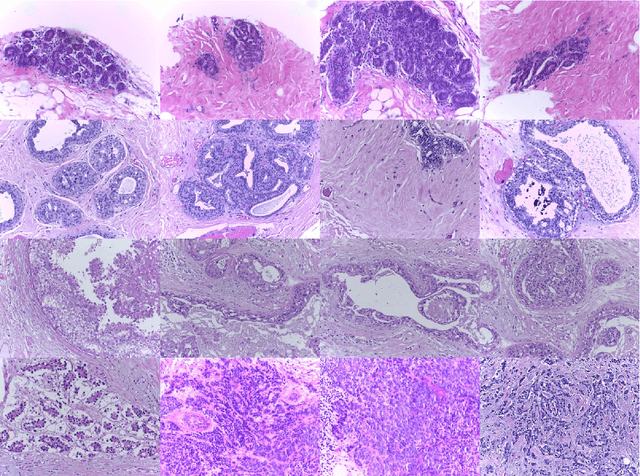
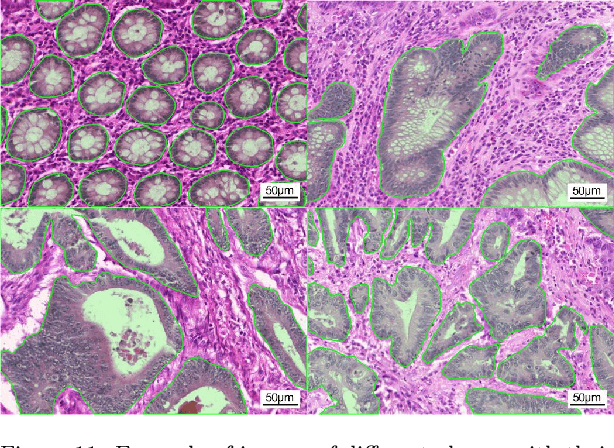
Using state-of-the-art deep learning models for the computer-assisted diagnosis of diseases like cancer raises several challenges related to the nature and availability of labeled histology images. In particular, cancer grading and localization in these images normally relies on both image- and pixel-level labels, the latter requiring a costly annotation process. In this survey, deep weakly-supervised learning (WSL) architectures are investigated to identify and locate diseases in histology image, without the need for pixel-level annotations. Given a training dataset with globally-annotated images, these models allow to simultaneously classify histology images, while localizing the corresponding regions of interest. These models are organized into two main approaches -- (1) bottom-up approaches (based on forward-pass information through a network, either by spatial pooling of representations/scores, or by detecting class regions), and (2) top-down approaches (based on backward-pass information within a network, inspired by human visual attention). Since relevant WSL models have mainly been developed in the computer vision community, and validated on natural scene images, we assess the extent to which they apply to histology images which have challenging properties, e.g., large size, non-salient and highly unstructured regions, stain heterogeneity, and coarse/ambiguous labels. The most relevant deep WSL models (e.g., CAM, WILDCAT and Deep MIL) are compared experimentally in terms of accuracy (classification and pixel-level localization) on several public benchmark histology datasets for breast and colon cancer (BACH ICIAR 2018, BreakHis, CAMELYON16, and GlaS). Results indicate that several deep learning models, and in particular WILDCAT and deep MIL can provide a high level of classification accuracy, although pixel-wise localization of cancer regions remains an issue for such images.
Image denoising with multi-layer perceptrons, part 1: comparison with existing algorithms and with bounds
Nov 09, 2012
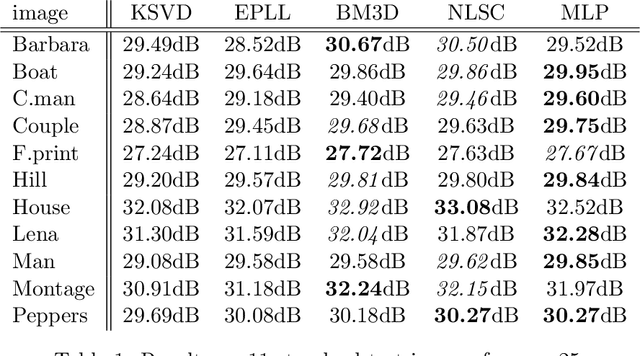
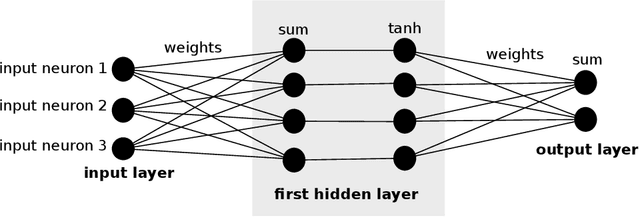
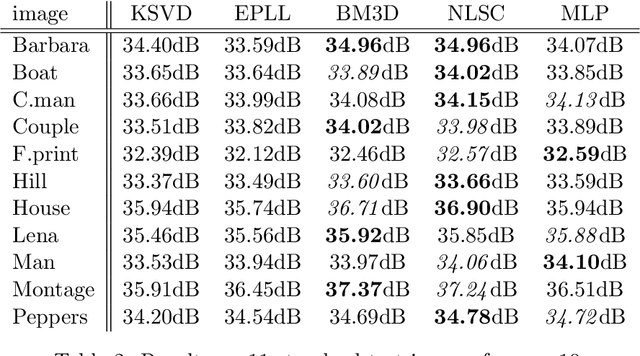
Image denoising can be described as the problem of mapping from a noisy image to a noise-free image. The best currently available denoising methods approximate this mapping with cleverly engineered algorithms. In this work we attempt to learn this mapping directly with plain multi layer perceptrons (MLP) applied to image patches. We will show that by training on large image databases we are able to outperform the current state-of-the-art image denoising methods. In addition, our method achieves results that are superior to one type of theoretical bound and goes a large way toward closing the gap with a second type of theoretical bound. Our approach is easily adapted to less extensively studied types of noise, such as mixed Poisson-Gaussian noise, JPEG artifacts, salt-and-pepper noise and noise resembling stripes, for which we achieve excellent results as well. We will show that combining a block-matching procedure with MLPs can further improve the results on certain images. In a second paper, we detail the training trade-offs and the inner mechanisms of our MLPs.
Training Deep Learning models with small datasets
Dec 14, 2019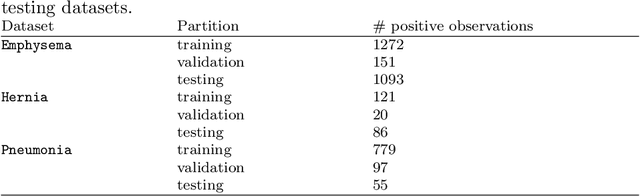
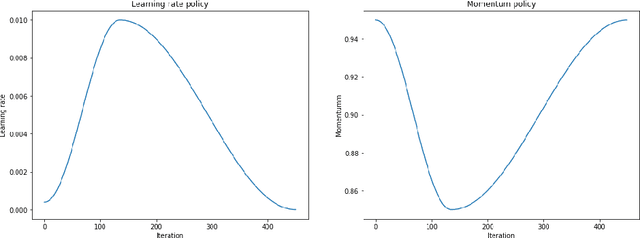
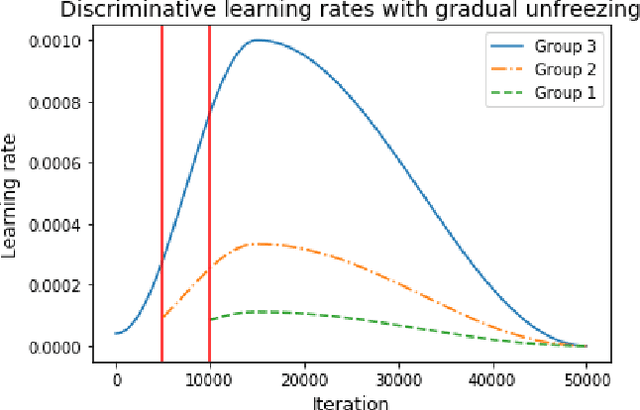
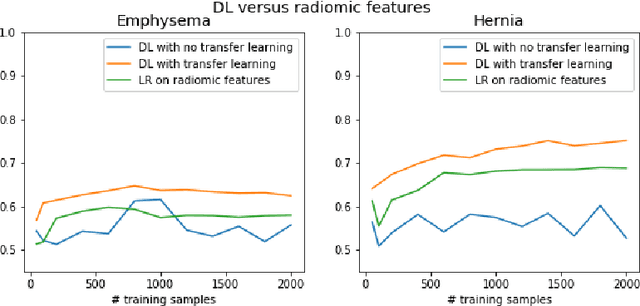
The growing use of Machine Learning has produced significant advances in many fields. For image-based tasks, however, the use of deep learning remains challenging in small datasets. In this article, we review, evaluate and compare the current state of the art techniques in training neural networks to elucidate which techniques work best for small datasets. We further propose a path forward for the improvement of model accuracy in medical imaging applications. We observed best results from one cycle training, discriminative learning rates with gradual freezing and parameter modification after transfer learning. We also established that when datasets are small, transfer learning plays an important role beyond parameter initialization by reusing previously learned features. Surprisingly we observed that there is little advantage in using pre-trained networks in images from another part of the body compared to Imagenet. On the contrary, if images from the same part of the body are available then transfer learning can produce a significant improvement in performance with as little as 50 images in the training data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge