"Image": models, code, and papers
Towards Robust and Expressive Whole-body Human Pose and Shape Estimation
Dec 14, 2023
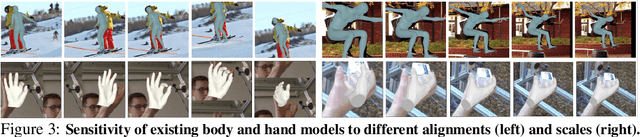


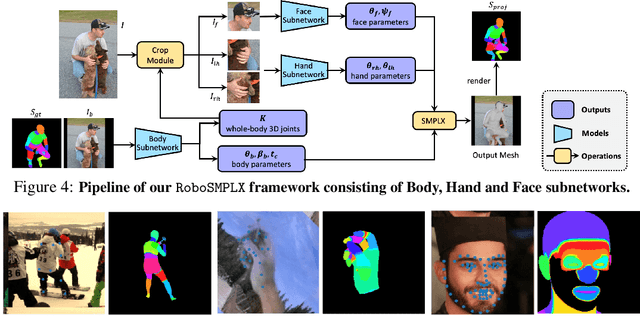
Whole-body pose and shape estimation aims to jointly predict different behaviors (e.g., pose, hand gesture, facial expression) of the entire human body from a monocular image. Existing methods often exhibit degraded performance under the complexity of in-the-wild scenarios. We argue that the accuracy and reliability of these models are significantly affected by the quality of the predicted \textit{bounding box}, e.g., the scale and alignment of body parts. The natural discrepancy between the ideal bounding box annotations and model detection results is particularly detrimental to the performance of whole-body pose and shape estimation. In this paper, we propose a novel framework to enhance the robustness of whole-body pose and shape estimation. Our framework incorporates three new modules to address the above challenges from three perspectives: \textbf{1) Localization Module} enhances the model's awareness of the subject's location and semantics within the image space. \textbf{2) Contrastive Feature Extraction Module} encourages the model to be invariant to robust augmentations by incorporating contrastive loss with dedicated positive samples. \textbf{3) Pixel Alignment Module} ensures the reprojected mesh from the predicted camera and body model parameters are accurate and pixel-aligned. We perform comprehensive experiments to demonstrate the effectiveness of our proposed framework on body, hands, face and whole-body benchmarks. Codebase is available at \url{https://github.com/robosmplx/robosmplx}.
Factorization Vision Transformer: Modeling Long Range Dependency with Local Window Cost
Dec 14, 2023Transformers have astounding representational power but typically consume considerable computation which is quadratic with image resolution. The prevailing Swin transformer reduces computational costs through a local window strategy. However, this strategy inevitably causes two drawbacks: (1) the local window-based self-attention hinders global dependency modeling capability; (2) recent studies point out that local windows impair robustness. To overcome these challenges, we pursue a preferable trade-off between computational cost and performance. Accordingly, we propose a novel factorization self-attention mechanism (FaSA) that enjoys both the advantages of local window cost and long-range dependency modeling capability. By factorizing the conventional attention matrix into sparse sub-attention matrices, FaSA captures long-range dependencies while aggregating mixed-grained information at a computational cost equivalent to the local window-based self-attention. Leveraging FaSA, we present the factorization vision transformer (FaViT) with a hierarchical structure. FaViT achieves high performance and robustness, with linear computational complexity concerning input image spatial resolution. Extensive experiments have shown FaViT's advanced performance in classification and downstream tasks. Furthermore, it also exhibits strong model robustness to corrupted and biased data and hence demonstrates benefits in favor of practical applications. In comparison to the baseline model Swin-T, our FaViT-B2 significantly improves classification accuracy by 1% and robustness by 7%, while reducing model parameters by 14%. Our code will soon be publicly available at https://github.com/q2479036243/FaViT.
Morphological Profiling for Drug Discovery in the Era of Deep Learning
Dec 13, 2023Morphological profiling is a valuable tool in phenotypic drug discovery. The advent of high-throughput automated imaging has enabled the capturing of a wide range of morphological features of cells or organisms in response to perturbations at the single-cell resolution. Concurrently, significant advances in machine learning and deep learning, especially in computer vision, have led to substantial improvements in analyzing large-scale high-content images at high-throughput. These efforts have facilitated understanding of compound mechanism-of-action (MOA), drug repurposing, characterization of cell morphodynamics under perturbation, and ultimately contributing to the development of novel therapeutics. In this review, we provide a comprehensive overview of the recent advances in the field of morphological profiling. We summarize the image profiling analysis workflow, survey a broad spectrum of analysis strategies encompassing feature engineering- and deep learning-based approaches, and introduce publicly available benchmark datasets. We place a particular emphasis on the application of deep learning in this pipeline, covering cell segmentation, image representation learning, and multimodal learning. Additionally, we illuminate the application of morphological profiling in phenotypic drug discovery and highlight potential challenges and opportunities in this field.
RadOcc: Learning Cross-Modality Occupancy Knowledge through Rendering Assisted Distillation
Dec 19, 20233D occupancy prediction is an emerging task that aims to estimate the occupancy states and semantics of 3D scenes using multi-view images. However, image-based scene perception encounters significant challenges in achieving accurate prediction due to the absence of geometric priors. In this paper, we address this issue by exploring cross-modal knowledge distillation in this task, i.e., we leverage a stronger multi-modal model to guide the visual model during training. In practice, we observe that directly applying features or logits alignment, proposed and widely used in bird's-eyeview (BEV) perception, does not yield satisfactory results. To overcome this problem, we introduce RadOcc, a Rendering assisted distillation paradigm for 3D Occupancy prediction. By employing differentiable volume rendering, we generate depth and semantic maps in perspective views and propose two novel consistency criteria between the rendered outputs of teacher and student models. Specifically, the depth consistency loss aligns the termination distributions of the rendered rays, while the semantic consistency loss mimics the intra-segment similarity guided by vision foundation models (VLMs). Experimental results on the nuScenes dataset demonstrate the effectiveness of our proposed method in improving various 3D occupancy prediction approaches, e.g., our proposed methodology enhances our baseline by 2.2% in the metric of mIoU and achieves 50% in Occ3D benchmark.
Adaptive Compression of the Latent Space in Variational Autoencoders
Dec 11, 2023Variational Autoencoders (VAEs) are powerful generative models that have been widely used in various fields, including image and text generation. However, one of the known challenges in using VAEs is the model's sensitivity to its hyperparameters, such as the latent space size. This paper presents a simple extension of VAEs for automatically determining the optimal latent space size during the training process by gradually decreasing the latent size through neuron removal and observing the model performance. The proposed method is compared to traditional hyperparameter grid search and is shown to be significantly faster while still achieving the best optimal dimensionality on four image datasets. Furthermore, we show that the final performance of our method is comparable to training on the optimal latent size from scratch, and might thus serve as a convenient substitute.
BeautifulPrompt: Towards Automatic Prompt Engineering for Text-to-Image Synthesis
Nov 12, 2023Recently, diffusion-based deep generative models (e.g., Stable Diffusion) have shown impressive results in text-to-image synthesis. However, current text-to-image models often require multiple passes of prompt engineering by humans in order to produce satisfactory results for real-world applications. We propose BeautifulPrompt, a deep generative model to produce high-quality prompts from very simple raw descriptions, which enables diffusion-based models to generate more beautiful images. In our work, we first fine-tuned the BeautifulPrompt model over low-quality and high-quality collecting prompt pairs. Then, to ensure that our generated prompts can generate more beautiful images, we further propose a Reinforcement Learning with Visual AI Feedback technique to fine-tune our model to maximize the reward values of the generated prompts, where the reward values are calculated based on the PickScore and the Aesthetic Scores. Our results demonstrate that learning from visual AI feedback promises the potential to improve the quality of generated prompts and images significantly. We further showcase the integration of BeautifulPrompt to a cloud-native AI platform to provide better text-to-image generation service in the cloud.
Synthesizing Black-box Anti-forensics DeepFakes with High Visual Quality
Dec 17, 2023DeepFake, an AI technology for creating facial forgeries, has garnered global attention. Amid such circumstances, forensics researchers focus on developing defensive algorithms to counter these threats. In contrast, there are techniques developed for enhancing the aggressiveness of DeepFake, e.g., through anti-forensics attacks, to disrupt forensic detectors. However, such attacks often sacrifice image visual quality for improved undetectability. To address this issue, we propose a method to generate novel adversarial sharpening masks for launching black-box anti-forensics attacks. Unlike many existing arts, with such perturbations injected, DeepFakes could achieve high anti-forensics performance while exhibiting pleasant sharpening visual effects. After experimental evaluations, we prove that the proposed method could successfully disrupt the state-of-the-art DeepFake detectors. Besides, compared with the images processed by existing DeepFake anti-forensics methods, the visual qualities of anti-forensics DeepFakes rendered by the proposed method are significantly refined.
Data-driven Crop Growth Simulation on Time-varying Generated Images using Multi-conditional Generative Adversarial Networks
Dec 06, 2023Image-based crop growth modeling can substantially contribute to precision agriculture by revealing spatial crop development over time, which allows an early and location-specific estimation of relevant future plant traits, such as leaf area or biomass. A prerequisite for realistic and sharp crop image generation is the integration of multiple growth-influencing conditions in a model, such as an image of an initial growth stage, the associated growth time, and further information about the field treatment. We present a two-stage framework consisting first of an image prediction model and second of a growth estimation model, which both are independently trained. The image prediction model is a conditional Wasserstein generative adversarial network (CWGAN). In the generator of this model, conditional batch normalization (CBN) is used to integrate different conditions along with the input image. This allows the model to generate time-varying artificial images dependent on multiple influencing factors of different kinds. These images are used by the second part of the framework for plant phenotyping by deriving plant-specific traits and comparing them with those of non-artificial (real) reference images. For various crop datasets, the framework allows realistic, sharp image predictions with a slight loss of quality from short-term to long-term predictions. Simulations of varying growth-influencing conditions performed with the trained framework provide valuable insights into how such factors relate to crop appearances, which is particularly useful in complex, less explored crop mixture systems. Further results show that adding process-based simulated biomass as a condition increases the accuracy of the derived phenotypic traits from the predicted images. This demonstrates the potential of our framework to serve as an interface between an image- and process-based crop growth model.
Adversarial Medical Image with Hierarchical Feature Hiding
Dec 04, 2023Deep learning based methods for medical images can be easily compromised by adversarial examples (AEs), posing a great security flaw in clinical decision-making. It has been discovered that conventional adversarial attacks like PGD which optimize the classification logits, are easy to distinguish in the feature space, resulting in accurate reactive defenses. To better understand this phenomenon and reassess the reliability of the reactive defenses for medical AEs, we thoroughly investigate the characteristic of conventional medical AEs. Specifically, we first theoretically prove that conventional adversarial attacks change the outputs by continuously optimizing vulnerable features in a fixed direction, thereby leading to outlier representations in the feature space. Then, a stress test is conducted to reveal the vulnerability of medical images, by comparing with natural images. Interestingly, this vulnerability is a double-edged sword, which can be exploited to hide AEs. We then propose a simple-yet-effective hierarchical feature constraint (HFC), a novel add-on to conventional white-box attacks, which assists to hide the adversarial feature in the target feature distribution. The proposed method is evaluated on three medical datasets, both 2D and 3D, with different modalities. The experimental results demonstrate the superiority of HFC, \emph{i.e.,} it bypasses an array of state-of-the-art adversarial medical AE detectors more efficiently than competing adaptive attacks, which reveals the deficiencies of medical reactive defense and allows to develop more robust defenses in future.
Exploring the Feasibility of Generating Realistic 3D Models of Endangered Species Using DreamGaussian: An Analysis of Elevation Angle's Impact on Model Generation
Dec 15, 2023Many species face the threat of extinction. It's important to study these species and gather information about them as much as possible to preserve biodiversity. Due to the rarity of endangered species, there is a limited amount of data available, making it difficult to apply data requiring generative AI methods to this domain. We aim to study the feasibility of generating consistent and real-like 3D models of endangered animals using limited data. Such a phenomenon leads us to utilize zero-shot stable diffusion models that can generate a 3D model out of a single image of the target species. This paper investigates the intricate relationship between elevation angle and the output quality of 3D model generation, focusing on the innovative approach presented in DreamGaussian. DreamGaussian, a novel framework utilizing Generative Gaussian Splatting along with novel mesh extraction and refinement algorithms, serves as the focal point of our study. We conduct a comprehensive analysis, analyzing the effect of varying elevation angles on DreamGaussian's ability to reconstruct 3D scenes accurately. Through an empirical evaluation, we demonstrate how changes in elevation angle impact the generated images' spatial coherence, structural integrity, and perceptual realism. We observed that giving a correct elevation angle with the input image significantly affects the result of the generated 3D model. We hope this study to be influential for the usability of AI to preserve endangered animals; while the penultimate aim is to obtain a model that can output biologically consistent 3D models via small samples, the qualitative interpretation of an existing state-of-the-art model such as DreamGaussian will be a step forward in our goal.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge