"Image": models, code, and papers
Image-based Guidance of Autonomous Aircraft for Wildfire Surveillance and Prediction
Mar 01, 2019
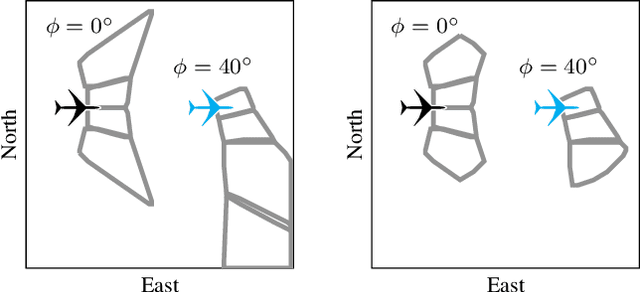
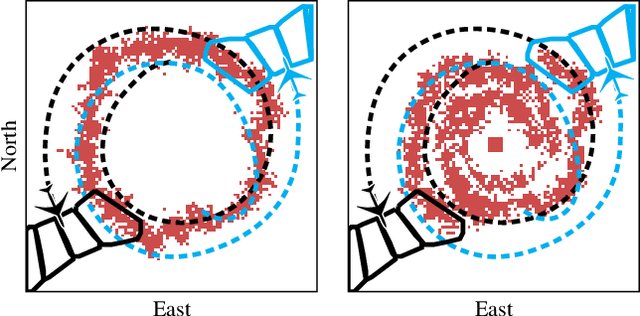
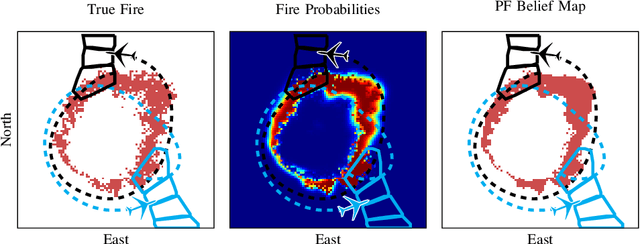
Small unmanned aircraft can help firefighters combat wildfires by providing real-time surveillance of the growing fires. However, guiding the aircraft autonomously given only wildfire images is a challenging problem. This work models noisy images obtained from on-board cameras and proposes two approaches to filtering the wildfire images. The first approach uses a simple Kalman filter to reduce noise and update a belief map in observed areas. The second approach uses a particle filter to predict wildfire growth and uses observations to estimate uncertainties relating to wildfire expansion. The belief maps are used to train a deep reinforcement learning controller, which learns a policy to navigate the aircraft to survey the wildfire while avoiding flight directly over the fire. Simulation results show that the proposed controllers precisely guide the aircraft and accurately estimate wildfire growth, and a study of observation noise demonstrates the robustness of the particle filter approach.
PathVQA: 30000+ Questions for Medical Visual Question Answering
Mar 07, 2020
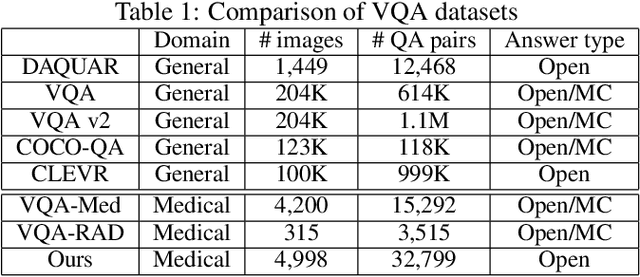
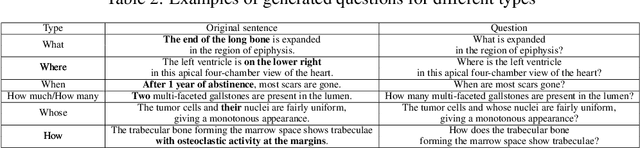
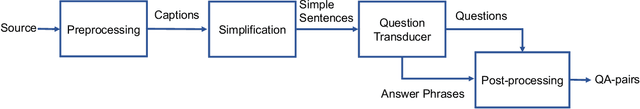
Is it possible to develop an "AI Pathologist" to pass the board-certified examination of the American Board of Pathology? To achieve this goal, the first step is to create a visual question answering (VQA) dataset where the AI agent is presented with a pathology image together with a question and is asked to give the correct answer. Our work makes the first attempt to build such a dataset. Different from creating general-domain VQA datasets where the images are widely accessible and there are many crowdsourcing workers available and capable of generating question-answer pairs, developing a medical VQA dataset is much more challenging. First, due to privacy concerns, pathology images are usually not publicly available. Second, only well-trained pathologists can understand pathology images, but they barely have time to help create datasets for AI research. To address these challenges, we resort to pathology textbooks and online digital libraries. We develop a semi-automated pipeline to extract pathology images and captions from textbooks and generate question-answer pairs from captions using natural language processing. We collect 32,799 open-ended questions from 4,998 pathology images where each question is manually checked to ensure correctness. To our best knowledge, this is the first dataset for pathology VQA. Our dataset will be released publicly to promote research in medical VQA.
On the Validity of Bayesian Neural Networks for Uncertainty Estimation
Dec 03, 2019
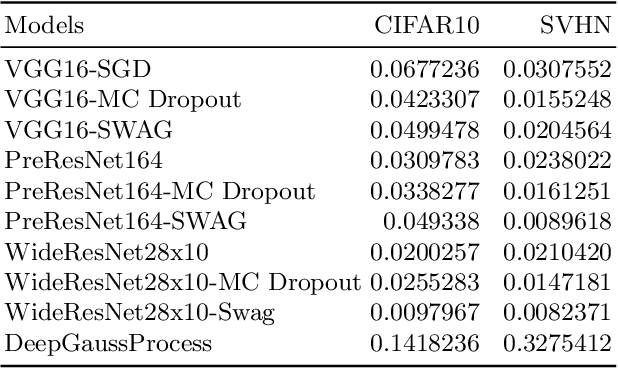
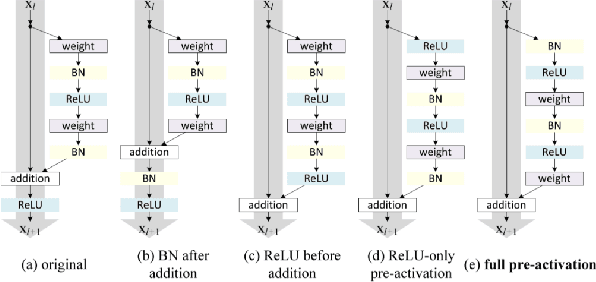
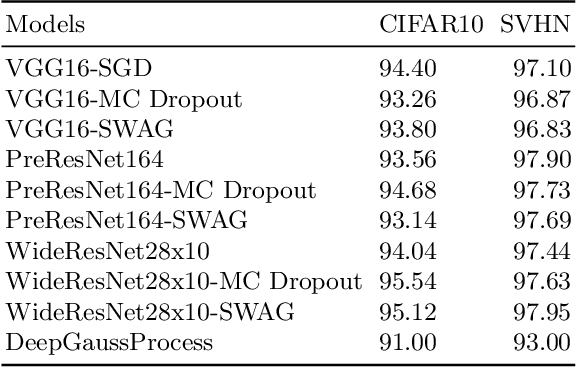
Deep neural networks (DNN) are versatile parametric models utilised successfully in a diverse number of tasks and domains. However, they have limitations---particularly from their lack of robustness and over-sensitivity to out of distribution samples. Bayesian Neural Networks, due to their formulation under the Bayesian framework, provide a principled approach to building neural networks that address these limitations. This paper describes a study that empirically evaluates and compares Bayesian Neural Networks to their equivalent point estimate Deep Neural Networks to quantify the predictive uncertainty induced by their parameters, as well as their performance in view of this uncertainty. In this study, we evaluated and compared three point estimate deep neural networks against comparable Bayesian neural network alternatives using two well-known benchmark image classification datasets (CIFAR-10 and SVHN).
Accelerated Motion-Aware MR Imaging via Motion Prediction from K-Space Center
Aug 26, 2019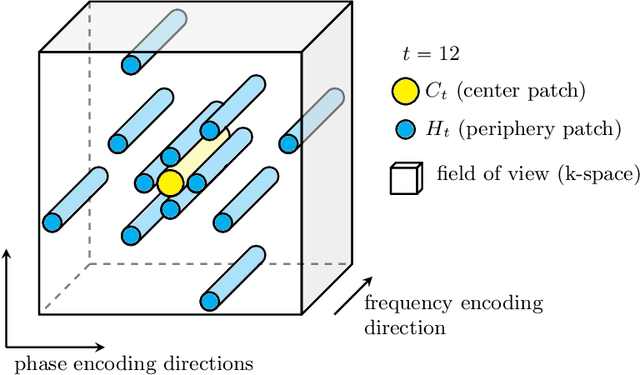
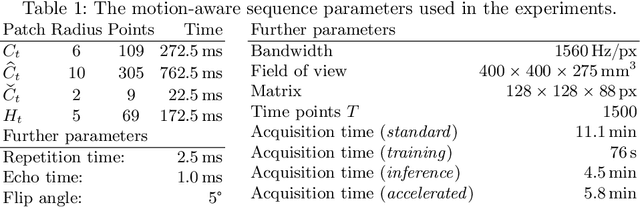

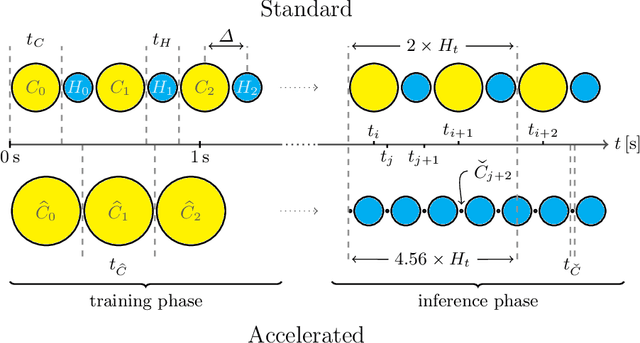
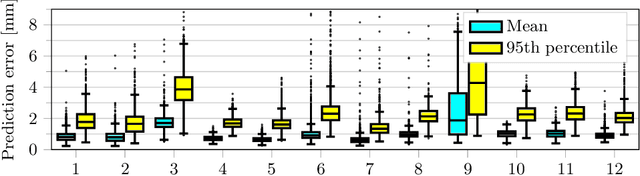
Motion has been a challenge for magnetic resonance (MR) imaging ever since the MR has been invented. Especially in volumetric imaging of thoracic and abdominal organs, motion-awareness is essential for reducing motion artifacts in the final image. A recently proposed MR imaging approach copes with motion by observing the motion patterns during the acquisition. Repetitive scanning of the k-space center region enables the extraction of the patient motion while acquiring the remaining part of the k-space. Due to highly redundant measurements of the center, the required scanning time of over 11 min and the reconstruction time of 2 h exceed clinical applicability though. We propose an accelerated motion-aware MR imaging method where the motion is inferred from small-sized k-space center patches and an initial training phase during which the characteristic movements are modeled. Thereby, acquisition times are reduced by a factor of almost 2 and reconstruction times by two orders of magnitude. Moreover, we improve the existing motion-aware approach with a systematic temporal shift correction to achieve a sharper image reconstruction. We tested our method on 12 volunteers and scanned their lungs and abdomen under free breathing. We achieved equivalent to higher reconstruction quality using the motion-prediction compared to the slower existing approach.
Total Denoising: Unsupervised Learning of 3D Point Cloud Cleaning
Apr 16, 2019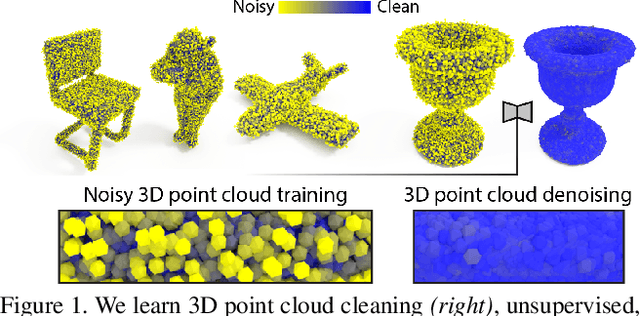
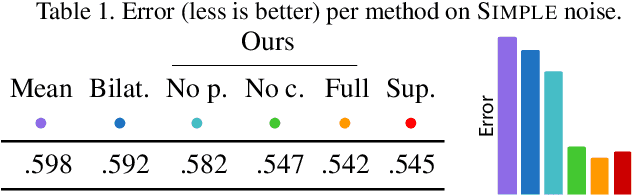

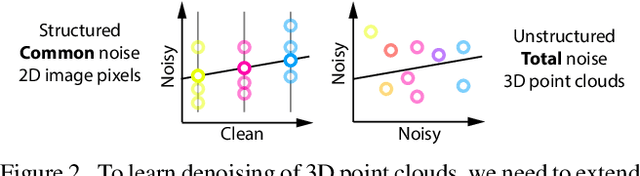
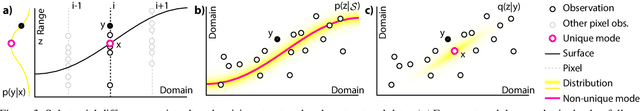
We show that denoising of 3D point clouds can be learned unsupervised, directly from noisy 3D point cloud data only. This is achieved by extending recent ideas from learning of unsupervised image denoisers to unstructured 3D point clouds. Unsupervised image denoisers operate under the assumption that a noisy pixel observation is a random realization of a distribution around a clean pixel value, which allows appropriate learning on this distribution to eventually converge to the correct value. Regrettably, this assumption is not valid for unstructured points: 3D point clouds are subject to total noise, i. e., deviations in all coordinates, with no reliable pixel grid. Thus, an observation can be the realization of an entire manifold of clean 3D points, which makes a na\"ive extension of unsupervised image denoisers to 3D point clouds impractical. Overcoming this, we introduce a spatial prior term, that steers converges to the unique closest out of the many possible modes on a manifold. Our results demonstrate unsupervised denoising performance similar to that of supervised learning with clean data when given enough training examples - whereby we do not need any pairs of noisy and clean training data.
Attend To Count: Crowd Counting with Adaptive Capacity Multi-scale CNNs
Aug 26, 2019
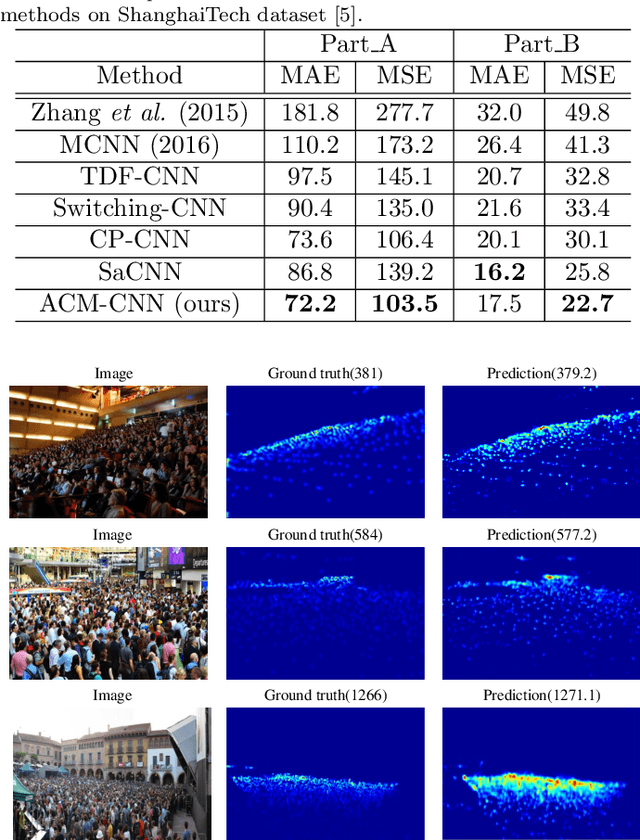
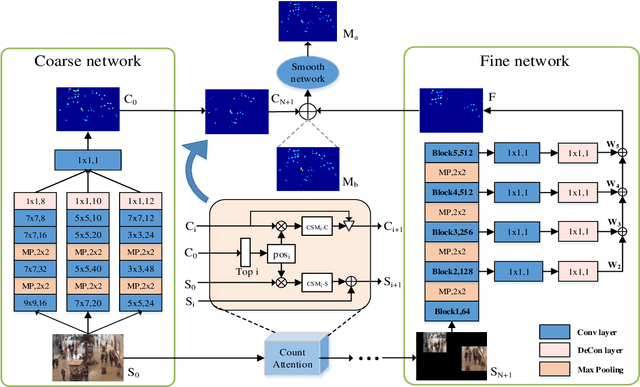
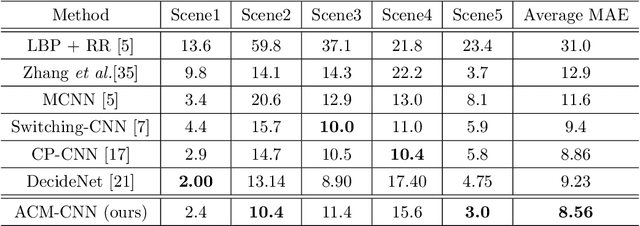
Crowd counting is a challenging task due to the large variations in crowd distributions. Previous methods tend to tackle the whole image with a single fixed structure, which is unable to handle diverse complicated scenes with different crowd densities. Hence, we propose the Adaptive Capacity Multi-scale convolutional neural networks (ACM-CNN), a novel crowd counting approach which can assign different capacities to different portions of the input. The intuition is that the model should focus on important regions of the input image and optimize its capacity allocation conditioning on the crowd intensive degree. ACM-CNN consists of three types of modules: a coarse network, a fine network, and a smooth network. The coarse network is used to explore the areas that need to be focused via count attention mechanism, and generate a rough feature map. Then the fine network processes the areas of interest into a fine feature map. To alleviate the sense of division caused by fusion, the smooth network is designed to combine two feature maps organically to produce high-quality density maps. Extensive experiments are conducted on five mainstream datasets. The results demonstrate the effectiveness of the proposed model for both density estimation and crowd counting tasks.
ATIS + SpiNNaker: a Fully Event-based Visual Tracking Demonstration
Dec 03, 2019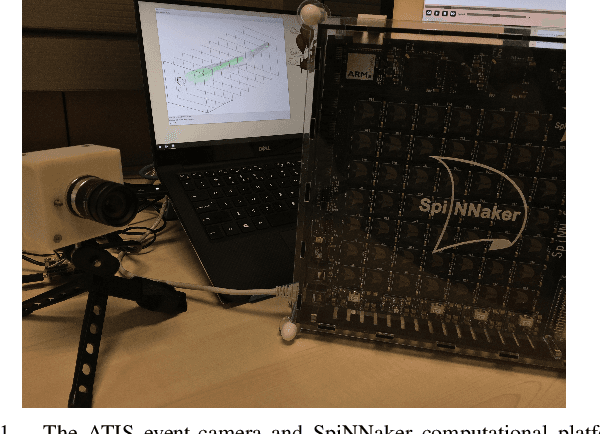
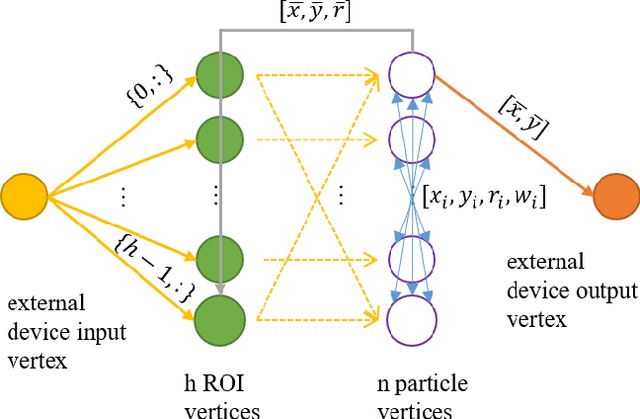
The Asynchronous Time-based Image Sensor (ATIS) and the Spiking Neural Network Architecture (SpiNNaker) are both neuromorphic technologies that "unconventionally" use binary spikes to represent information. The ATIS produces spikes to represent the change in light falling on the sensor, and the SpiNNaker is a massively parallel computing platform that asynchronously sends spikes between cores for processing. In this demonstration we show these two hardware used together to perform a visual tracking task. We aim to show the hardware and software architecture that integrates the ATIS and SpiNNaker together in a robot middle-ware that makes processing agnostic to the platform (CPU or SpiNNaker). We also aim to describe the algorithm, why it is suitable for the "unconventional" sensor and processing platform including the advantages as well as challenges faced.
Feature Fusion Encoder Decoder Network For Automatic Liver Lesion Segmentation
Mar 28, 2019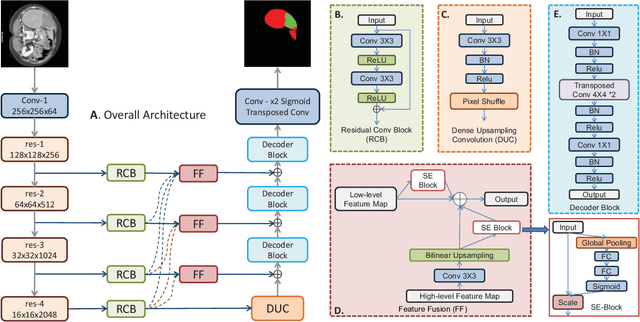
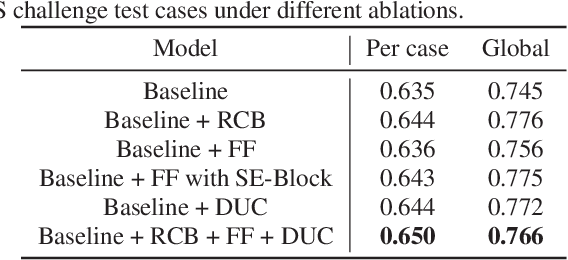
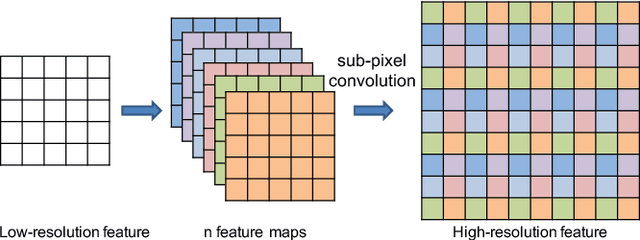
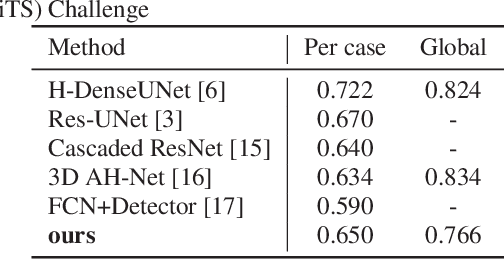
Liver lesion segmentation is a difficult yet critical task for medical image analysis. Recently, deep learning based image segmentation methods have achieved promising performance, which can be divided into three categories: 2D, 2.5D and 3D, based on the dimensionality of the models. However, 2.5D and 3D methods can have very high complexity and 2D methods may not perform satisfactorily. To obtain competitive performance with low complexity, in this paper, we propose a Feature-fusion Encoder-Decoder Network (FED-Net) based 2D segmentation model to tackle the challenging problem of liver lesion segmentation from CT images. Our feature fusion method is based on the attention mechanism, which fuses high-level features carrying semantic information with low-level features having image details. Additionally, to compensate for the information loss during the upsampling process, a dense upsampling convolution and a residual convolutional structure are proposed. We tested our method on the dataset of MICCAI 2017 Liver Tumor Segmentation (LiTS) Challenge and achieved competitive results compared with other state-of-the-art methods.
Test-Time Training for Out-of-Distribution Generalization
Sep 29, 2019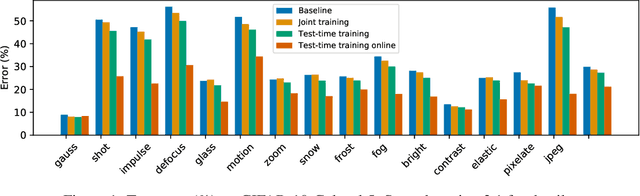
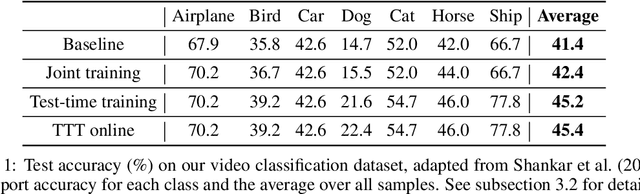
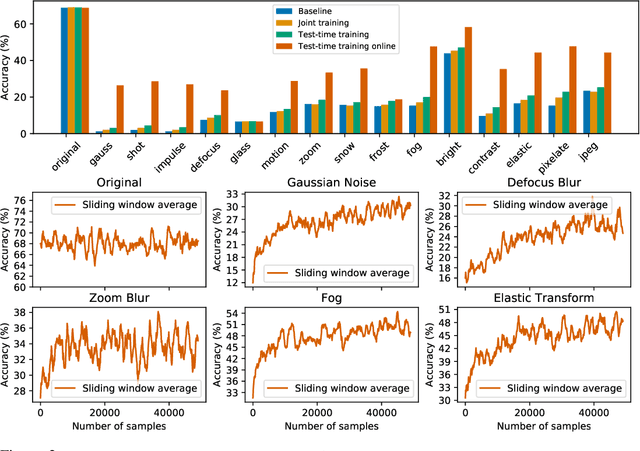
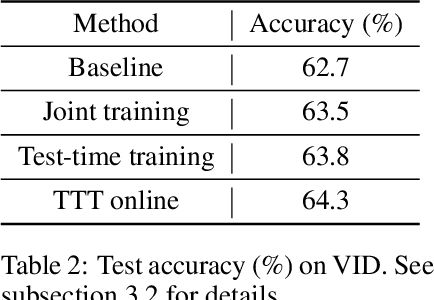
We introduce a general approach, called test-time training, for improving the performance of predictive models when test and training data come from different distributions. Test-time training turns a single unlabeled test instance into a self-supervised learning problem, on which we update the model parameters before making a prediction on the test sample. We show that this simple idea leads to surprising improvements on diverse image classification benchmarks aimed at evaluating robustness to distribution shifts. Theoretical investigations on a convex model reveal helpful intuitions for when we can expect our approach to help.
Sparse Deep Stacking Network for Image Classification
Jan 05, 2015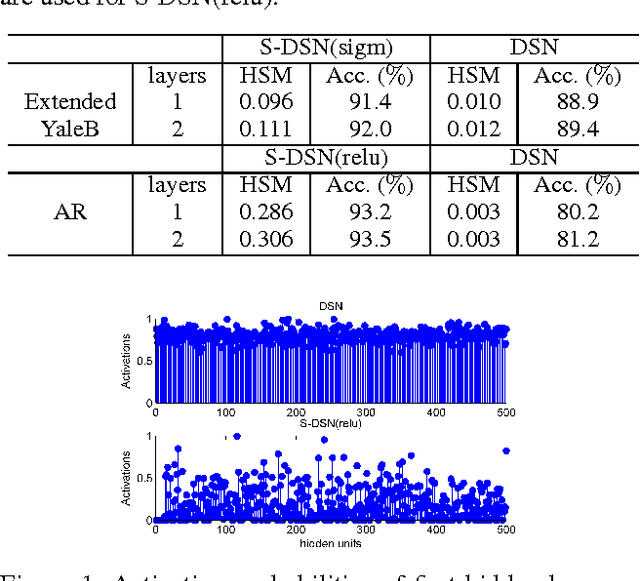
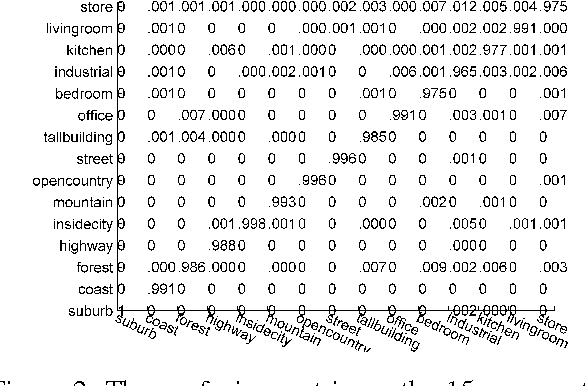
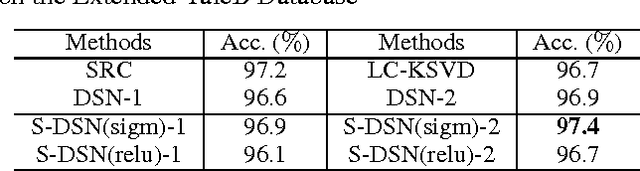

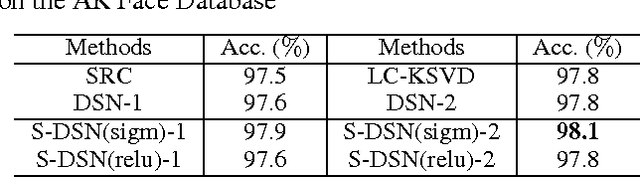
Sparse coding can learn good robust representation to noise and model more higher-order representation for image classification. However, the inference algorithm is computationally expensive even though the supervised signals are used to learn compact and discriminative dictionaries in sparse coding techniques. Luckily, a simplified neural network module (SNNM) has been proposed to directly learn the discriminative dictionaries for avoiding the expensive inference. But the SNNM module ignores the sparse representations. Therefore, we propose a sparse SNNM module by adding the mixed-norm regularization (l1/l2 norm). The sparse SNNM modules are further stacked to build a sparse deep stacking network (S-DSN). In the experiments, we evaluate S-DSN with four databases, including Extended YaleB, AR, 15 scene and Caltech101. Experimental results show that our model outperforms related classification methods with only a linear classifier. It is worth noting that we reach 98.8% recognition accuracy on 15 scene.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge