"Image": models, code, and papers
Augmented Semantic Signatures of Airborne LiDAR Point Clouds for Determining Change in Time-varying Data
Apr 29, 2020
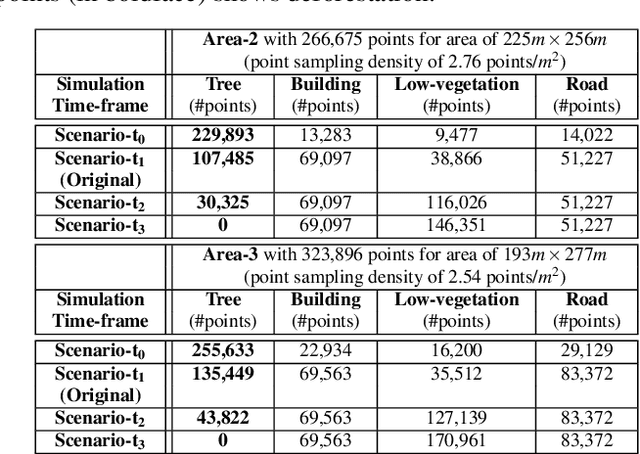
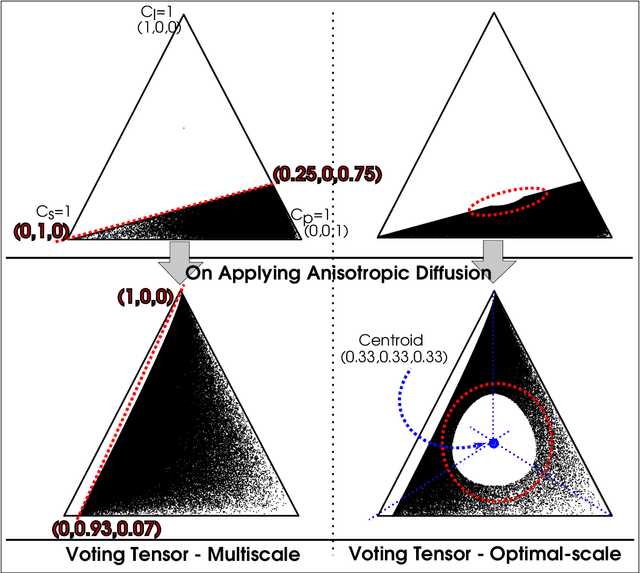
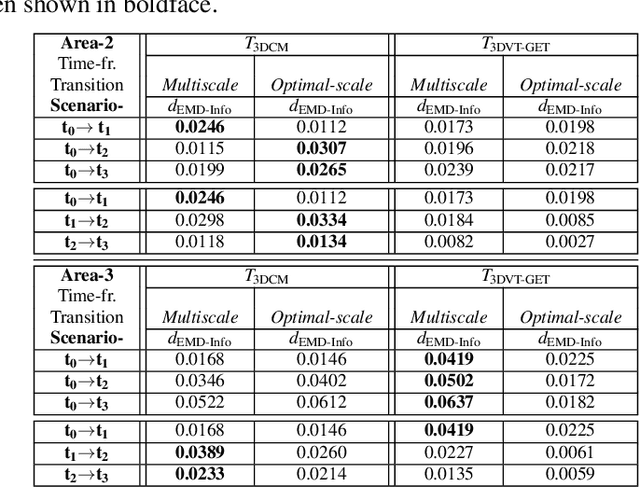
LiDAR point clouds provide rich geometric information, which is particularly useful for the analysis of complex scenes of urban regions. Finding structural and semantic differences between two different three-dimensional point clouds, say, of the same region but acquired at different time instances is an important problem. Usually, the data capture has inconsistencies when taken at different time instances, e.g., sampling densities, and the orientation of the flight path. Hence, change detection involves computationally expensive registration and segmentation. We are interested in capturing the relative differences in the geometric uncertainty and semantic content of the point cloud without the registration process. Hence, we propose an orientation-invariant geometric signature of the point cloud, which integrates its probabilistic geometric and semantic classifications. We study different properties of the geometric signature, which are image-based encoding of geometric uncertainty and semantic content. We explore different metrics to determine differences between these signatures, which in turn compare point clouds without performing point-to-point registration. We have observed that a point cloud with four semantic classes, namely, buildings, trees, road, and low-vegetation, that the tree class shows a characteristic pattern. Thus, we use a case study of airborne LiDAR point clouds where the visual and the quantitative comparisons of the geometric signatures of point clouds are useful in demonstrating changes during a thematic event, such as progressive deforestation, in the topography of an urban region. Our results show that the differences in the signatures corroborate with the geometric and semantic differences of the point clouds.
Local Search is a Remarkably Strong Baseline for Neural Architecture Search
Apr 29, 2020



Neural Architecture Search (NAS), i.e., the automation of neural network design, has gained much popularity in recent years with increasingly complex search algorithms being proposed. Yet, solid comparisons with simple baselines are often missing. At the same time, recent retrospective studies have found many new algorithms to be no better than random search (RS). In this work we consider, for the first time, a simple Local Search (LS) algorithm for NAS. We particularly consider a multi-objective NAS formulation, with network accuracy and network complexity as two objectives, as understanding the trade-off between these two objectives is arguably the most interesting aspect of NAS. The proposed LS algorithm is compared with RS and two evolutionary algorithms (EAs), as these are often heralded as being ideal for multi-objective optimization. To promote reproducibility, we create and release two benchmark datasets containing 200K saved network evaluations for two established image classification tasks, CIFAR-10 and CIFAR-100. Our benchmarks are designed to be complementary to existing benchmarks, especially in that they are better suited for multi-objective search. We additionally consider a version of the problem with a much larger architecture space. While we find and show that the considered algorithms explore the search space in fundamentally different ways, we also find that LS substantially outperforms RS and even performs nearly as good as state-of-the-art EAs. We believe that this provides strong evidence that LS is truly a competitive baseline for NAS against which new NAS algorithms should be benchmarked.
Radioactive data: tracing through training
Feb 03, 2020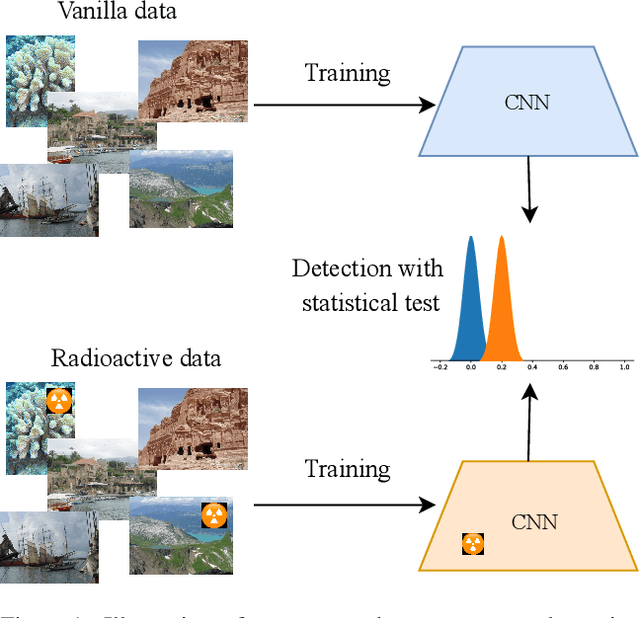
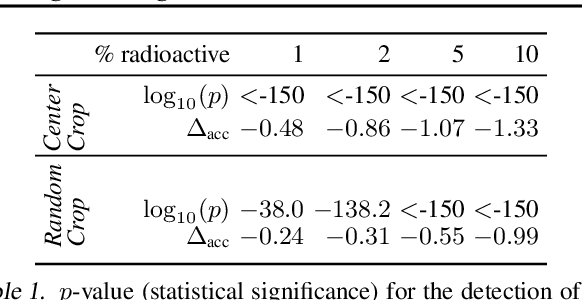
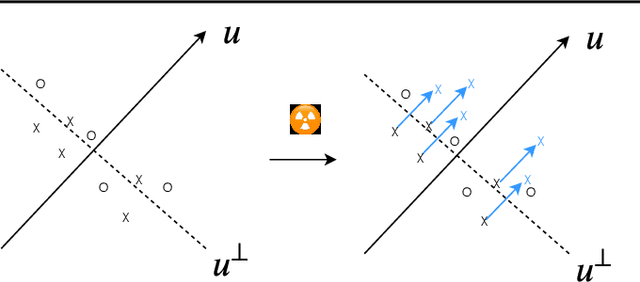
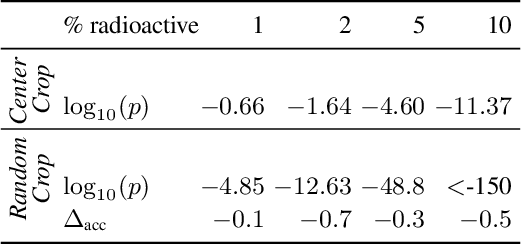
We want to detect whether a particular image dataset has been used to train a model. We propose a new technique, \emph{radioactive data}, that makes imperceptible changes to this dataset such that any model trained on it will bear an identifiable mark. The mark is robust to strong variations such as different architectures or optimization methods. Given a trained model, our technique detects the use of radioactive data and provides a level of confidence (p-value). Our experiments on large-scale benchmarks (Imagenet), using standard architectures (Resnet-18, VGG-16, Densenet-121) and training procedures, show that we can detect usage of radioactive data with high confidence (p<10^-4) even when only 1% of the data used to trained our model is radioactive. Our method is robust to data augmentation and the stochasticity of deep network optimization. As a result, it offers a much higher signal-to-noise ratio than data poisoning and backdoor methods.
Multi-scale GANs for Memory-efficient Generation of High Resolution Medical Images
Jul 02, 2019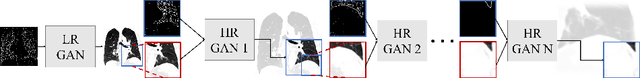
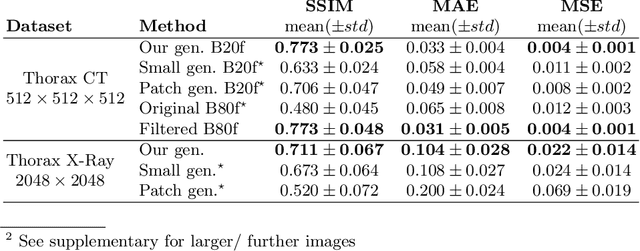
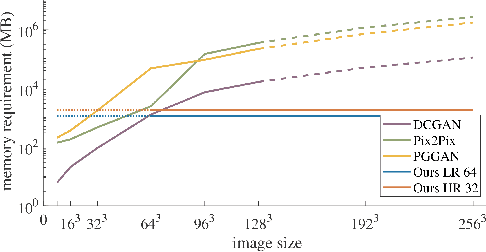
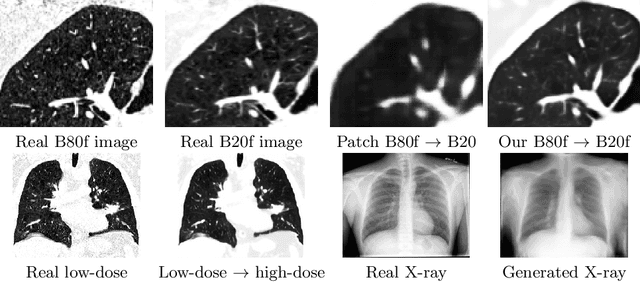
Currently generative adversarial networks (GANs) are rarely applied to medical images of large sizes, especially 3D volumes, due to their large computational demand. We propose a novel multi-scale patch-based GAN approach to generate large high resolution 2D and 3D images. Our key idea is to first learn a low-resolution version of the image and then generate patches of successively growing resolutions conditioned on previous scales. In a domain translation use-case scenario, 3D thorax CTs of size 512x512x512 and thorax X-rays of size 2048x2048 are generated and we show that, due to the constant GPU memory demand of our method, arbitrarily large images of high resolution can be generated. Moreover, compared to common patch-based approaches, our multi-resolution scheme enables better image quality and prevents patch artifacts.
Cross-Modal Relationship Inference for Grounding Referring Expressions
Jun 11, 2019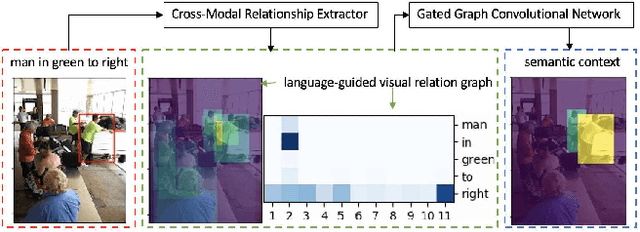
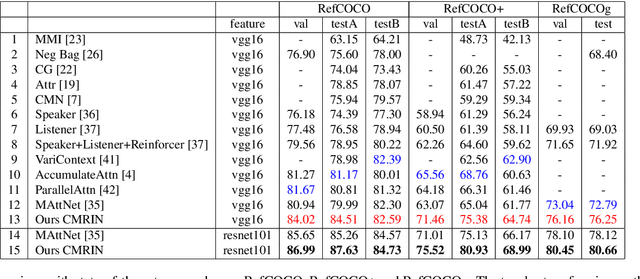
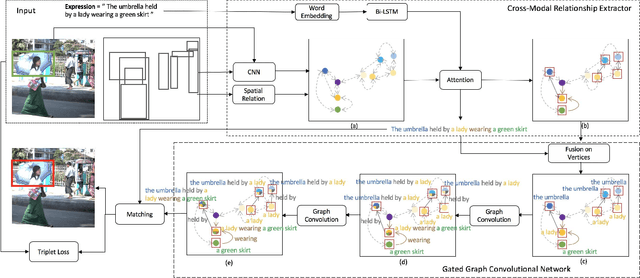
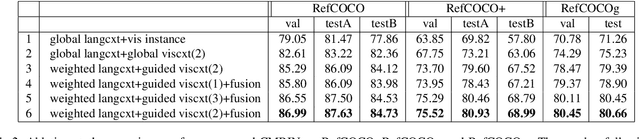
Grounding referring expressions is a fundamental yet challenging task facilitating human-machine communication in the physical world. It locates the target object in an image on the basis of the comprehension of the relationships between referring natural language expressions and the image. A feasible solution for grounding referring expressions not only needs to extract all the necessary information (i.e., objects and the relationships among them) in both the image and referring expressions, but also compute and represent multimodal contexts from the extracted information. Unfortunately, existing work on grounding referring expressions cannot extract multi-order relationships from the referring expressions accurately and the contexts they obtain have discrepancies with the contexts described by referring expressions. In this paper, we propose a Cross-Modal Relationship Extractor (CMRE) to adaptively highlight objects and relationships, that have connections with a given expression, with a cross-modal attention mechanism, and represent the extracted information as a language-guided visual relation graph. In addition, we propose a Gated Graph Convolutional Network (GGCN) to compute multimodal semantic contexts by fusing information from different modes and propagating multimodal information in the structured relation graph. Experiments on various common benchmark datasets show that our Cross-Modal Relationship Inference Network, which consists of CMRE and GGCN, outperforms all existing state-of-the-art methods.
One-Shot Informed Robotic Visual Search in the Wild
Mar 22, 2020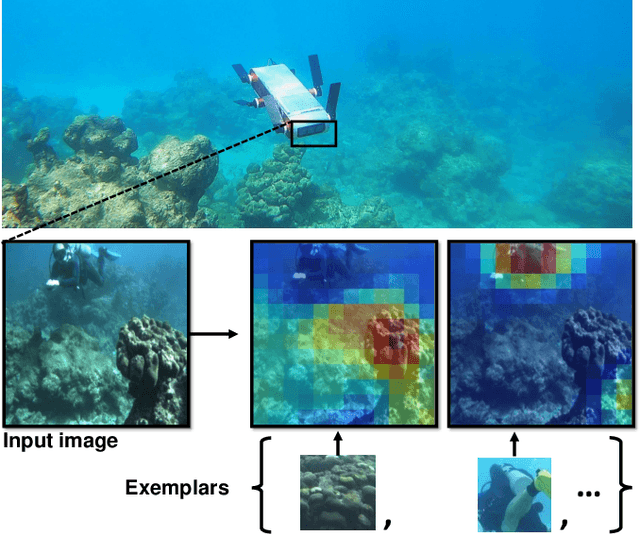
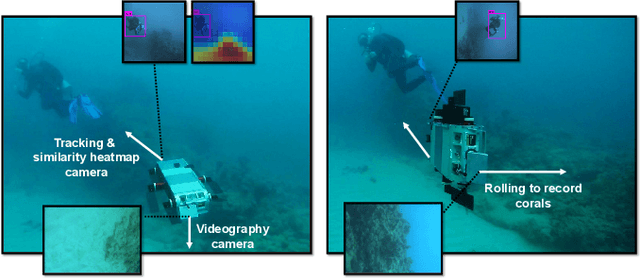
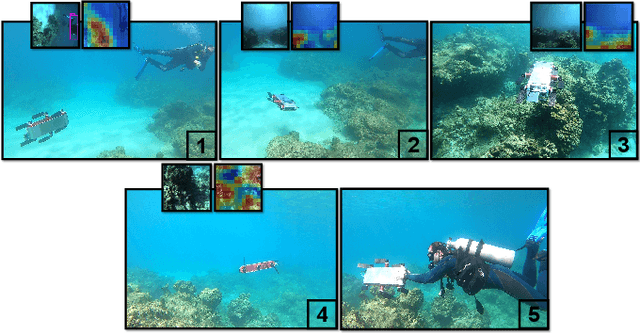
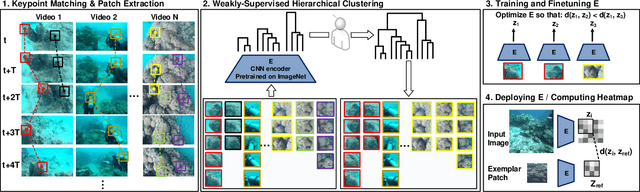


We consider the task of underwater robot navigation for the purpose of collecting scientifically-relevant video data for environmental monitoring. The majority of field robots that currently perform monitoring tasks in unstructured natural environments navigate via path-tracking a pre-specified sequence of waypoints. Although this navigation method is often necessary, it is limiting because the robot does not have a model of what the scientist deems to be relevant visual observations. Thus, the robot can neither visually search for particular types of objects, nor focus its attention on parts of the scene that might be more relevant than the pre-specified waypoints and viewpoints. In this paper we propose a method that enables informed visual navigation via a learned visual similarity operator that guides the robot's visual search towards parts of the scene that look like an exemplar image, which is given by the user as a high-level specification for data collection. We propose and evaluate a weakly-supervised video representation learning method that outperforms ImageNet embeddings for similarity tasks in the underwater domain. We also demonstrate the deployment of this similarity operator during informed visual navigation in collaborative environmental monitoring scenarios, in large-scale field trials, where the robot and a human scientist jointly search for relevant visual content.
Linear Spatial Pyramid Matching Using Non-convex and non-negative Sparse Coding for Image Classification
Apr 27, 2015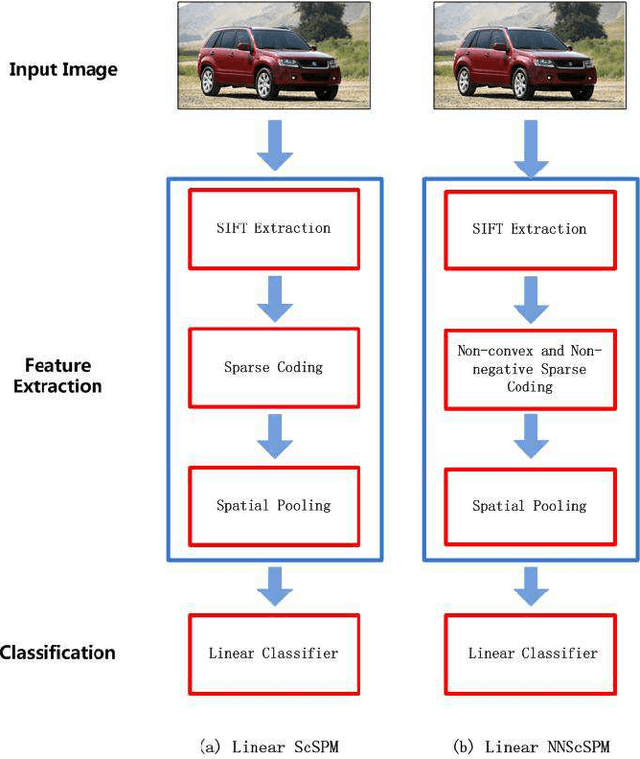
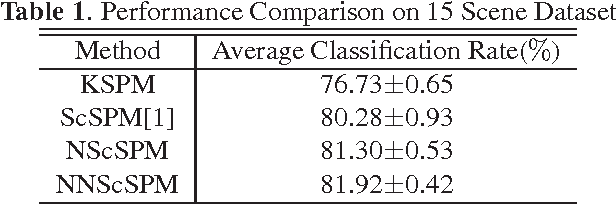
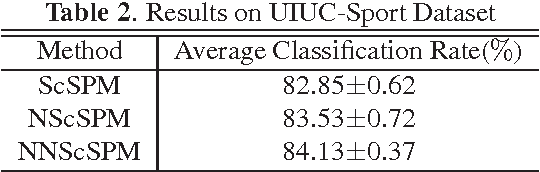
Recently sparse coding have been highly successful in image classification mainly due to its capability of incorporating the sparsity of image representation. In this paper, we propose an improved sparse coding model based on linear spatial pyramid matching(SPM) and Scale Invariant Feature Transform (SIFT ) descriptors. The novelty is the simultaneous non-convex and non-negative characters added to the sparse coding model. Our numerical experiments show that the improved approach using non-convex and non-negative sparse coding is superior than the original ScSPM[1] on several typical databases.
End-to-end Segmentation with Recurrent Attention Neural Network
Dec 05, 2018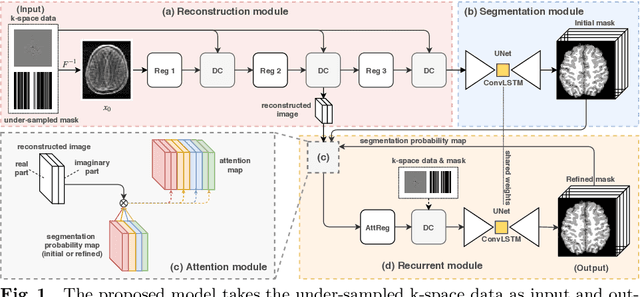
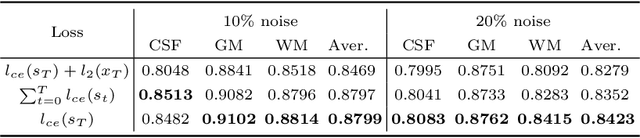
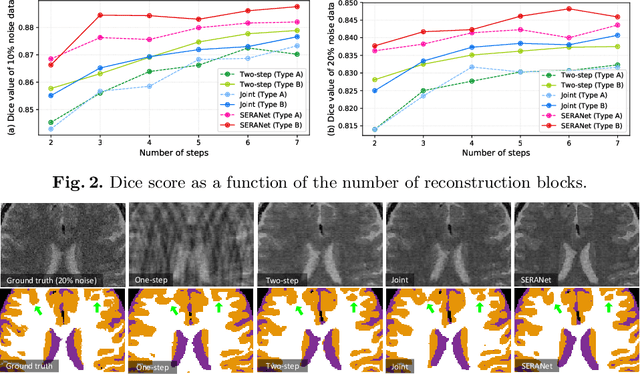
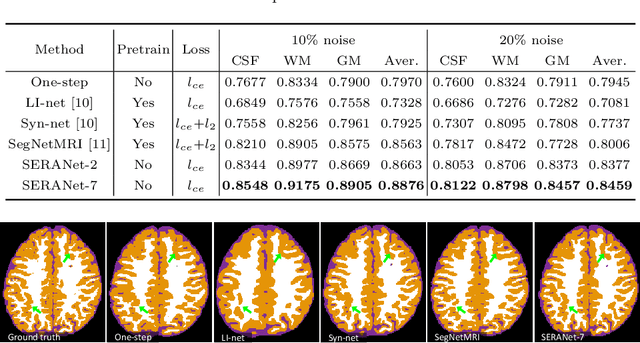
Image segmentation quality depends heavily on the quality of the image. For many medical imaging modalities, image reconstruction is required to convert acquired raw data to images before any analysis. However, imperfect reconstruction with artifacts and loss of information is almost inevitable, which compromises the final performance of segmentation. In this study, we present a novel end-to-end deep learning framework that performs magnetic resonance brain image segmentation directly from the raw data. The end-to-end framework consists a unique task-driven attention module that recurrently utilizes intermediate segmentation result to facilitate image-domain feature extraction from the raw data for segmentation, thus closely bridging the reconstruction and the segmentation tasks. In addition, we introduce a novel workflow to generate labeled training data for segmentation by exploiting imaging modality simulators and digital phantoms. Extensive experiment results show that the proposed method outperforms the state-of-the-art methods.
Neither Global Nor Local: A Hierarchical Robust Subspace Clustering For Image Data
May 17, 2019
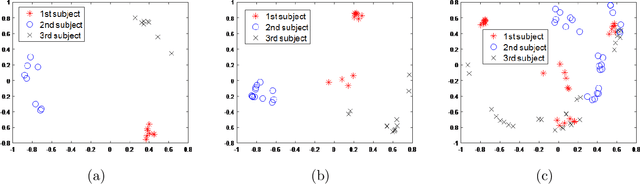
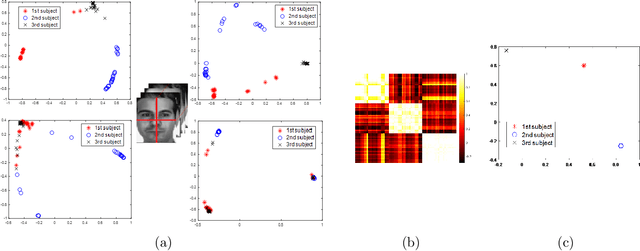
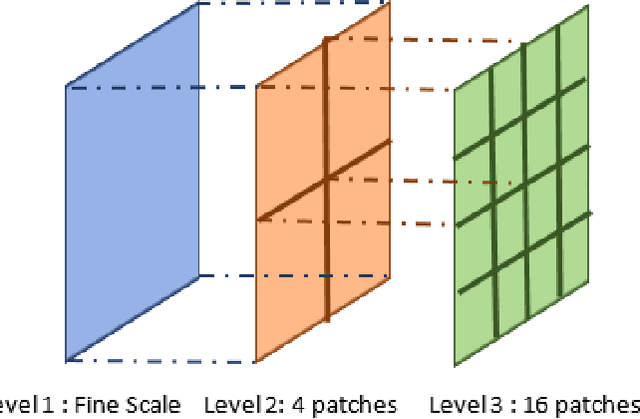
In this paper, we consider the problem of subspace clustering in presence of contiguous noise, occlusion and disguise. We argue that self-expressive representation of data in current state-of-the-art approaches is severely sensitive to occlusions and complex real-world noises. To alleviate this problem, we propose a hierarchical framework that brings robustness of local patches-based representations and discriminant property of global representations together. This approach consists of 1) a top-down stage, in which the input data is subject to repeated division to smaller patches and 2) a bottom-up stage, in which the low rank embedding of local patches in field of view of a corresponding patch in upper level are merged on a Grassmann manifold. This summarized information provides two key information for the corresponding patch on the upper level: cannot-links and recommended-links. This information is employed for computing a self-expressive representation of each patch at upper levels using a weighted sparse group lasso optimization problem. Numerical results on several real data sets confirm the efficiency of our approach.
Leveraging Big Data Analytics in Healthcare Enhancement: Trends, Challenges and Opportunities
Apr 05, 2020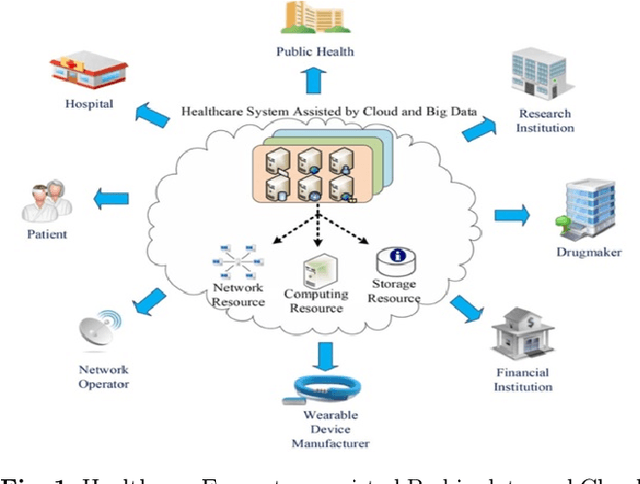
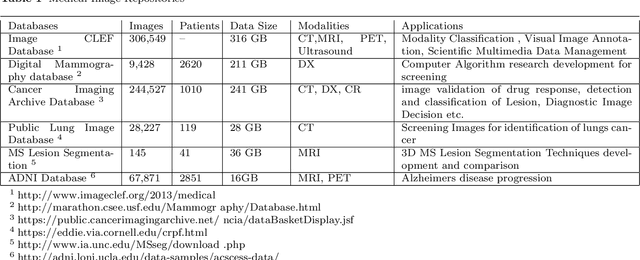
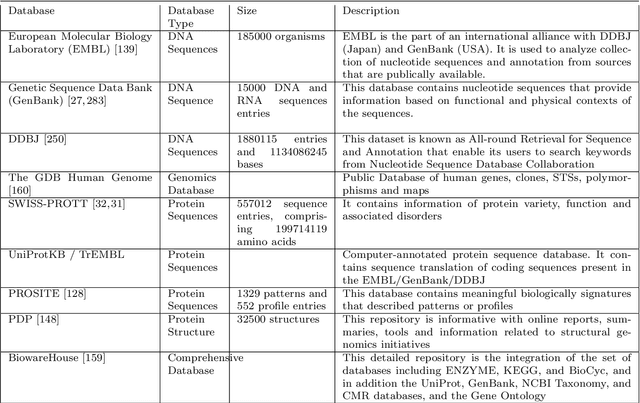
Clinicians decisions are becoming more and more evidence-based meaning in no other field the big data analytics so promising as in healthcare. Due to the sheer size and availability of healthcare data, big data analytics has revolutionized this industry and promises us a world of opportunities. It promises us the power of early detection, prediction, prevention and helps us to improve the quality of life. Researchers and clinicians are working to inhibit big data from having a positive impact on health in the future. Different tools and techniques are being used to analyze, process, accumulate, assimilate and manage large amount of healthcare data either in structured or unstructured form. In this paper, we would like to address the need of big data analytics in healthcare: why and how can it help to improve life?. We present the emerging landscape of big data and analytical techniques in the five sub-disciplines of healthcare i.e.medical image analysis and imaging informatics, bioinformatics, clinical informatics, public health informatics and medical signal analytics. We presents different architectures, advantages and repositories of each discipline that draws an integrated depiction of how distinct healthcare activities are accomplished in the pipeline to facilitate individual patients from multiple perspectives. Finally the paper ends with the notable applications and challenges in adoption of big data analytics in healthcare.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge