"Image": models, code, and papers
PAINTER: a spatio-spectral image reconstruction algorithm for optical interferometry
Sep 27, 2014
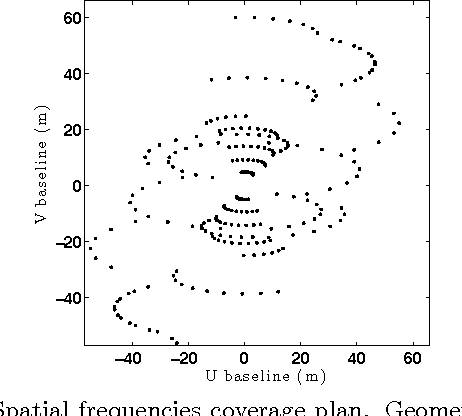
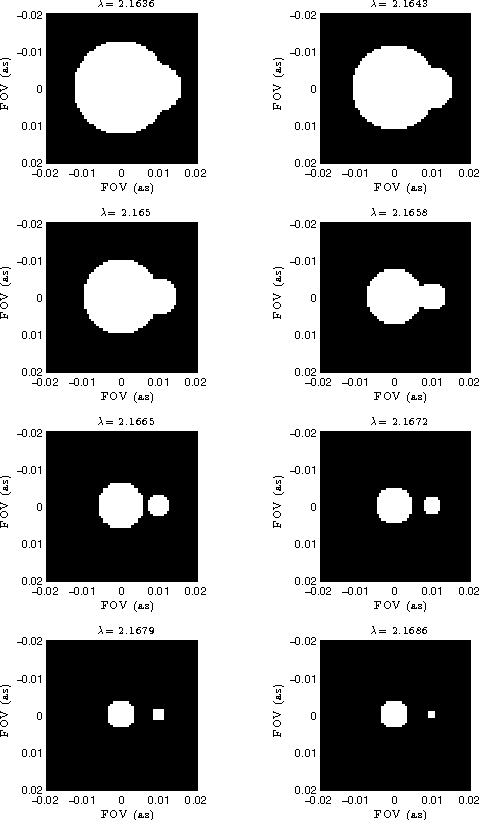
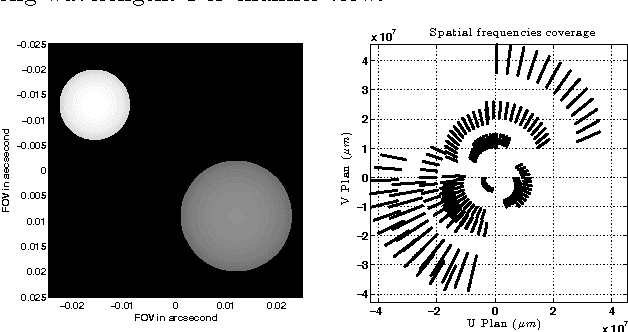
Astronomical optical interferometers sample the Fourier transform of the intensity distribution of a source at the observation wavelength. Because of rapid perturbations caused by atmospheric turbulence, the phases of the complex Fourier samples (visibilities) cannot be directly exploited. Consequently, specific image reconstruction methods have been devised in the last few decades. Modern polychromatic optical interferometric instruments are now paving the way to multiwavelength imaging. This paper is devoted to the derivation of a spatio-spectral (3D) image reconstruction algorithm, coined PAINTER (Polychromatic opticAl INTErferometric Reconstruction software). The algorithm relies on an iterative process, which alternates estimation of polychromatic images and of complex visibilities. The complex visibilities are not only estimated from squared moduli and closure phases, but also differential phases, which helps to better constrain the polychromatic reconstruction. Simulations on synthetic data illustrate the efficiency of the algorithm and in particular the relevance of injecting a differential phases model in the reconstruction.
Conditionally Deep Hybrid Neural Networks Across Edge and Cloud
May 21, 2020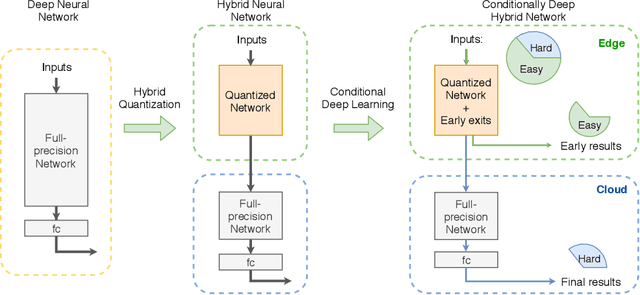
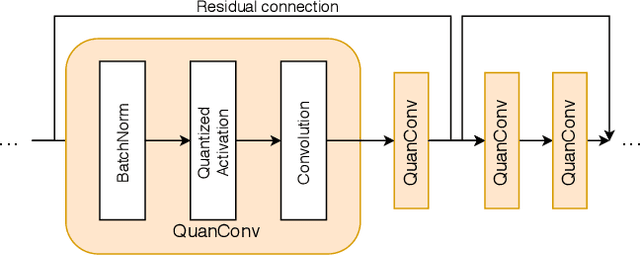
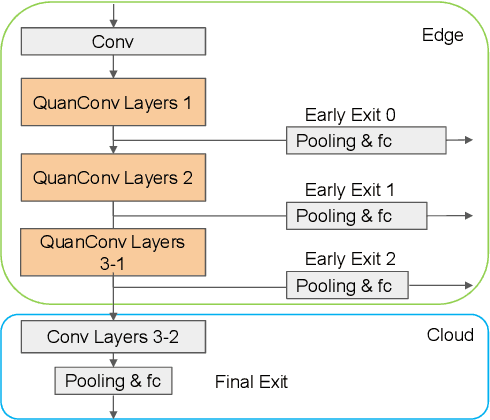
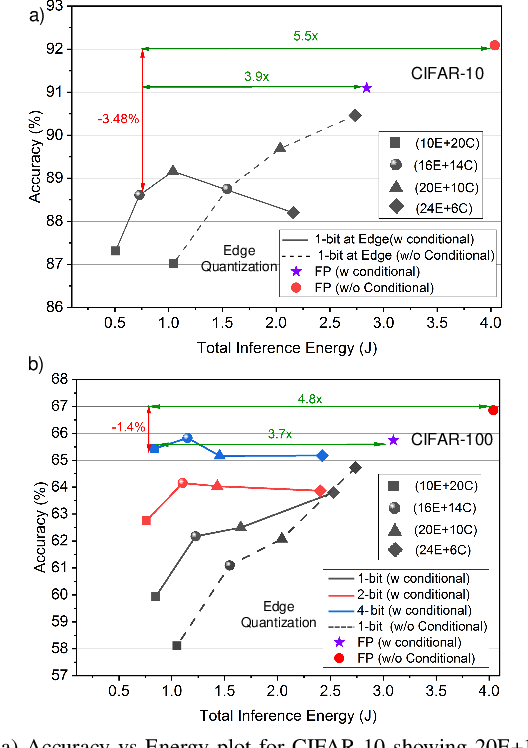
The pervasiveness of "Internet-of-Things" in our daily life has led to a recent surge in fog computing, encompassing a collaboration of cloud computing and edge intelligence. To that effect, deep learning has been a major driving force towards enabling such intelligent systems. However, growing model sizes in deep learning pose a significant challenge towards deployment in resource-constrained edge devices. Moreover, in a distributed intelligence environment, efficient workload distribution is necessary between edge and cloud systems. To address these challenges, we propose a conditionally deep hybrid neural network for enabling AI-based fog computing. The proposed network can be deployed in a distributed manner, consisting of quantized layers and early exits at the edge and full-precision layers on the cloud. During inference, if an early exit has high confidence in the classification results, it would allow samples to exit at the edge, and the deeper layers on the cloud are activated conditionally, which can lead to improved energy efficiency and inference latency. We perform an extensive design space exploration with the goal of minimizing energy consumption at the edge while achieving state-of-the-art classification accuracies on image classification tasks. We show that with binarized layers at the edge, the proposed conditional hybrid network can process 65% of inferences at the edge, leading to 5.5x computational energy reduction with minimal accuracy degradation on CIFAR-10 dataset. For the more complex dataset CIFAR-100, we observe that the proposed network with 4-bit quantization at the edge achieves 52% early classification at the edge with 4.8x energy reduction. The analysis gives us insights on designing efficient hybrid networks which achieve significantly higher energy efficiency than full-precision networks for edge-cloud based distributed intelligence systems.
HeatNet: Bridging the Day-Night Domain Gap in Semantic Segmentation with Thermal Images
Mar 10, 2020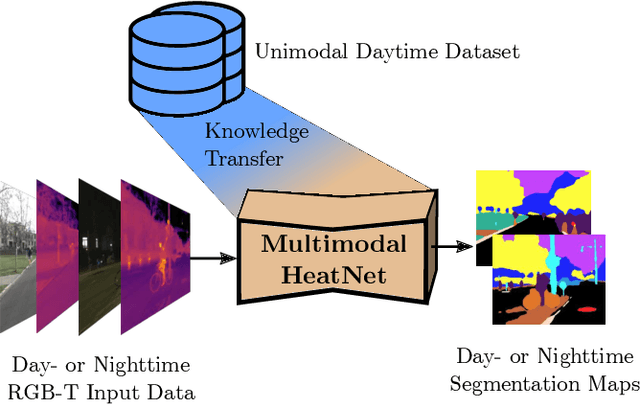
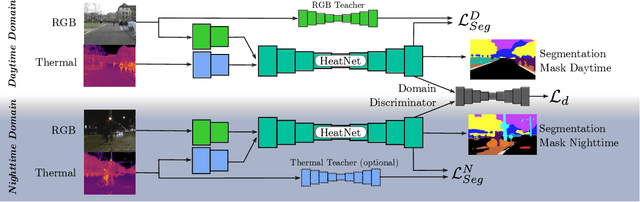


The majority of learning-based semantic segmentation methods are optimized for daytime scenarios and favorable lighting conditions. Real-world driving scenarios, however, entail adverse environmental conditions such as nighttime illumination or glare which remain a challenge for existing approaches. In this work, we propose a multimodal semantic segmentation model that can be applied during daytime and nighttime. To this end, besides RGB images, we leverage thermal images, making our network significantly more robust. We avoid the expensive annotation of nighttime images by leveraging an existing daytime RGB-dataset and propose a teacher-student training approach that transfers the dataset's knowledge to the nighttime domain. We further employ a domain adaptation method to align the learned feature spaces across the domains and propose a novel two-stage training scheme. Furthermore, due to a lack of thermal data for autonomous driving, we present a new dataset comprising over 20,000 time-synchronized and aligned RGB-thermal image pairs. In this context, we also present a novel target-less calibration method that allows for automatic robust extrinsic and intrinsic thermal camera calibration. Among others, we employ our new dataset to show state-of-the-art results for nighttime semantic segmentation.
TripletUNet: Multi-Task U-Net with Online Voxel-Wise Learning for Precise CT Prostate Segmentation
May 21, 2020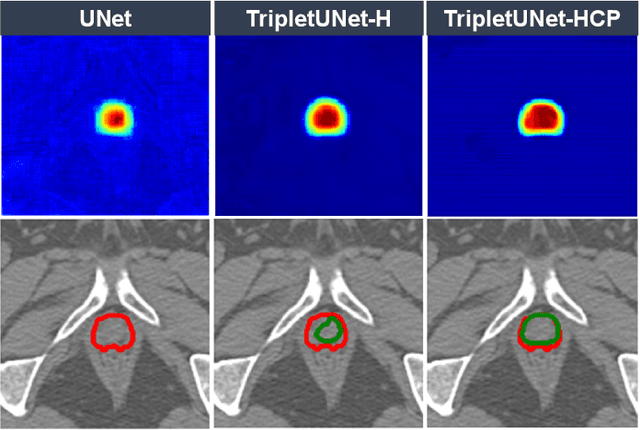
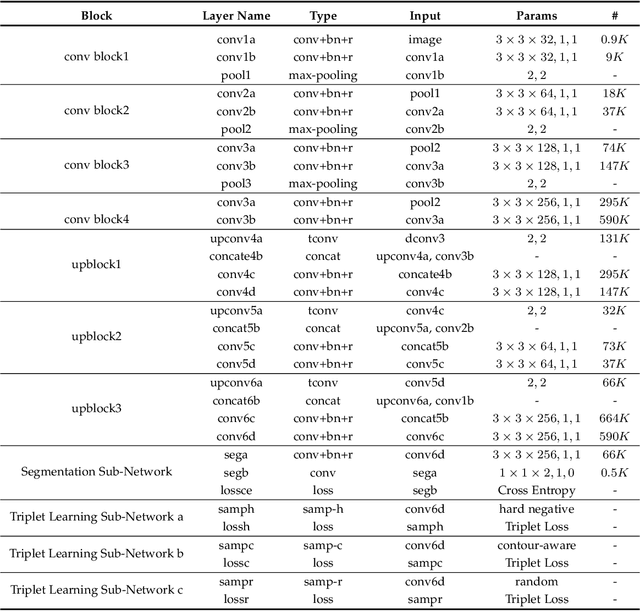
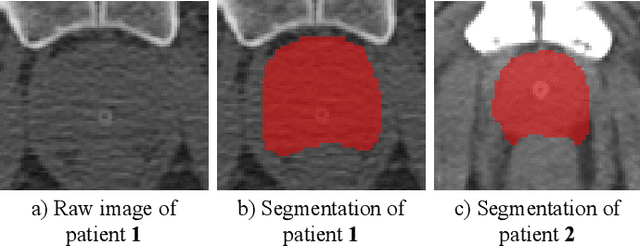
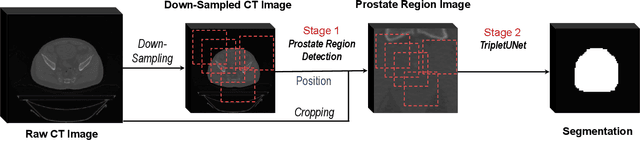
Fully convolutional networks (FCNs), including U-Net and V-Net, are widely-used network architecture for semantic segmentation in recent studies. However, conventional FCNs are typically trained by the cross-entropy loss or dice loss, in which the relationships among voxels are neglected. This often results in non-smooth neighborhoods in the output segmentation map. This problem becomes more serious in CT prostate segmentation as CT images are usually of low tissue contrast. To address this problem, we propose a two-stage framework. The first stage quickly localizes the prostate region. Then, the second stage precisely segments the prostate by a multi-task FCN-based on the U-Net architecture. We introduce a novel online voxel-triplet learning module through metric learning and voxel feature embeddings in the multi-task network. The proposed network has two branches guided by two tasks: 1) a segmentation sub-network aiming to generate prostate segmentation, and 2) a triplet learning sub-network aiming to improve the quality of the learned feature space supervised by a mixed of triplet and pair-wise loss function. The triplet learning sub-network samples triplets from the inter-mediate heatmap. Unlike conventional deep triplet learning methods that generate triplets before the training phase, our proposed voxel-triplets are sampled in an online manner and operates in an end-to-end fashion via multi-task learning. To evaluate the proposed method, we implement comprehensive experiments on a CT image dataset consisting of 339 patients. The ablation studies show that our method can effectively learn more representative voxel-level features compared with the conventional FCN network. And the comparisons show that the proposed method outperforms the state-of-the-art methods by a large margin.
Efficient and Accurate Gaussian Image Filtering Using Running Sums
Jul 25, 2011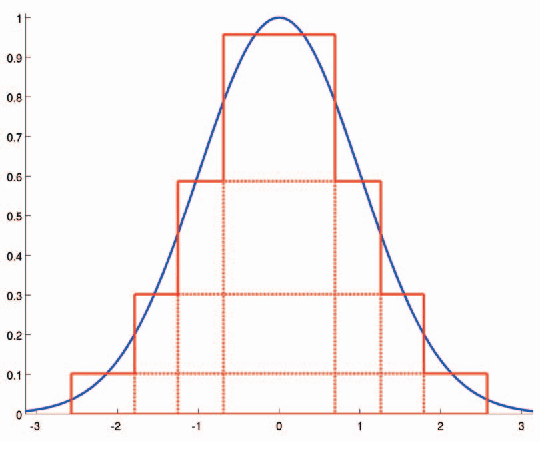
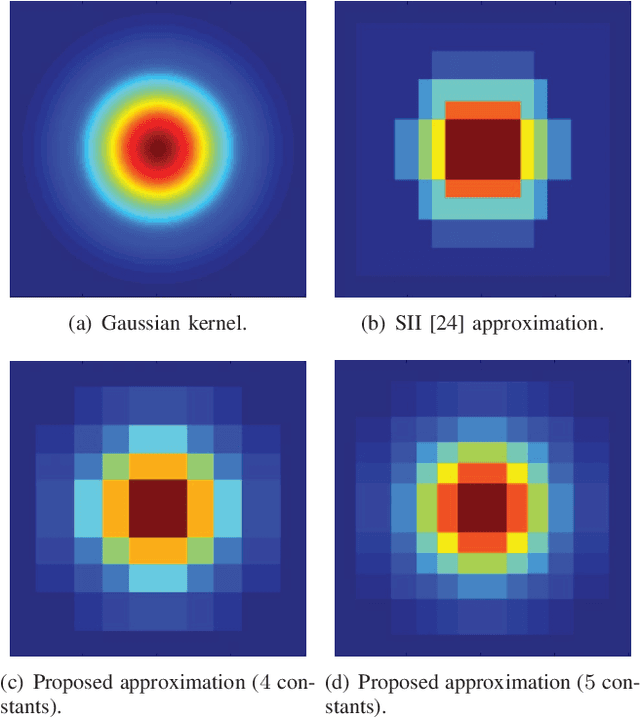
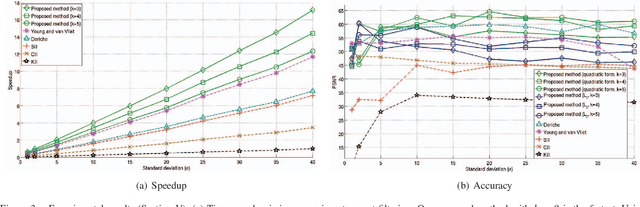
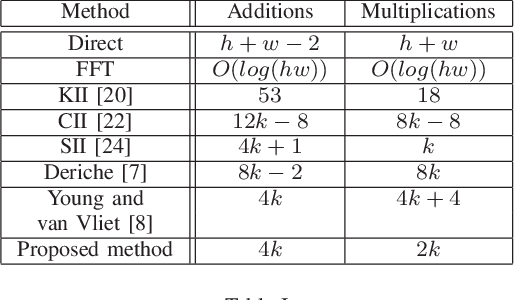
This paper presents a simple and efficient method to convolve an image with a Gaussian kernel. The computation is performed in a constant number of operations per pixel using running sums along the image rows and columns. We investigate the error function used for kernel approximation and its relation to the properties of the input signal. Based on natural image statistics we propose a quadratic form kernel error function so that the output image l2 error is minimized. We apply the proposed approach to approximate the Gaussian kernel by linear combination of constant functions. This results in very efficient Gaussian filtering method. Our experiments show that the proposed technique is faster than state of the art methods while preserving a similar accuracy.
A Biologically Interpretable Two-stage Deep Neural Network (BIT-DNN) For Hyperspectral Imagery Classification
Apr 19, 2020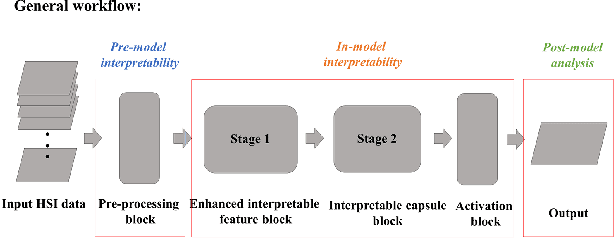
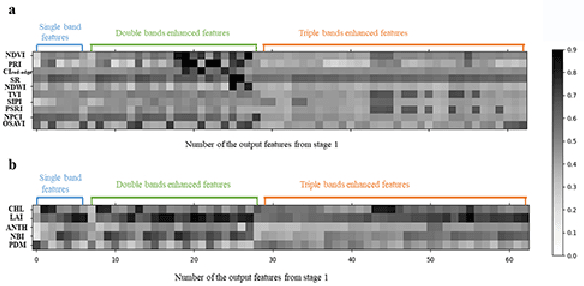
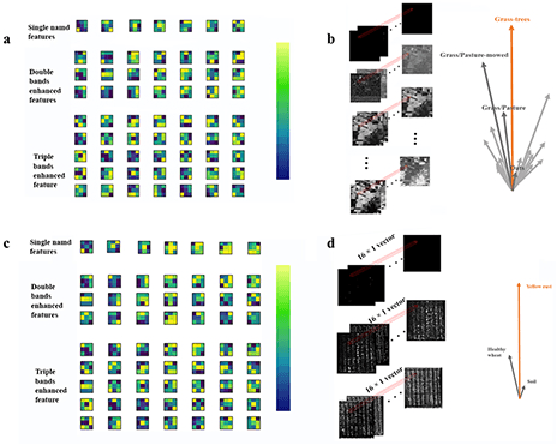
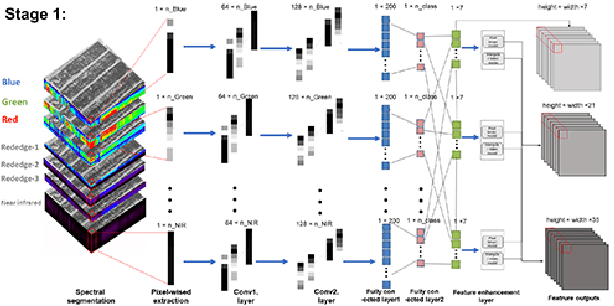
Spectral-spatial based deep learning models have recently proven to be effective in hyperspectral image (HSI) classification for various earth monitoring applications such as land cover classification and agricultural monitoring. However, due to the nature of "black-box" model representation, how to explain and interpret the learning process and the model decision remains an open problem. This study proposes an interpretable deep learning model -- a biologically interpretable two-stage deep neural network (BIT-DNN), by integrating biochemical and biophysical associated information into the proposed framework, capable of achieving both high accuracy and interpretability on HSI based classification tasks. The proposed model introduces a two-stage feature learning process. In the first stage, an enhanced interpretable feature block extracts low-level spectral features associated with the biophysical and biochemical attributes of the target entities; and in the second stage, an interpretable capsule block extracts and encapsulates the high-level joint spectral-spatial features into the featured tensors representing the hierarchical structure of the biophysical and biochemical attributes of the target ground entities, which provides the model an improved performance on classification and intrinsic interpretability. We have tested and evaluated the model using two real HSI datasets for crop type recognition and crop disease recognition tasks and compared it with six state-of-the-art machine learning models. The results demonstrate that the proposed model has competitive advantages in terms of both classification accuracy and model interpretability.
An analysis on the use of autoencoders for representation learning: fundamentals, learning task case studies, explainability and challenges
May 21, 2020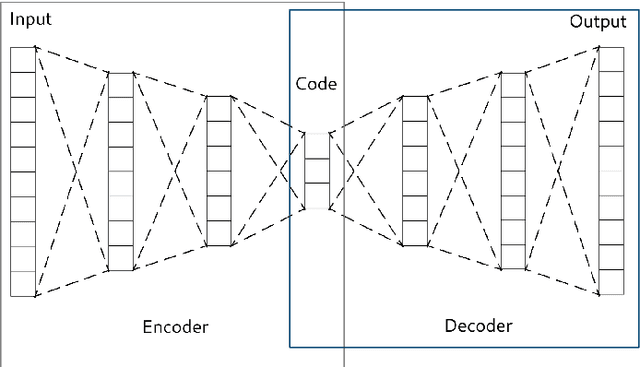
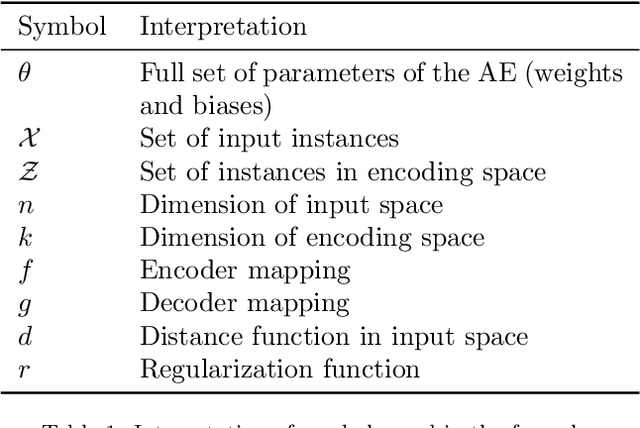
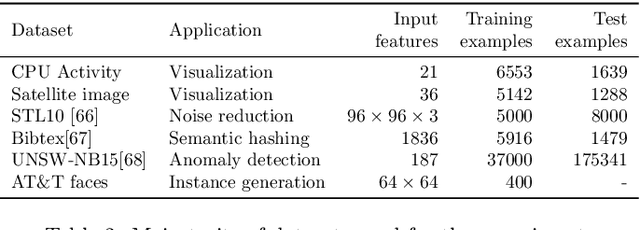
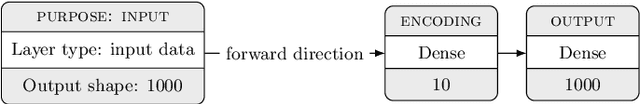
In many machine learning tasks, learning a good representation of the data can be the key to building a well-performant solution. This is because most learning algorithms operate with the features in order to find models for the data. For instance, classification performance can improve if the data is mapped to a space where classes are easily separated, and regression can be facilitated by finding a manifold of data in the feature space. As a general rule, features are transformed by means of statistical methods such as principal component analysis, or manifold learning techniques such as Isomap or locally linear embedding. From a plethora of representation learning methods, one of the most versatile tools is the autoencoder. In this paper we aim to demonstrate how to influence its learned representations to achieve the desired learning behavior. To this end, we present a series of learning tasks: data embedding for visualization, image denoising, semantic hashing, detection of abnormal behaviors and instance generation. We model them from the representation learning perspective, following the state of the art methodologies in each field. A solution is proposed for each task employing autoencoders as the only learning method. The theoretical developments are put into practice using a selection of datasets for the different problems and implementing each solution, followed by a discussion of the results in each case study and a brief explanation of other six learning applications. We also explore the current challenges and approaches to explainability in the context of autoencoders. All of this helps conclude that, thanks to alterations in their structure as well as their objective function, autoencoders may be the core of a possible solution to many problems which can be modeled as a transformation of the feature space.
Learning to be Global Optimizer
Mar 10, 2020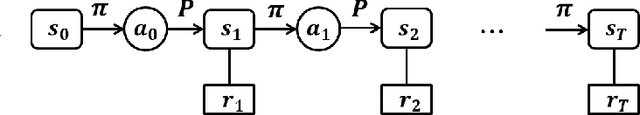
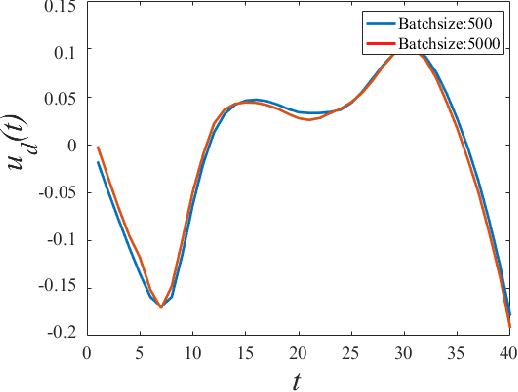
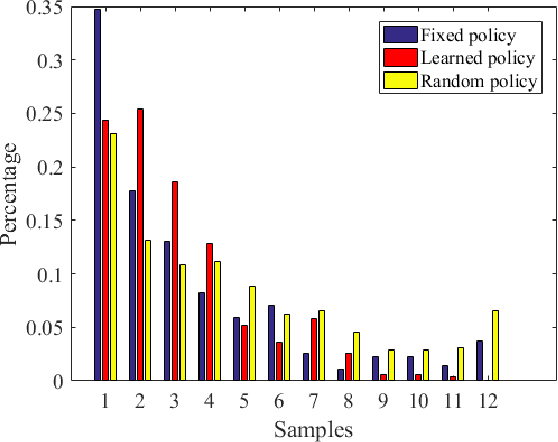
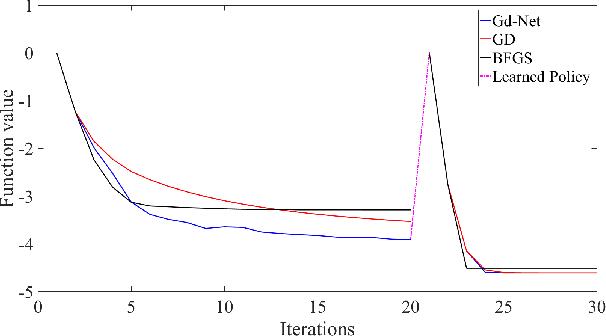
The advancement of artificial intelligence has cast a new light on the development of optimization algorithm. This paper proposes to learn a two-phase (including a minimization phase and an escaping phase) global optimization algorithm for smooth non-convex functions. For the minimization phase, a model-driven deep learning method is developed to learn the update rule of descent direction, which is formalized as a nonlinear combination of historical information, for convex functions. We prove that the resultant algorithm with the proposed adaptive direction guarantees convergence for convex functions. Empirical study shows that the learned algorithm significantly outperforms some well-known classical optimization algorithms, such as gradient descent, conjugate descent and BFGS, and performs well on ill-posed functions. The escaping phase from local optimum is modeled as a Markov decision process with a fixed escaping policy. We further propose to learn an optimal escaping policy by reinforcement learning. The effectiveness of the escaping policies is verified by optimizing synthesized functions and training a deep neural network for CIFAR image classification. The learned two-phase global optimization algorithm demonstrates a promising global search capability on some benchmark functions and machine learning tasks.
Fuzzy-based Propagation of Prior Knowledge to Improve Large-Scale Image Analysis Pipelines
Aug 03, 2016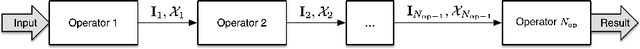
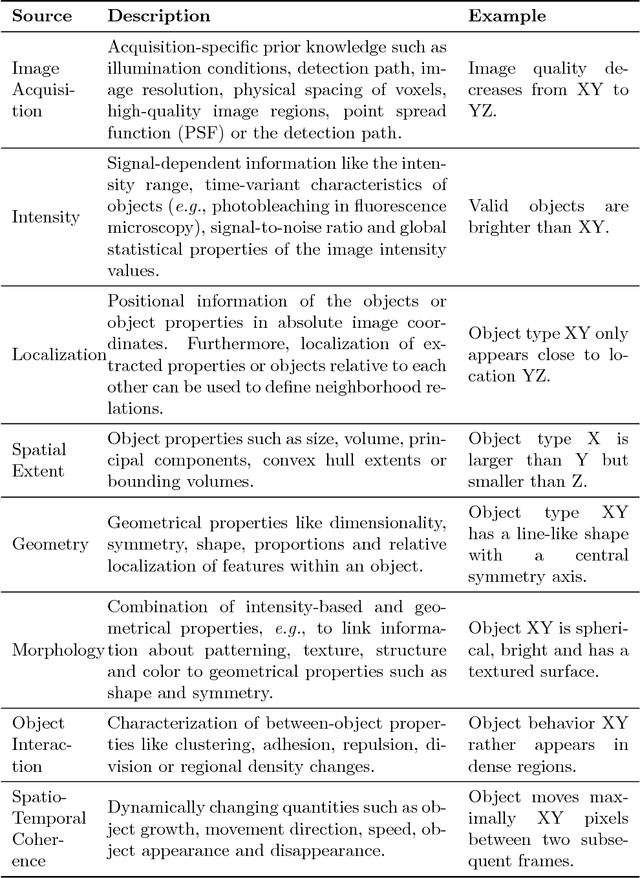
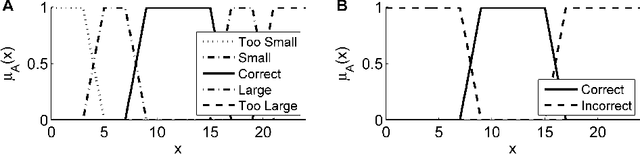
Many automatically analyzable scientific questions are well-posed and offer a variety of information about the expected outcome a priori. Although often being neglected, this prior knowledge can be systematically exploited to make automated analysis operations sensitive to a desired phenomenon or to evaluate extracted content with respect to this prior knowledge. For instance, the performance of processing operators can be greatly enhanced by a more focused detection strategy and the direct information about the ambiguity inherent in the extracted data. We present a new concept for the estimation and propagation of uncertainty involved in image analysis operators. This allows using simple processing operators that are suitable for analyzing large-scale 3D+t microscopy images without compromising the result quality. On the foundation of fuzzy set theory, we transform available prior knowledge into a mathematical representation and extensively use it enhance the result quality of various processing operators. All presented concepts are illustrated on a typical bioimage analysis pipeline comprised of seed point detection, segmentation, multiview fusion and tracking. Furthermore, the functionality of the proposed approach is validated on a comprehensive simulated 3D+t benchmark data set that mimics embryonic development and on large-scale light-sheet microscopy data of a zebrafish embryo. The general concept introduced in this contribution represents a new approach to efficiently exploit prior knowledge to improve the result quality of image analysis pipelines. Especially, the automated analysis of terabyte-scale microscopy data will benefit from sophisticated and efficient algorithms that enable a quantitative and fast readout. The generality of the concept, however, makes it also applicable to practically any other field with processing strategies that are arranged as linear pipelines.
On Modelling Label Uncertainty in Deep Neural Networks: Automatic Estimation of Intra-observer Variability in 2D Echocardiography Quality Assessment
Nov 02, 2019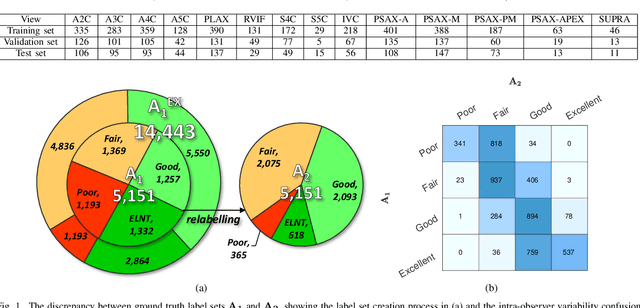
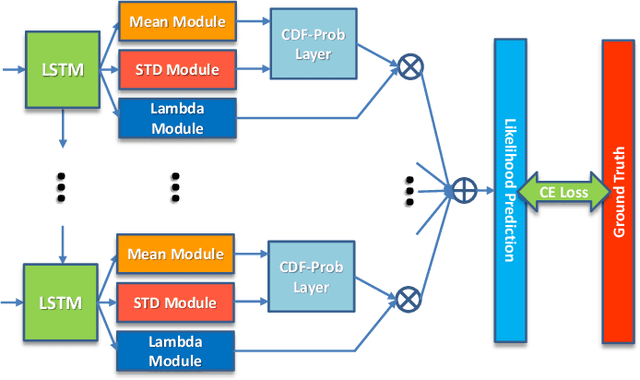
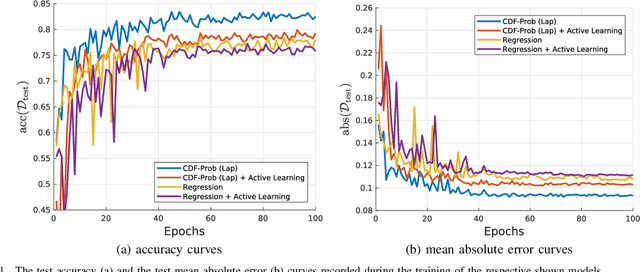
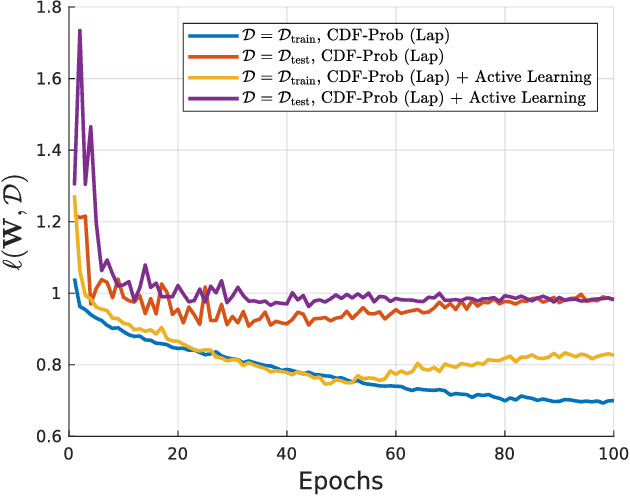
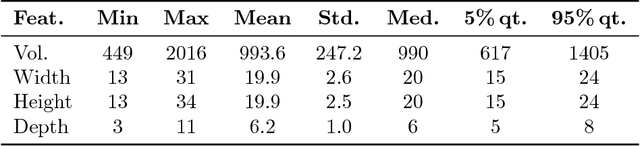
Uncertainty of labels in clinical data resulting from intra-observer variability can have direct impact on the reliability of assessments made by deep neural networks. In this paper, we propose a method for modelling such uncertainty in the context of 2D echocardiography (echo), which is a routine procedure for detecting cardiovascular disease at point-of-care. Echo imaging quality and acquisition time is highly dependent on the operator's experience level. Recent developments have shown the possibility of automating echo image quality quantification by mapping an expert's assessment of quality to the echo image via deep learning techniques. Nevertheless, the observer variability in the expert's assessment can impact the quality quantification accuracy. Here, we aim to model the intra-observer variability in echo quality assessment as an aleatoric uncertainty modelling regression problem with the introduction of a novel method that handles the regression problem with categorical labels. A key feature of our design is that only a single forward pass is sufficient to estimate the level of uncertainty for the network output. Compared to the $0.11 \pm 0.09$ absolute error (in a scale from 0 to 1) archived by the conventional regression method, the proposed method brings the error down to $0.09 \pm 0.08$, where the improvement is statistically significant and equivalents to $5.7\%$ test accuracy improvement. The simplicity of the proposed approach means that it could be generalized to other applications of deep learning in medical imaging, where there is often uncertainty in clinical labels.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge