"Image": models, code, and papers
CALC2.0: Combining Appearance, Semantic and Geometric Information for Robust and Efficient Visual Loop Closure
Oct 30, 2019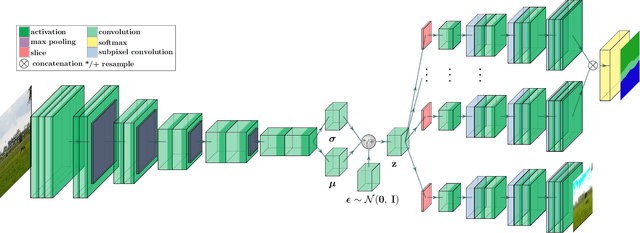
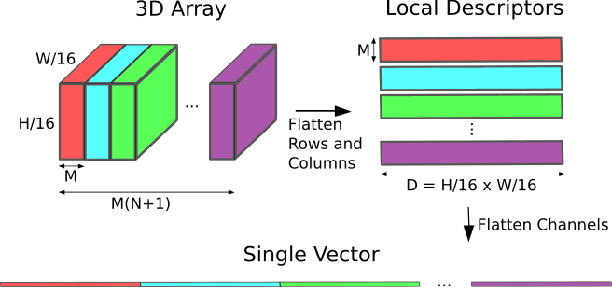

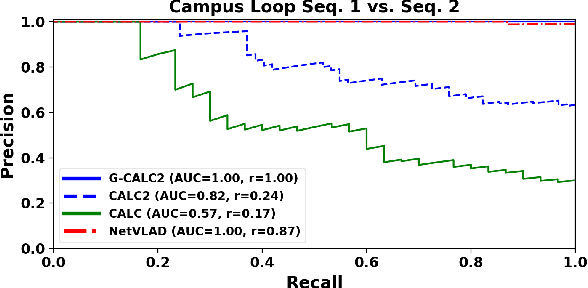
Traditional attempts for loop closure detection typically use hand-crafted features, relying on geometric and visual information only, whereas more modern approaches tend to use semantic, appearance or geometric features extracted from deep convolutional neural networks (CNNs). While these approaches are successful in many applications, they do not utilize all of the information that a monocular image provides, and many of them, particularly the deep-learning based methods, require user-chosen thresholding to actually close loops -- which may impact generality in practical applications. In this work, we address these issues by extracting all three modes of information from a custom deep CNN trained specifically for the task of place recognition. Our network is built upon a combination of a semantic segmentator, Variational Autoencoder (VAE) and triplet embedding network. The network is trained to construct a global feature space to describe both the visual appearance and semantic layout of an image. Then local keypoints are extracted from maximally-activated regions of low-level convolutional feature maps, and keypoint descriptors are extracted from these feature maps in a novel way that incorporates ideas from successful hand-crafted features. These keypoints are matched globally for loop closure candidates, and then used as a final geometric check to refute false positives. As a result, the proposed loop closure detection system requires no touchy thresholding, and is highly robust to false positives -- achieving better precision-recall curves than the state-of-the-art NetVLAD, and with real-time speeds.
Fuzzy-based Propagation of Prior Knowledge to Improve Large-Scale Image Analysis Pipelines
Aug 03, 2016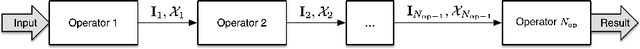
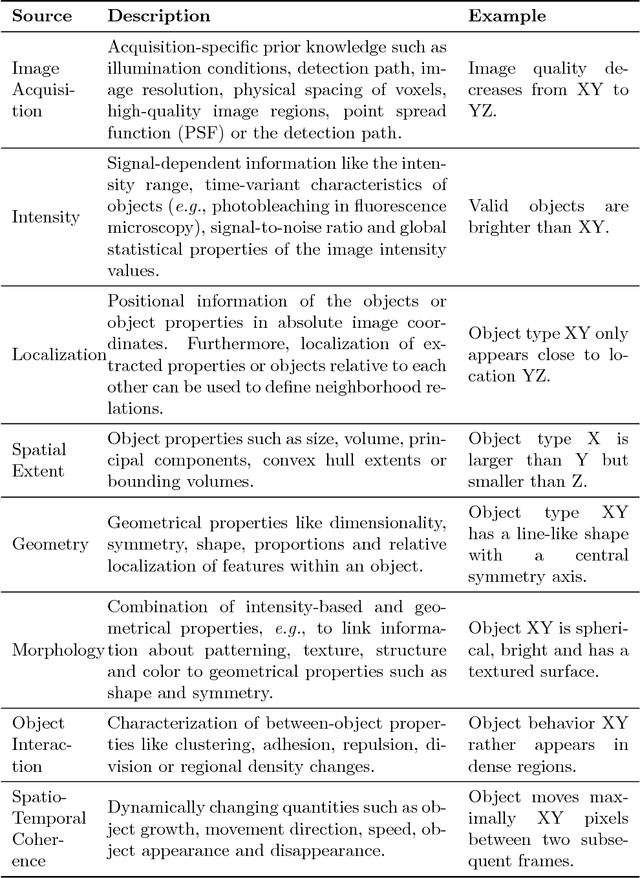
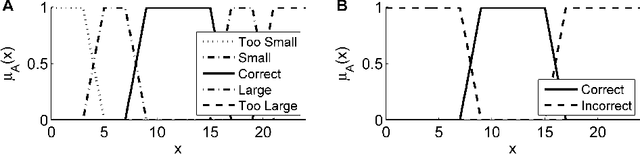
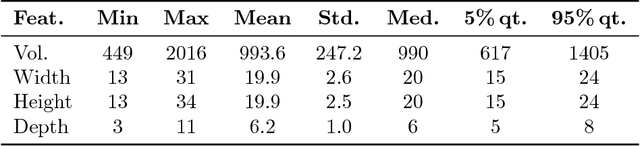
Many automatically analyzable scientific questions are well-posed and offer a variety of information about the expected outcome a priori. Although often being neglected, this prior knowledge can be systematically exploited to make automated analysis operations sensitive to a desired phenomenon or to evaluate extracted content with respect to this prior knowledge. For instance, the performance of processing operators can be greatly enhanced by a more focused detection strategy and the direct information about the ambiguity inherent in the extracted data. We present a new concept for the estimation and propagation of uncertainty involved in image analysis operators. This allows using simple processing operators that are suitable for analyzing large-scale 3D+t microscopy images without compromising the result quality. On the foundation of fuzzy set theory, we transform available prior knowledge into a mathematical representation and extensively use it enhance the result quality of various processing operators. All presented concepts are illustrated on a typical bioimage analysis pipeline comprised of seed point detection, segmentation, multiview fusion and tracking. Furthermore, the functionality of the proposed approach is validated on a comprehensive simulated 3D+t benchmark data set that mimics embryonic development and on large-scale light-sheet microscopy data of a zebrafish embryo. The general concept introduced in this contribution represents a new approach to efficiently exploit prior knowledge to improve the result quality of image analysis pipelines. Especially, the automated analysis of terabyte-scale microscopy data will benefit from sophisticated and efficient algorithms that enable a quantitative and fast readout. The generality of the concept, however, makes it also applicable to practically any other field with processing strategies that are arranged as linear pipelines.
A linear method for camera pair self-calibration and multi-view reconstruction with geometrically verified correspondences
Jun 28, 2019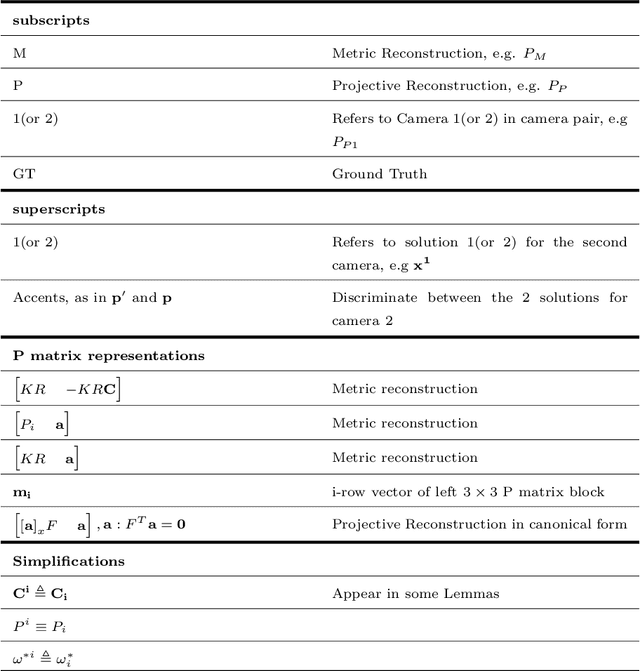
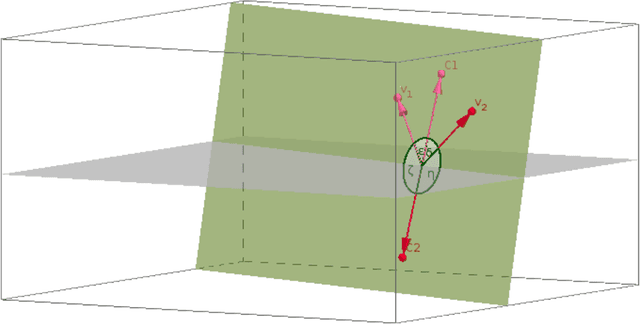
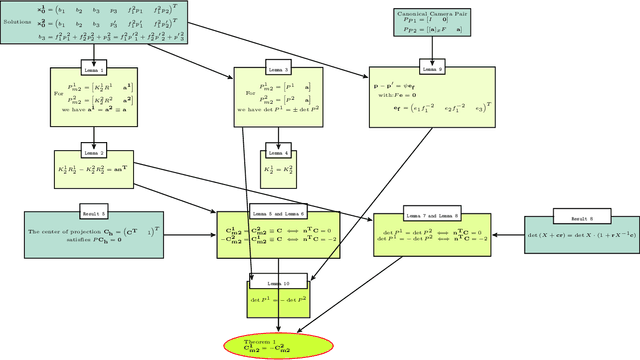
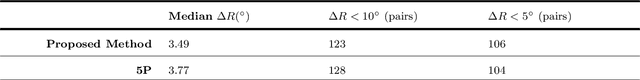
We examine 3D reconstruction of architectural scenes in unordered sets of uncalibrated images. We introduce a linear method to self-calibrate and find the metric reconstruction of a camera pair. We assume unknown and different focal lengths but otherwise known internal camera parameters and a known projective reconstruction of the camera pair. We recover two possible camera configurations in space and use the Cheirality condition, that all 3D scene points are in front of both cameras, to disambiguate the solution. We show in two Theorems, first that the two solutions are in mirror positions and then the relations between their viewing directions. Our new method performs on par (median rotation error $\Delta R = 3.49^{\circ}$) with the standard approach of Kruppa equations ($\Delta R = 3.77^{\circ}$) for self-calibration and 5-Point algorithm for calibrated metric reconstruction of a camera pair. We reject erroneous image correspondences by introducing a method to examine whether point correspondences appear in the same order along $x, y$ image axes in image pairs. We evaluate this method by its precision and recall and show that it improves the robustness of point matches in architectural and general scenes. Finally, we integrate all the introduced methods to a 3D reconstruction pipeline. We utilize the numerous camera pair metric recontructions using rotation-averaging algorithms and a novel method to average focal length estimates.
Multimodal Style Transfer via Graph Cuts
Apr 09, 2019
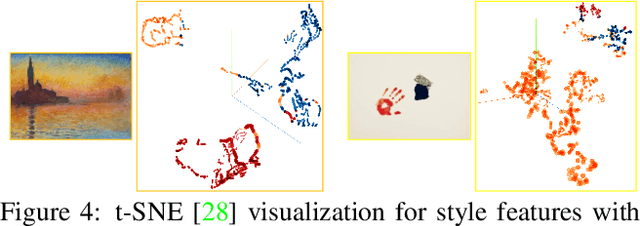
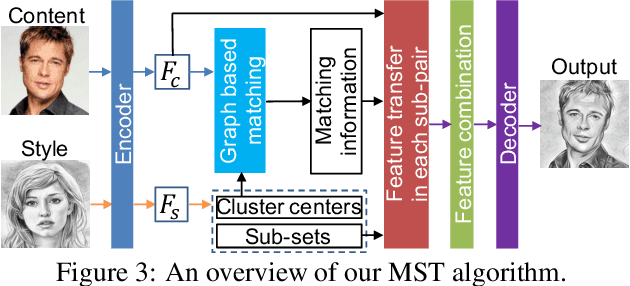

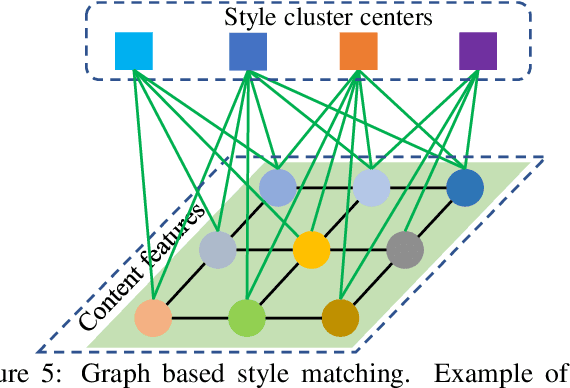
An assumption widely used in recent neural style transfer methods is that image styles can be described by global statics of deep features like Gram or covariance matrices. Alternative approaches have represented styles by decomposing them into local pixel or neural patches. Despite the recent progress, most existing methods treat the semantic patterns of style image uniformly, resulting unpleasing results on complex styles. In this paper, we introduce a more flexible and general universal style transfer technique: multimodal style transfer (MST). MST explicitly considers the matching of semantic patterns in content and style images. Specifically, the style image features are clustered into sub-style components, which are matched with local content features under a graph cut formulation. A reconstruction network is trained to transfer each sub-style and render the final stylized result. Extensive experiments demonstrate the superior effectiveness, robustness and flexibility of MST.
Best of Both Worlds: AutoML Codesign of a CNN and its Hardware Accelerator
Mar 06, 2020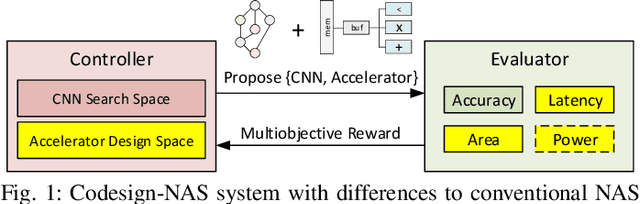
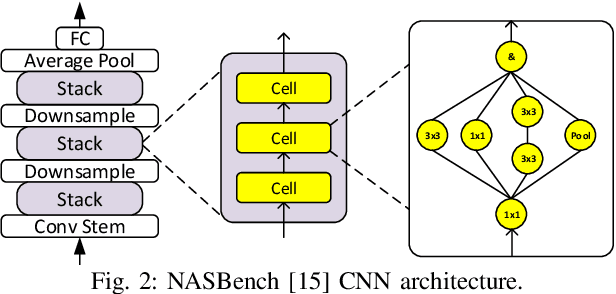
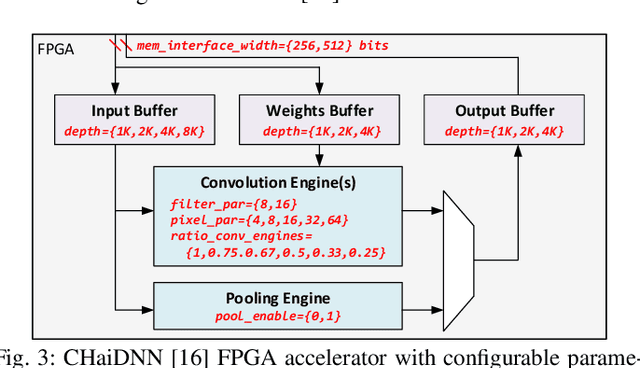
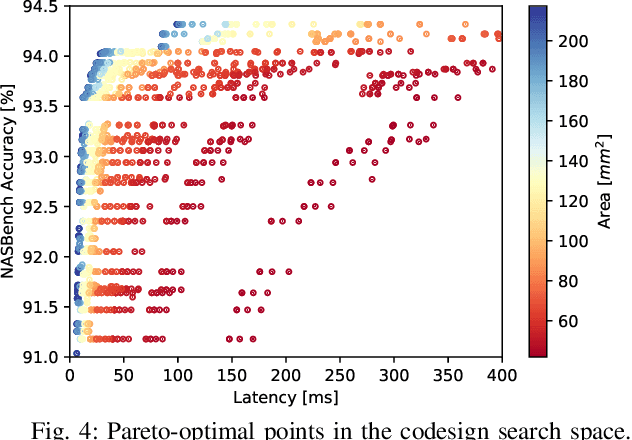
Neural architecture search (NAS) has been very successful at outperforming human-designed convolutional neural networks (CNN) in accuracy, and when hardware information is present, latency as well. However, NAS-designed CNNs typically have a complicated topology, therefore, it may be difficult to design a custom hardware (HW) accelerator for such CNNs. We automate HW-CNN codesign using NAS by including parameters from both the CNN model and the HW accelerator, and we jointly search for the best model-accelerator pair that boosts accuracy and efficiency. We call this Codesign-NAS. In this paper we focus on defining the Codesign-NAS multiobjective optimization problem, demonstrating its effectiveness, and exploring different ways of navigating the codesign search space. For CIFAR-10 image classification, we enumerate close to 4 billion model-accelerator pairs, and find the Pareto frontier within that large search space. This allows us to evaluate three different reinforcement-learning-based search strategies. Finally, compared to ResNet on its most optimal HW accelerator from within our HW design space, we improve on CIFAR-100 classification accuracy by 1.3% while simultaneously increasing performance/area by 41% in just~1000 GPU-hours of running Codesign-NAS.
Memory-efficient Learning for Large-scale Computational Imaging
Dec 11, 2019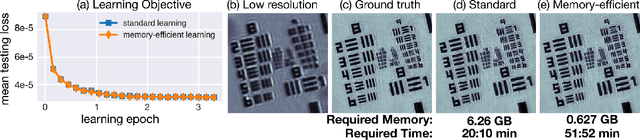
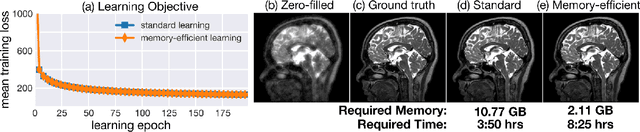
Computational imaging systems jointly design computation and hardware to retrieve information which is not traditionally accessible with standard imaging systems. Recently, critical aspects such as experimental design and image priors are optimized through deep neural networks formed by the unrolled iterations of classical physics-based reconstructions (termed physics-based networks). However, for real-world large-scale systems, computing gradients via backpropagation restricts learning due to memory limitations of graphical processing units. In this work, we propose a memory-efficient learning procedure that exploits the reversibility of the network's layers to enable data-driven design for large-scale computational imaging. We demonstrate our methods practicality on two large-scale systems: super-resolution optical microscopy and multi-channel magnetic resonance imaging.
Detection and skeletonization of single neurons and tracer injections using topological methods
Mar 20, 2020Neuroscientific data analysis has traditionally relied on linear algebra and stochastic process theory. However, the tree-like shapes of neurons cannot be described easily as points in a vector space (the subtraction of two neuronal shapes is not a meaningful operation), and methods from computational topology are better suited to their analysis. Here we introduce methods from Discrete Morse (DM) Theory to extract the tree-skeletons of individual neurons from volumetric brain image data, and to summarize collections of neurons labelled by tracer injections. Since individual neurons are topologically trees, it is sensible to summarize the collection of neurons using a consensus tree-shape that provides a richer information summary than the traditional regional 'connectivity matrix' approach. The conceptually elegant DM approach lacks hand-tuned parameters and captures global properties of the data as opposed to previous approaches which are inherently local. For individual skeletonization of sparsely labelled neurons we obtain substantial performance gains over state-of-the-art non-topological methods (over 10% improvements in precision and faster proofreading). The consensus-tree summary of tracer injections incorporates the regional connectivity matrix information, but in addition captures the collective collateral branching patterns of the set of neurons connected to the injection site, and provides a bridge between single-neuron morphology and tracer-injection data.
Meta Segmentation Network for Ultra-Resolution Medical Images
Feb 19, 2020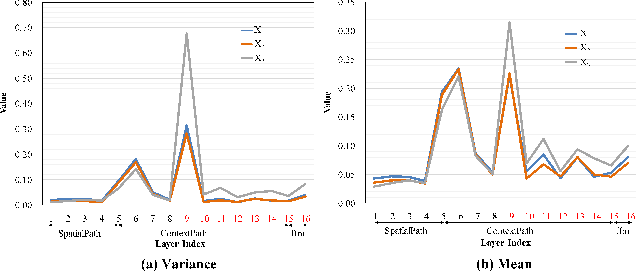
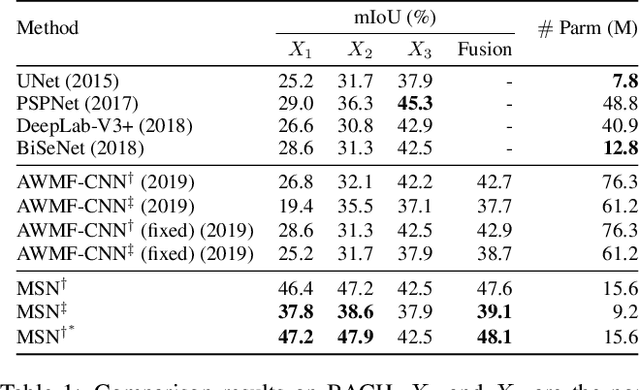
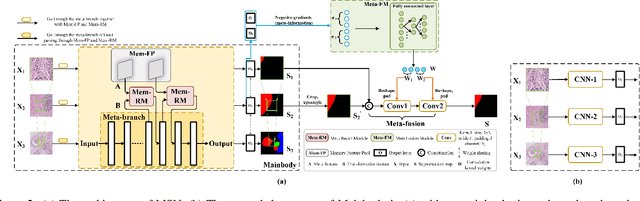
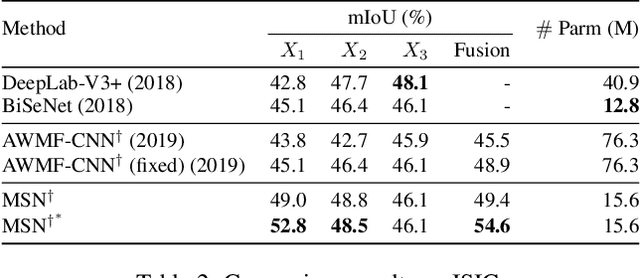
Despite recent progress on semantic segmentation, there still exist huge challenges in medical ultra-resolution image segmentation. The methods based on multi-branch structure can make a good balance between computational burdens and segmentation accuracy. However, the fusion structure in these methods require to be designed elaborately to achieve desirable result, which leads to model redundancy. In this paper, we propose Meta Segmentation Network (MSN) to solve this challenging problem. With the help of meta-learning, the fusion module of MSN is quite simple but effective. MSN can fast generate the weights of fusion layers through a simple meta-learner, requiring only a few training samples and epochs to converge. In addition, to avoid learning all branches from scratch, we further introduce a particular weight sharing mechanism to realize a fast knowledge adaptation and share the weights among multiple branches, resulting in the performance improvement and significant parameters reduction. The experimental results on two challenging ultra-resolution medical datasets BACH and ISIC show that MSN achieves the best performance compared with the state-of-the-art methods.
Visual Navigation Among Humans with Optimal Control as a Supervisor
Mar 20, 2020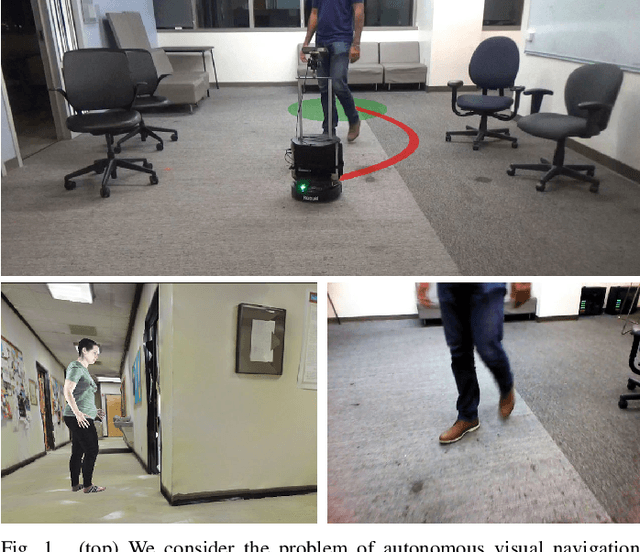
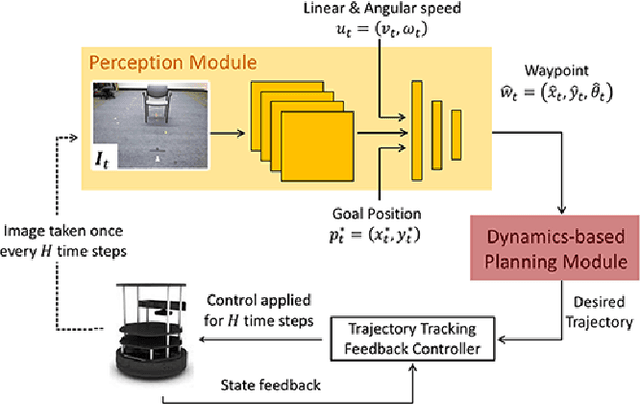
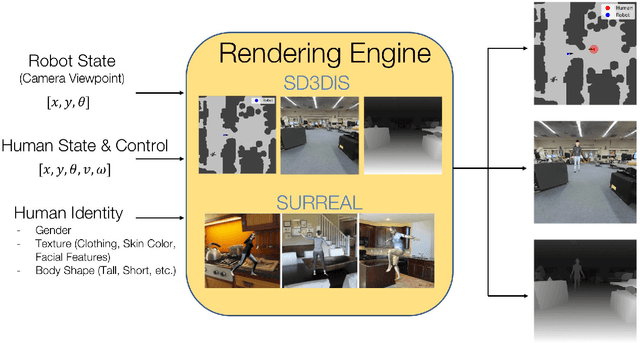

Real world navigation requires robots to operate in unfamiliar, dynamic environments, sharing spaces with humans. Navigating around humans is especially difficult because it requires predicting their future motion, which can be quite challenging. We propose a novel framework for navigation around humans which combines learning-based perception with model-based optimal control. Specifically, we train a Convolutional Neural Network (CNN)-based perception module which maps the robot's visual inputs to a waypoint, or next desired state. This waypoint is then input into planning and control modules which convey the robot safely and efficiently to the goal. To train the CNN we contribute a photo-realistic bench-marking dataset for autonomous robot navigation in the presence of humans. The CNN is trained using supervised learning on images rendered from our photo-realistic dataset. The proposed framework learns to anticipate and react to peoples' motion based only on a monocular RGB image, without explicitly predicting future human motion. Our method generalizes well to unseen buildings and humans in both simulation and real world environments. Furthermore, our experiments demonstrate that combining model-based control and learning leads to better and more data-efficient navigational behaviors as compared to a purely learning based approach. Videos describing our approach and experiments are available on the project website.
Reconstructing continuously heterogeneous structures from single particle cryo-EM with deep generative models
Sep 11, 2019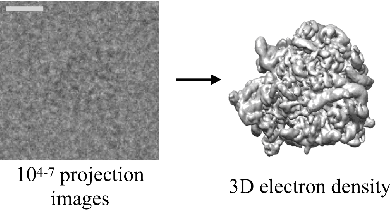
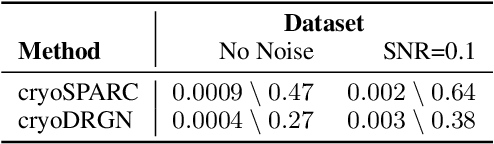
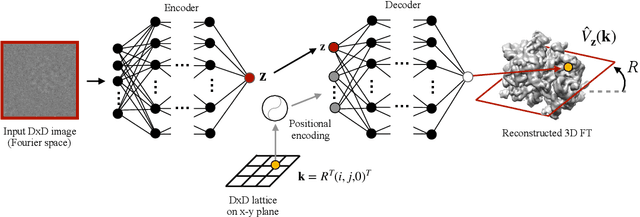
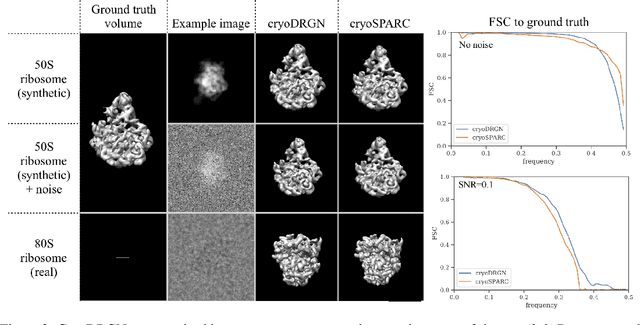
Cryo-electron microscopy (cryo-EM) is a powerful technique for determining the structure of proteins and other macromolecular complexes at near-atomic resolution. In single particle cryo-EM, the central problem is to reconstruct the three-dimensional structure of a macromolecule from $10^{4-7}$ noisy and randomly oriented two-dimensional projections. However, the imaged protein complexes may exhibit structural variability, which complicates reconstruction and is typically addressed using discrete clustering approaches that fail to capture the full range of protein dynamics. Here, we introduce a novel method for cryo-EM reconstruction that extends naturally to modeling continuous generative factors of structural heterogeneity. This method encodes structures in Fourier space using coordinate-based deep neural networks, and trains these networks from unlabeled 2D cryo-EM images by combining exact inference over image orientation with variational inference for structural heterogeneity. We demonstrate that the proposed method, termed cryoDRGN, can perform ab initio reconstruction of 3D protein complexes from simulated and real 2D cryo-EM image data. To our knowledge, cryoDRGN is the first neural network-based approach for cryo-EM reconstruction and the first end-to-end method for directly reconstructing continuous ensembles of protein structures from cryo-EM images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge