"Image": models, code, and papers
HeatNet: Bridging the Day-Night Domain Gap in Semantic Segmentation with Thermal Images
Mar 10, 2020
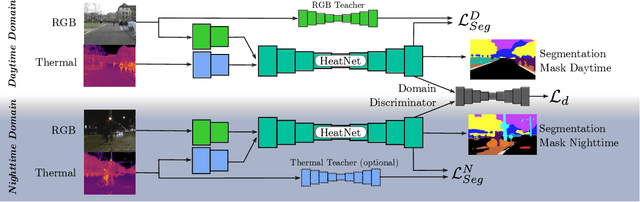


The majority of learning-based semantic segmentation methods are optimized for daytime scenarios and favorable lighting conditions. Real-world driving scenarios, however, entail adverse environmental conditions such as nighttime illumination or glare which remain a challenge for existing approaches. In this work, we propose a multimodal semantic segmentation model that can be applied during daytime and nighttime. To this end, besides RGB images, we leverage thermal images, making our network significantly more robust. We avoid the expensive annotation of nighttime images by leveraging an existing daytime RGB-dataset and propose a teacher-student training approach that transfers the dataset's knowledge to the nighttime domain. We further employ a domain adaptation method to align the learned feature spaces across the domains and propose a novel two-stage training scheme. Furthermore, due to a lack of thermal data for autonomous driving, we present a new dataset comprising over 20,000 time-synchronized and aligned RGB-thermal image pairs. In this context, we also present a novel target-less calibration method that allows for automatic robust extrinsic and intrinsic thermal camera calibration. Among others, we employ our new dataset to show state-of-the-art results for nighttime semantic segmentation.
Simultaneously Learning Architectures and Features of Deep Neural Networks
Jun 11, 2019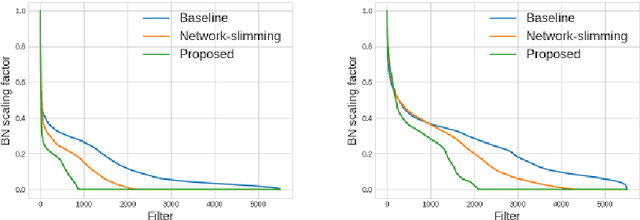
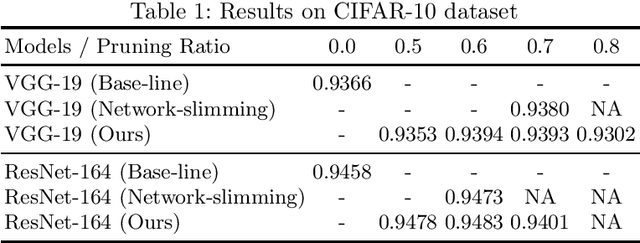
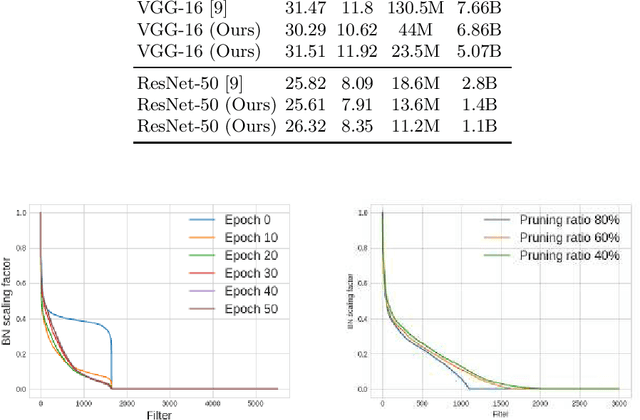
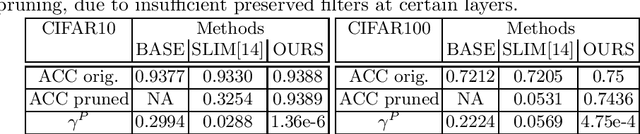
This paper presents a novel method which simultaneously learns the number of filters and network features repeatedly over multiple epochs. We propose a novel pruning loss to explicitly enforces the optimizer to focus on promising candidate filters while suppressing contributions of less relevant ones. In the meanwhile, we further propose to enforce the diversities between filters and this diversity-based regularization term improves the trade-off between model sizes and accuracies. It turns out the interplay between architecture and feature optimizations improves the final compressed models, and the proposed method is compared favorably to existing methods, in terms of both models sizes and accuracies for a wide range of applications including image classification, image compression and audio classification.
An analysis on the use of autoencoders for representation learning: fundamentals, learning task case studies, explainability and challenges
May 21, 2020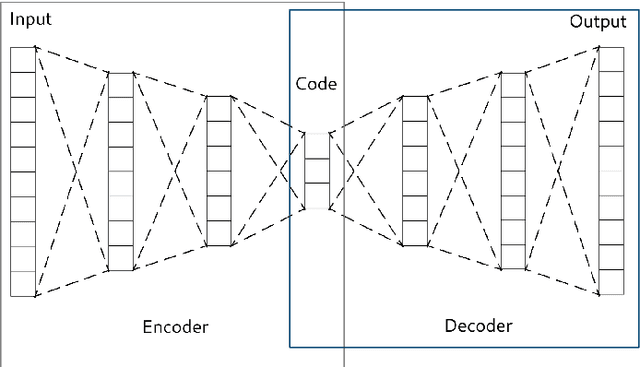
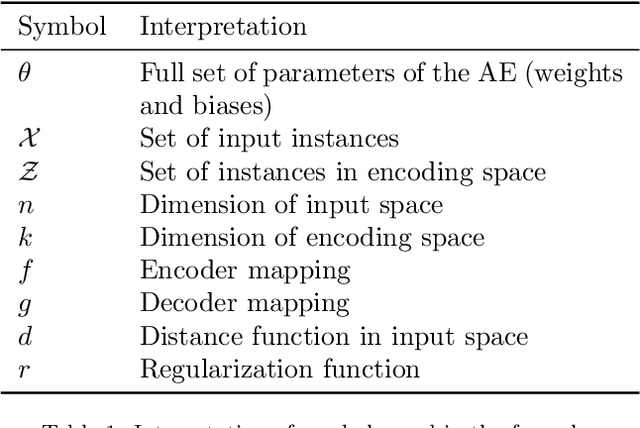
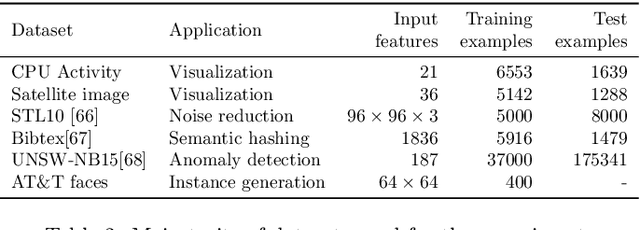
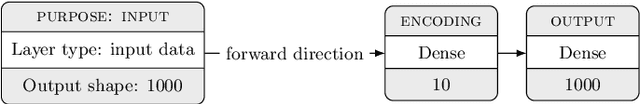
In many machine learning tasks, learning a good representation of the data can be the key to building a well-performant solution. This is because most learning algorithms operate with the features in order to find models for the data. For instance, classification performance can improve if the data is mapped to a space where classes are easily separated, and regression can be facilitated by finding a manifold of data in the feature space. As a general rule, features are transformed by means of statistical methods such as principal component analysis, or manifold learning techniques such as Isomap or locally linear embedding. From a plethora of representation learning methods, one of the most versatile tools is the autoencoder. In this paper we aim to demonstrate how to influence its learned representations to achieve the desired learning behavior. To this end, we present a series of learning tasks: data embedding for visualization, image denoising, semantic hashing, detection of abnormal behaviors and instance generation. We model them from the representation learning perspective, following the state of the art methodologies in each field. A solution is proposed for each task employing autoencoders as the only learning method. The theoretical developments are put into practice using a selection of datasets for the different problems and implementing each solution, followed by a discussion of the results in each case study and a brief explanation of other six learning applications. We also explore the current challenges and approaches to explainability in the context of autoencoders. All of this helps conclude that, thanks to alterations in their structure as well as their objective function, autoencoders may be the core of a possible solution to many problems which can be modeled as a transformation of the feature space.
Alternating Segmentation and Simulation for Contrast Adaptive Tissue Classification
Nov 17, 2018A key feature of magnetic resonance (MR) imaging is its ability to manipulate how the intrinsic tissue parameters of the anatomy ultimately contribute to the contrast properties of the final, acquired image. This flexibility, however, can lead to substantial challenges for segmentation algorithms, particularly supervised methods. These methods require atlases or training data, which are composed of MR image and labeled image pairs. In most cases, the training data are obtained with a fixed acquisition protocol, leading to suboptimal performance when an input data set that requires segmentation has differing contrast properties. This drawback is increasingly significant with the recent movement towards multi-center research studies involving multiple scanners and acquisition protocols. In this work, we propose a new framework for supervised segmentation approaches that is robust to contrast differences between the training MR image and the input image. Our approach uses a generative simulation model within the segmentation process to compensate for the contrast differences. We allow the contrast of the MR image in the training data to vary by simulating a new contrast from the corresponding label image. The model parameters are optimized by a cost function measuring the consistency between the input MR image and its simulation based on a current estimate of the segmentation labels. We provide a proof of concept of this approach by combining a supervised classifier with a simple simulation model, and apply the resulting algorithm to synthetic images and actual MR images.
Weakly Supervised Object Detection with 2D and 3D Regression Neural Networks
Jun 14, 2019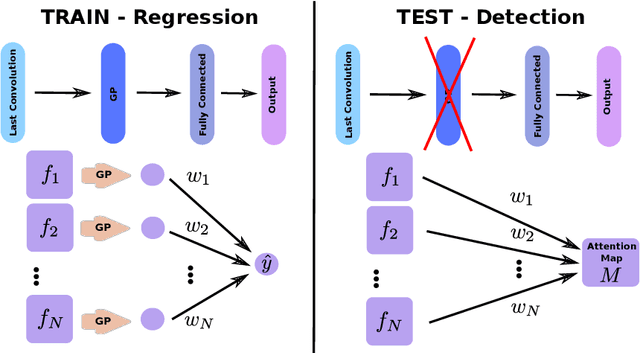
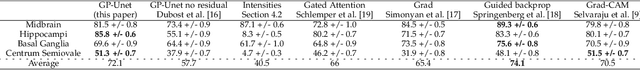
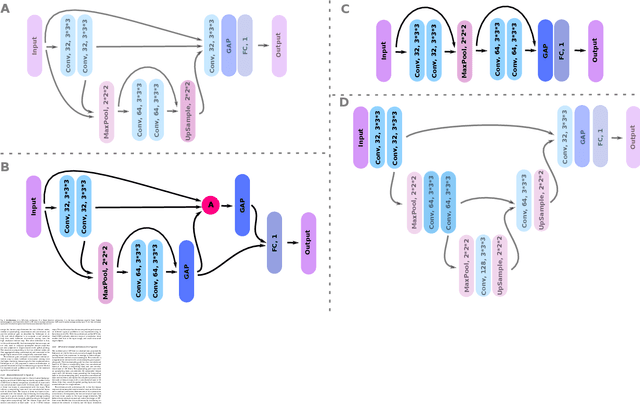
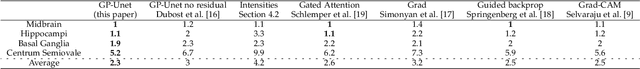
Weakly supervised detection methods can infer the location of target objects in an image without requiring location or appearance information during training. We propose a weakly supervised deep learning method for the detection of objects that appear at multiple locations in an image. The method computes attention maps using the last feature maps of an encoder-decoder network optimized only with global labels: the number of occurrences of the target object in an image. In contrast with previous approaches, attention maps are generated at full input resolution thanks to the decoder part. The proposed approach is compared to multiple state-of-the-art methods in two tasks: the detection of digits in MNIST-based datasets, and the real life application of detection of enlarged perivascular spaces -- a type of brain lesion -- in four brain regions in a dataset of 2202 3D brain MRI scans. In MNIST-based datasets, the proposed method outperforms the other methods. In the brain dataset, several weakly supervised detection methods come close to the human intrarater agreement in each region. The proposed method reaches the lowest number of false positive detections in all brain regions at the operating point, while its average sensitivity is similar to that of the other best methods.
A Fast, Semi-Automatic Brain Structure Segmentation Algorithm for Magnetic Resonance Imaging
Apr 21, 2019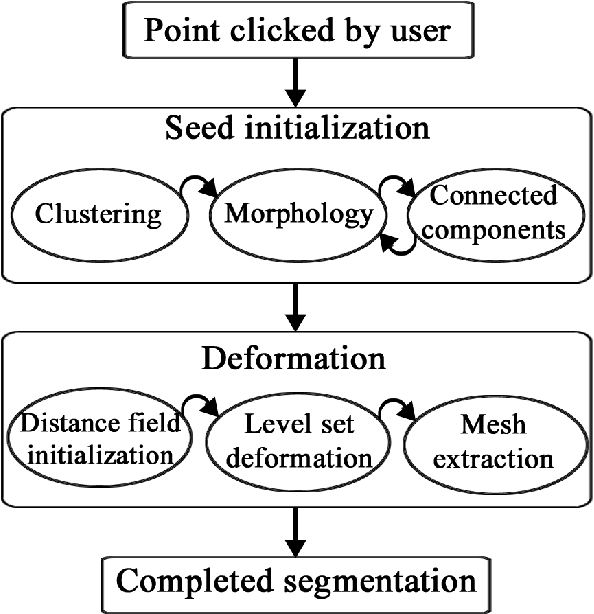
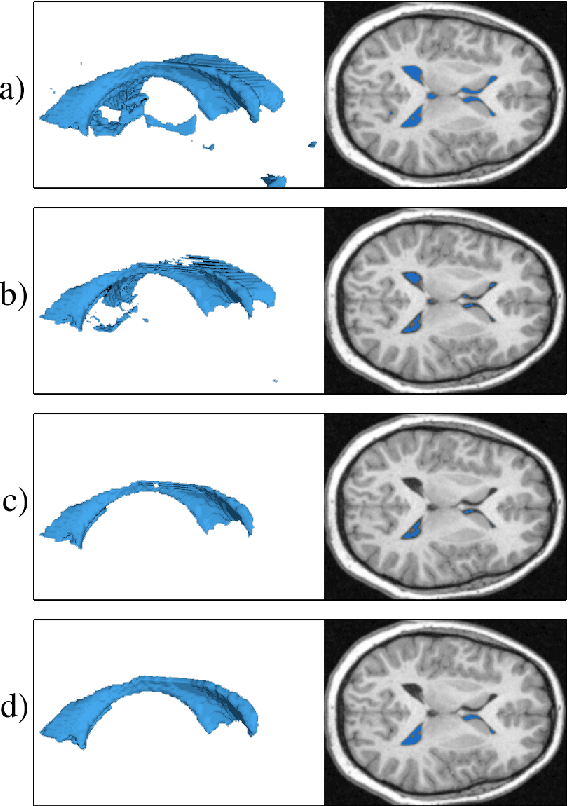

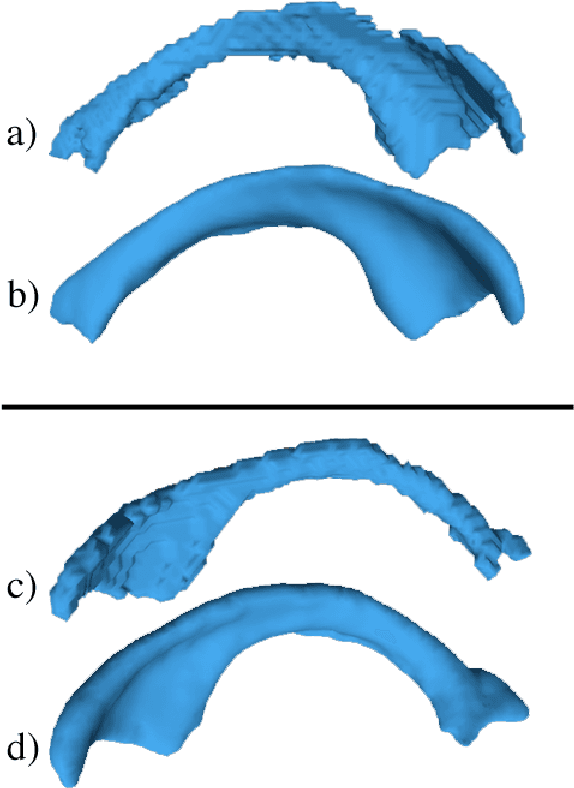
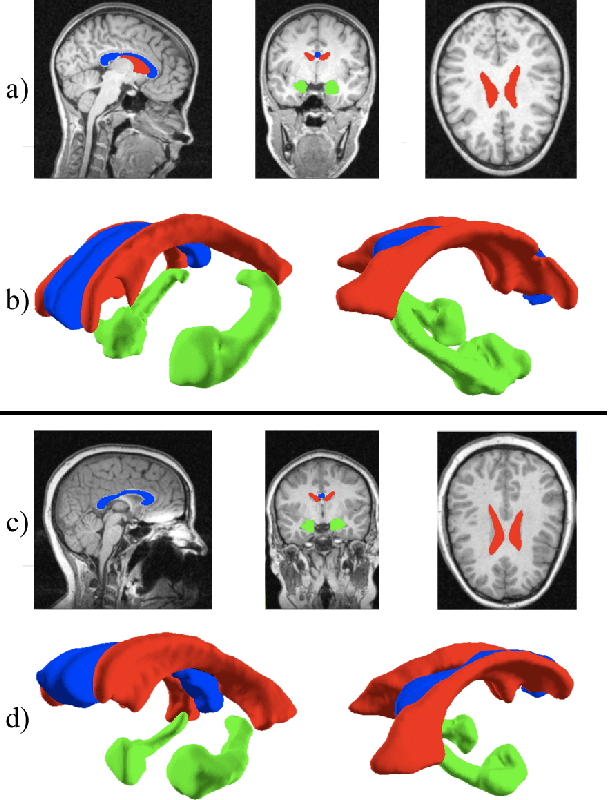
Medical image segmentation has become an essential technique in clinical and research-oriented applications. Because manual segmentation methods are tedious, and fully automatic segmentation lacks the flexibility of human intervention or correction, semi-automatic methods have become the preferred type of medical image segmentation. We present a hybrid, semi-automatic segmentation method in 3D that integrates both region-based and boundary-based procedures. Our method differs from previous hybrid methods in that we perform region-based and boundary-based approaches separately, which allows for more efficient segmentation. A region-based technique is used to generate an initial seed contour that roughly represents the boundary of a target brain structure, alleviating the local minima problem in the subsequent model deformation phase. The contour is deformed under a unique force equation independent of image edges. Experiments on MRI data show that this method can achieve high accuracy and efficiency primarily due to the unique seed initialization technique.
A Biologically Interpretable Two-stage Deep Neural Network (BIT-DNN) For Hyperspectral Imagery Classification
Apr 19, 2020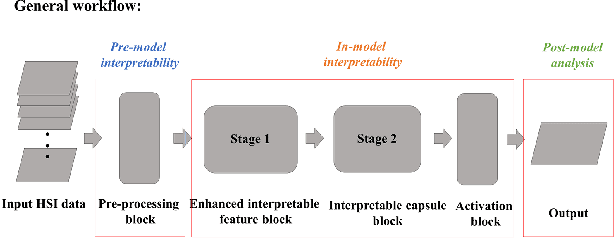
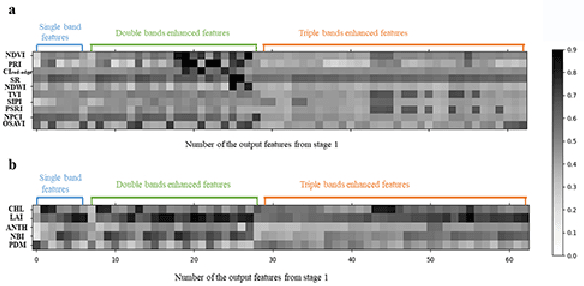
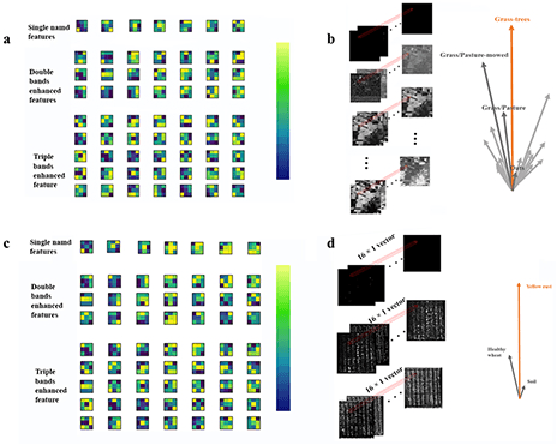
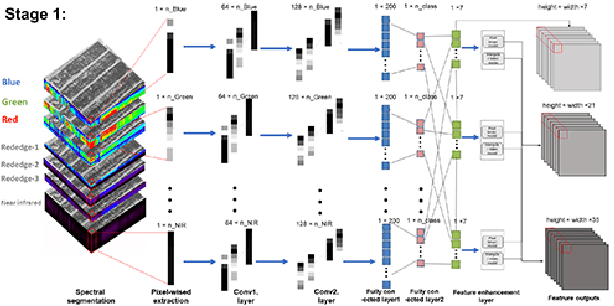
Spectral-spatial based deep learning models have recently proven to be effective in hyperspectral image (HSI) classification for various earth monitoring applications such as land cover classification and agricultural monitoring. However, due to the nature of "black-box" model representation, how to explain and interpret the learning process and the model decision remains an open problem. This study proposes an interpretable deep learning model -- a biologically interpretable two-stage deep neural network (BIT-DNN), by integrating biochemical and biophysical associated information into the proposed framework, capable of achieving both high accuracy and interpretability on HSI based classification tasks. The proposed model introduces a two-stage feature learning process. In the first stage, an enhanced interpretable feature block extracts low-level spectral features associated with the biophysical and biochemical attributes of the target entities; and in the second stage, an interpretable capsule block extracts and encapsulates the high-level joint spectral-spatial features into the featured tensors representing the hierarchical structure of the biophysical and biochemical attributes of the target ground entities, which provides the model an improved performance on classification and intrinsic interpretability. We have tested and evaluated the model using two real HSI datasets for crop type recognition and crop disease recognition tasks and compared it with six state-of-the-art machine learning models. The results demonstrate that the proposed model has competitive advantages in terms of both classification accuracy and model interpretability.
Learning to be Global Optimizer
Mar 10, 2020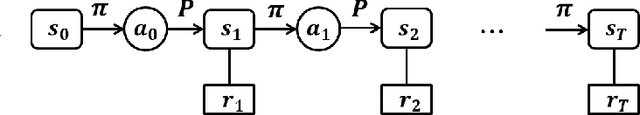
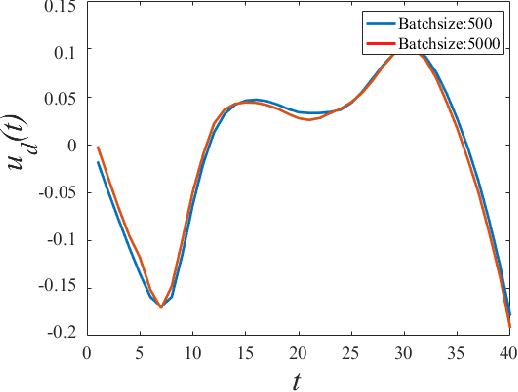
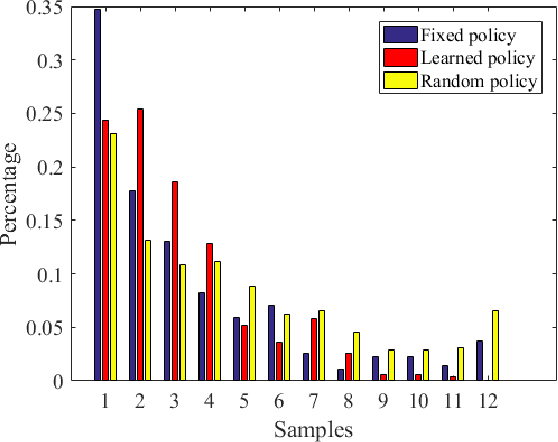
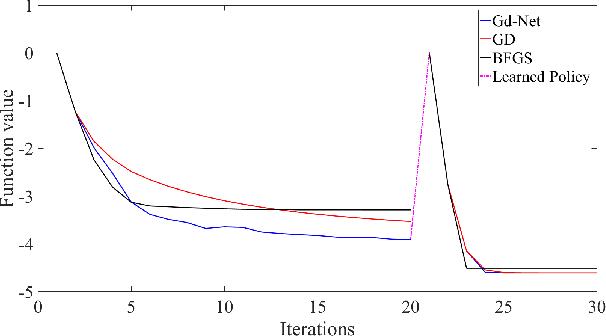
The advancement of artificial intelligence has cast a new light on the development of optimization algorithm. This paper proposes to learn a two-phase (including a minimization phase and an escaping phase) global optimization algorithm for smooth non-convex functions. For the minimization phase, a model-driven deep learning method is developed to learn the update rule of descent direction, which is formalized as a nonlinear combination of historical information, for convex functions. We prove that the resultant algorithm with the proposed adaptive direction guarantees convergence for convex functions. Empirical study shows that the learned algorithm significantly outperforms some well-known classical optimization algorithms, such as gradient descent, conjugate descent and BFGS, and performs well on ill-posed functions. The escaping phase from local optimum is modeled as a Markov decision process with a fixed escaping policy. We further propose to learn an optimal escaping policy by reinforcement learning. The effectiveness of the escaping policies is verified by optimizing synthesized functions and training a deep neural network for CIFAR image classification. The learned two-phase global optimization algorithm demonstrates a promising global search capability on some benchmark functions and machine learning tasks.
Intra-model Variability in COVID-19 Classification Using Chest X-ray Images
Apr 30, 2020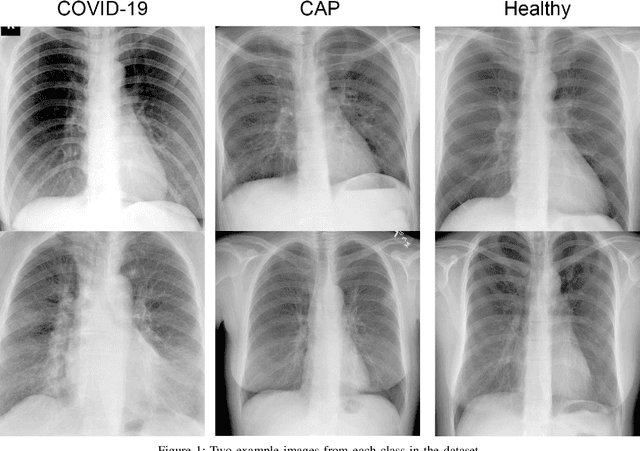
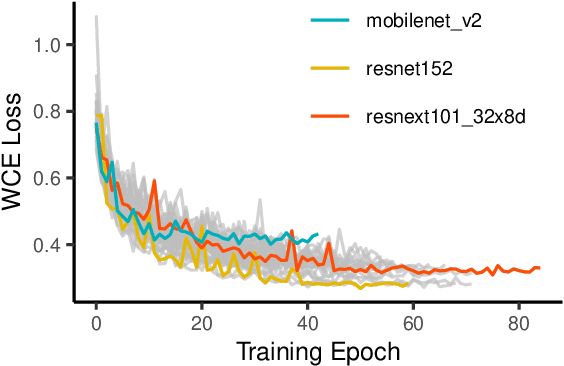
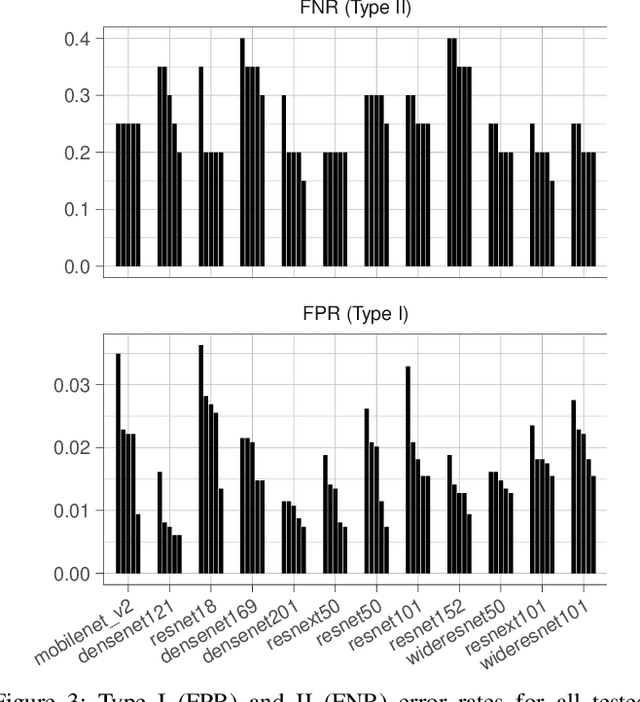
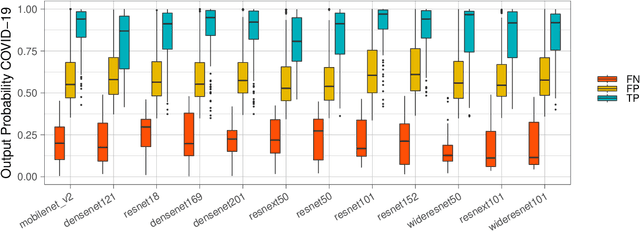
X-ray and computed tomography (CT) scanning technologies for COVID-19 screening have gained significant traction in AI research since the start of the coronavirus pandemic. Despite these continuous advancements for COVID-19 screening, many concerns remain about model reliability when used in a clinical setting. Much has been published, but with limited transparency in expected model performance. We set out to address this limitation through a set of experiments to quantify baseline performance metrics and variability for COVID-19 detection in chest x-ray for 12 common deep learning architectures. Specifically, we adopted an experimental paradigm controlling for train-validation-test split and model architecture where the source of prediction variability originates from model weight initialization, random data augmentation transformations, and batch shuffling. Each model architecture was trained 5 separate times on identical train-validation-test splits of a publicly available x-ray image dataset provided by Cohen et al. (2020). Results indicate that even within model architectures, model behavior varies in a meaningful way between trained models. Best performing models achieve a false negative rate of 3 out of 20 for detecting COVID-19 in a hold-out set. While these results show promise in using AI for COVID-19 screening, they further support the urgent need for diverse medical imaging datasets for model training in a way that yields consistent prediction outcomes. It is our hope that these modeling results accelerate work in building a more robust dataset and a viable screening tool for COVID-19.
Learning to Reconstruct Confocal Microscopy Stacks from Single Light Field Images
Mar 24, 2020We present a novel deep learning approach to reconstruct confocal microscopy stacks from single light field images. To perform the reconstruction, we introduce the LFMNet, a novel neural network architecture inspired by the U-Net design. It is able to reconstruct with high-accuracy a 112x112x57.6$\mu m^3$ volume (1287x1287x64 voxels) in 50ms given a single light field image of 1287x1287 pixels, thus dramatically reducing 720-fold the time for confocal scanning of assays at the same volumetric resolution and 64-fold the required storage. To prove the applicability in life sciences, our approach is evaluated both quantitatively and qualitatively on mouse brain slices with fluorescently labelled blood vessels. Because of the drastic reduction in scan time and storage space, our setup and method are directly applicable to real-time in vivo 3D microscopy. We provide analysis of the optical design, of the network architecture and of our training procedure to optimally reconstruct volumes for a given target depth range. To train our network, we built a data set of 362 light field images of mouse brain blood vessels and the corresponding aligned set of 3D confocal scans, which we use as ground truth. The data set will be made available for research purposes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge