"Image": models, code, and papers
Hierarchical Kinematic Human Mesh Recovery
Mar 09, 2020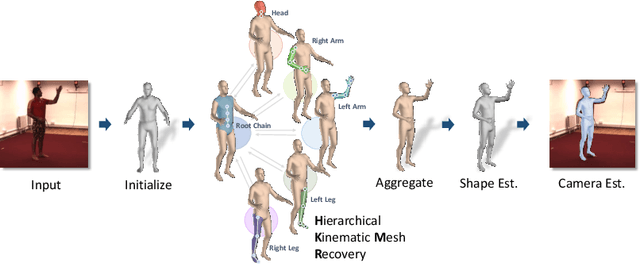
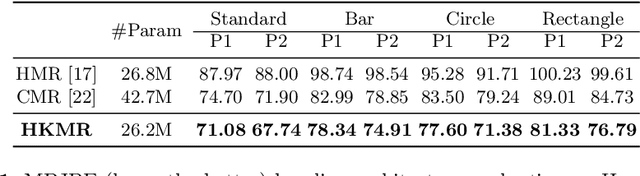
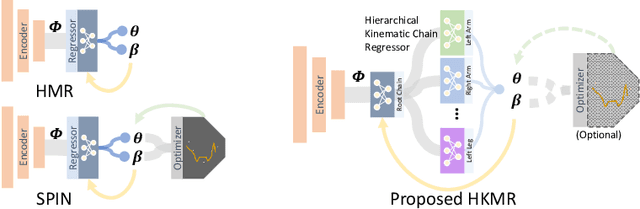

We consider the problem of estimating a parametric model of 3D human mesh from a single image. While there has been substantial recent progress in this area with direct regression of model parameters, these methods only implicitly exploit the human body kinematic structure, leading to sub-optimal use of the model prior. In this work, we address this gap by proposing a new technique for regression of human parametric model that is explicitly informed by the known hierarchical structure, including joint interdependencies of the model. This results in a strong prior-informed design of the regressor architecture and an associated hierarchical optimization that is flexible to be used in conjunction with the current standard frameworks for 3D human mesh recovery. We demonstrate these aspects by means of extensive experiments on standard benchmark datasets, showing how our proposed new design outperforms several existing and popular methods, establishing new state-of-the-art results. With our explicit consideration of joint interdependencies, our proposed method is equipped to infer joints even under data corruptions, which we demonstrate with experiments under varying degrees of occlusion.
Video from Stills: Lensless Imaging with Rolling Shutter
May 30, 2019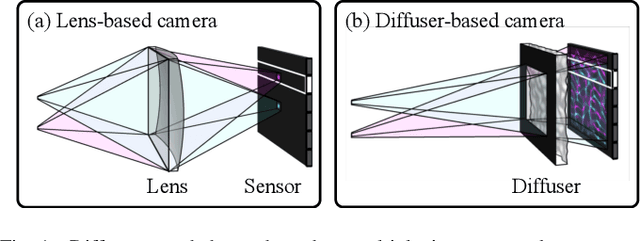
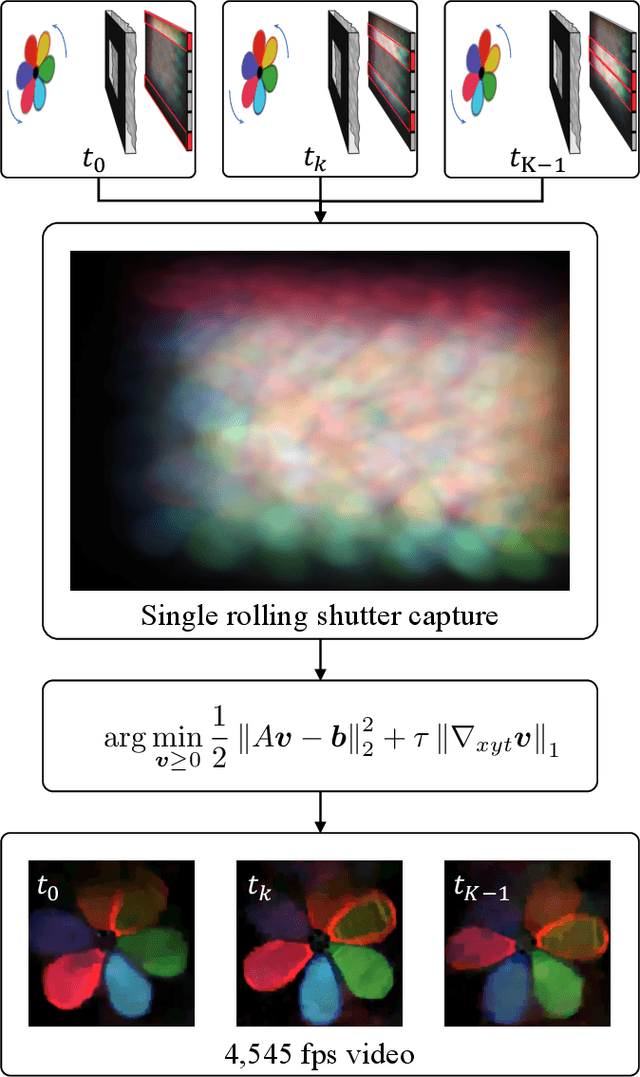

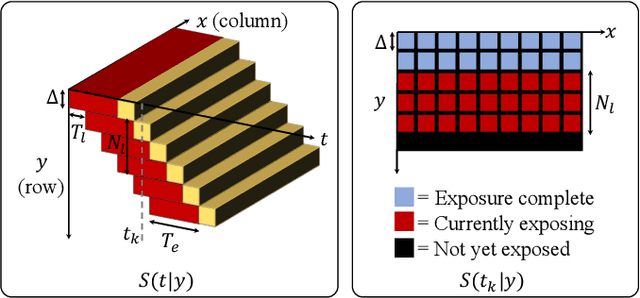
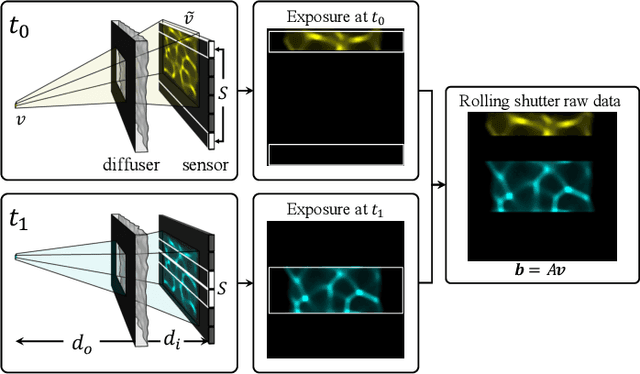
Because image sensor chips have a finite bandwidth with which to read out pixels, recording video typically requires a trade-off between frame rate and pixel count. Compressed sensing techniques can circumvent this trade-off by assuming that the image is compressible. Here, we propose using multiplexing optics to spatially compress the scene, enabling information about the whole scene to be sampled from a row of sensor pixels, which can be read off quickly via a rolling shutter CMOS sensor. Conveniently, such multiplexing can be achieved with a simple lensless, diffuser-based imaging system. Using sparse recovery methods, we are able to recover 140 video frames at over 4,500 frames per second, all from a single captured image with a rolling shutter sensor. Our proof-of-concept system uses easily-fabricated diffusers paired with an off-the-shelf sensor. The resulting prototype enables compressive encoding of high frame rate video into a single rolling shutter exposure, and exceeds the sampling-limited performance of an equivalent global shutter system for sufficiently sparse objects.
A Hierarchical Probabilistic U-Net for Modeling Multi-Scale Ambiguities
May 30, 2019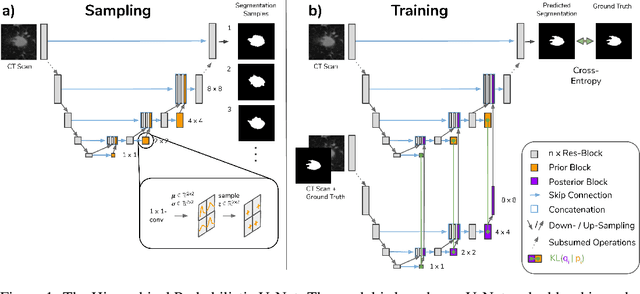
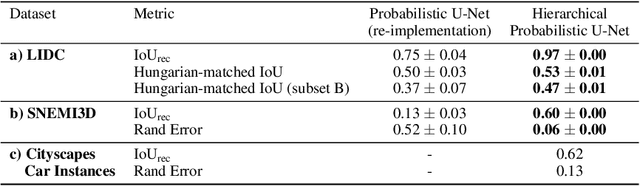
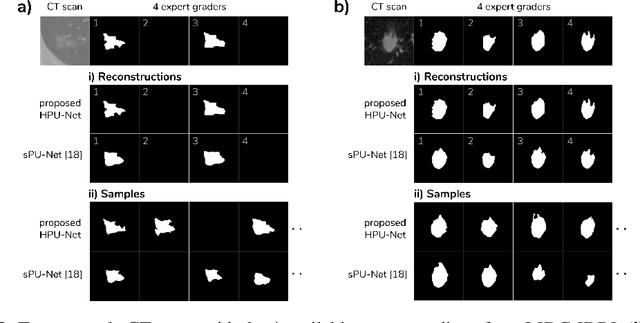
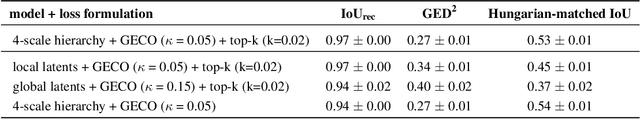
Medical imaging only indirectly measures the molecular identity of the tissue within each voxel, which often produces only ambiguous image evidence for target measures of interest, like semantic segmentation. This diversity and the variations of plausible interpretations are often specific to given image regions and may thus manifest on various scales, spanning all the way from the pixel to the image level. In order to learn a flexible distribution that can account for multiple scales of variations, we propose the Hierarchical Probabilistic U-Net, a segmentation network with a conditional variational auto-encoder (cVAE) that uses a hierarchical latent space decomposition. We show that this model formulation enables sampling and reconstruction of segmenations with high fidelity, i.e. with finely resolved detail, while providing the flexibility to learn complex structured distributions across scales. We demonstrate these abilities on the task of segmenting ambiguous medical scans as well as on instance segmentation of neurobiological and natural images. Our model automatically separates independent factors across scales, an inductive bias that we deem beneficial in structured output prediction tasks beyond segmentation.
Unified Multifaceted Feature Learning for Person Re-Identification
Nov 20, 2019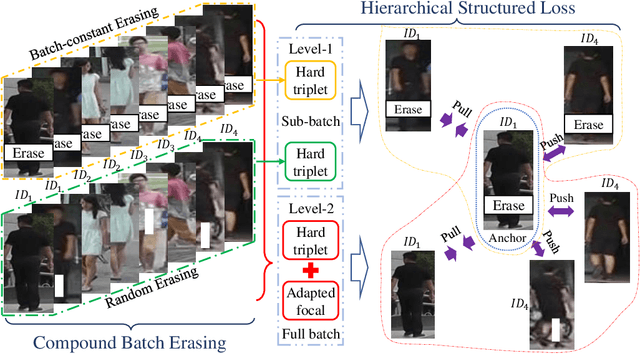
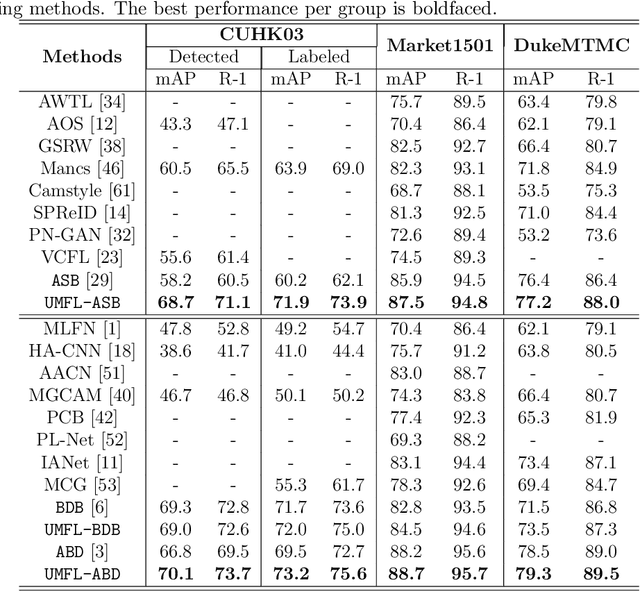
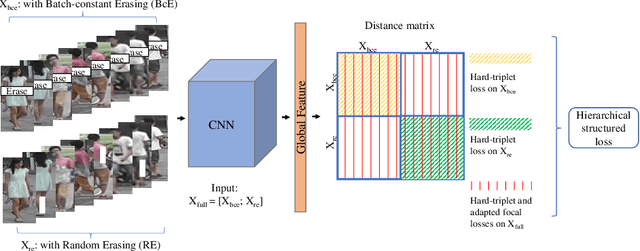
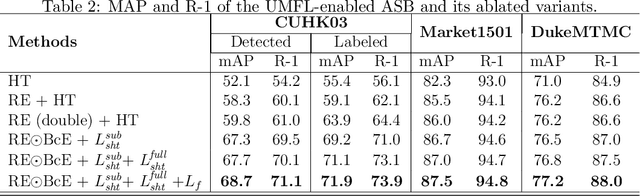
Person re-identification (ReID) aims at re-identifying persons from different viewpoints across multiple cameras, of which it is of great importance to learn multifaceted features expressed in different parts of a person, e.g., clothes, bags, and other accessories in the main body, appearance in the head, and shoes in the foot. To learn such features, existing methods are focused on the striping-based approach that builds multi-branch neural networks to learn local features in each part of the identities, with one-branch network dedicated to one part. This results in complex models with a large number of parameters. To address this issue, this paper proposes to learn the multifaceted features in a simple unified single-branch neural network. The Unified Multifaceted Feature Learning (UMFL) framework is introduced to fulfill this goal, which consists of two key collaborative modules: compound batch image erasing (including batch constant erasing and random erasing) and hierarchical structured loss. The loss structures the augmented images resulted by the two types of image erasing in a two-level hierarchy and enforces multifaceted attention to different parts. As we show in the extensive experimental results on four benchmark person ReID datasets, despite the use of significantly simplified network structure, our method performs substantially better than state-of-the-art competing methods. Our method can also effectively generalize to vehicle ReID, achieving similar improvement on two vehicle ReID datasets.
A Light Field Camera Calibration Method Using Sub-Aperture Related Bipartition Projection Model and 4D Corner Detection
Jan 11, 2020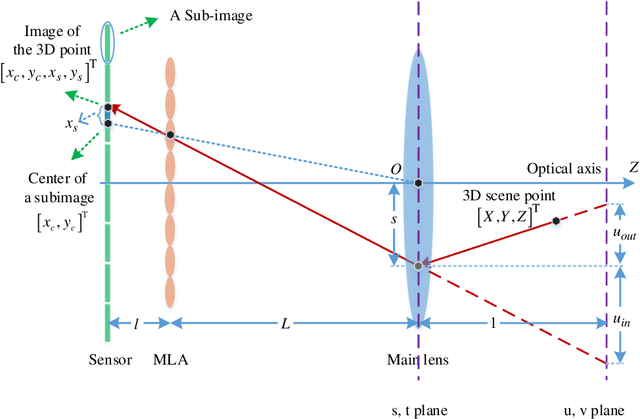
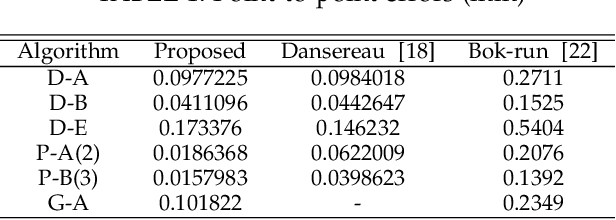
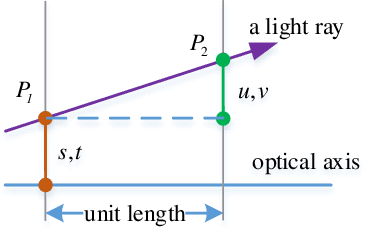
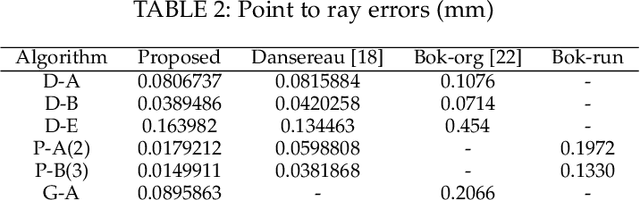
Accurate calibration of intrinsic parameters of the light field (LF) camera is the key issue of many applications, especially of the 3D reconstruction. In this paper, we propose the Sub-Aperture Related Bipartition (SARB) projection model to characterize the LF camera. This projection model is composed with two sets of parameters targeting on center view sub-aperture and relations between sub-apertures. Moreover, we also propose a corner point detection algorithm which fully utilizes the 4D LF information in the raw image. Experimental results have demonstrated the accuracy and robustness of the corner detection method. Both the 2D re-projection errors in the lateral direction and errors in the depth direction are minimized because two sets of parameters in SARB projection model are solved separately.
ANDA: A Novel Data Augmentation Technique Applied to Salient Object Detection
Oct 03, 2019



In this paper, we propose a novel data augmentation technique (ANDA) applied to the Salient Object Detection (SOD) context. Standard data augmentation techniques proposed in the literature, such as image cropping, rotation, flipping, and resizing, only generate variations of the existing examples, providing a limited generalization. Our method has the novelty of creating new images, by combining an object with a new background while retaining part of its salience in this new context; To do so, the ANDA technique relies on the linear combination between labeled salient objects and new backgrounds, generated by removing the original salient object in a process known as image inpainting. Our proposed technique allows for more precise control of the object's position and size while preserving background information. Aiming to evaluate our proposed method, we trained multiple deep neural networks and compared the effect that our technique has in each one. We also compared our method with other data augmentation techniques. Our findings show that depending on the network improvement can be up to 14.1% in the F-measure and decay of up to 2.6% in the Mean Absolute Error.
A Convolutional Forward and Back-Projection Model for Fan-Beam Geometry
Jul 24, 2019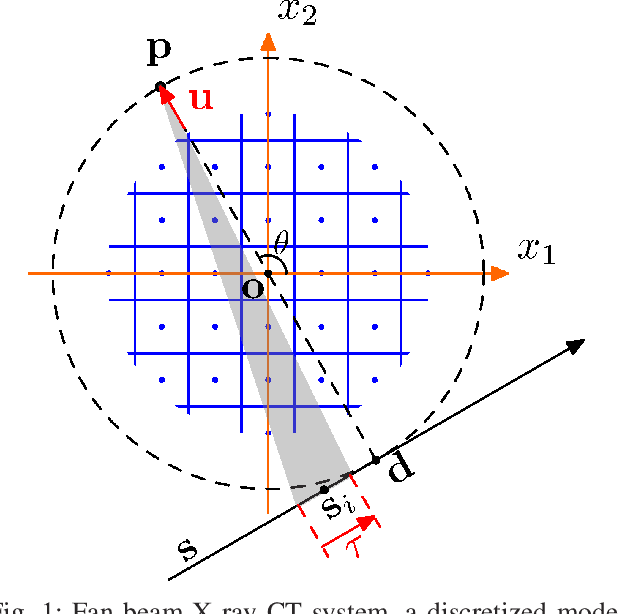
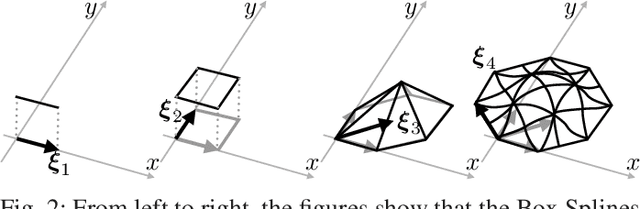
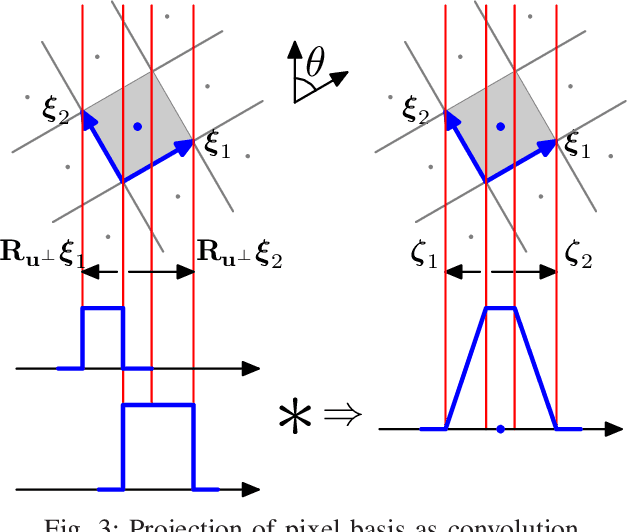
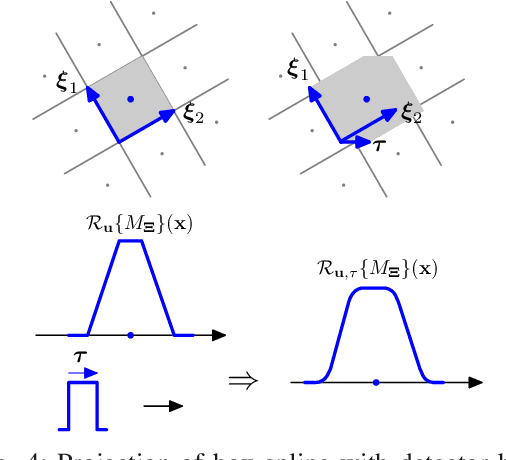
Iterative methods for tomographic image reconstruction have great potential for enabling high quality imaging from low-dose projection data. The computational burden of iterative reconstruction algorithms, however, has been an impediment in their adoption in practical CT reconstruction problems. We present an approach for highly efficient and accurate computation of forward model for image reconstruction in fan-beam geometry in X-ray CT. The efficiency of computations makes this approach suitable for large-scale optimization algorithms with on-the-fly, memory-less, computations of the forward and back-projection. Our experiments demonstrate the improvements in accuracy as well as efficiency of our model, specifically for first-order box splines (i.e., pixel-basis) compared to recently developed methods for this purpose, namely Look-up Table-based Ray Integration (LTRI) and Separable Footprints (SF) in 2-D.
Patch-based Non-Local Bayesian Networks for Blind Confocal Microscopy Denoising
Mar 25, 2020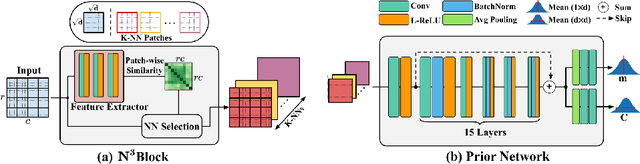
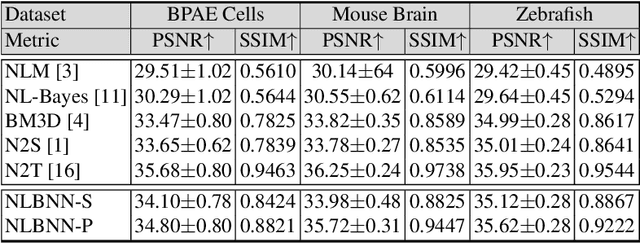
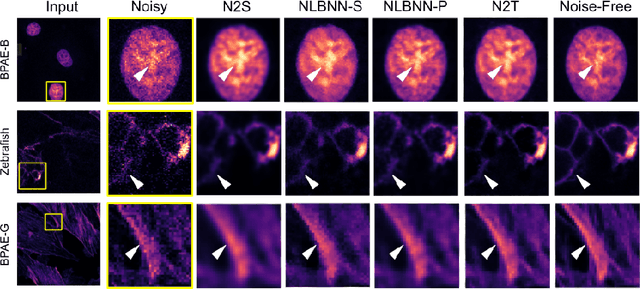
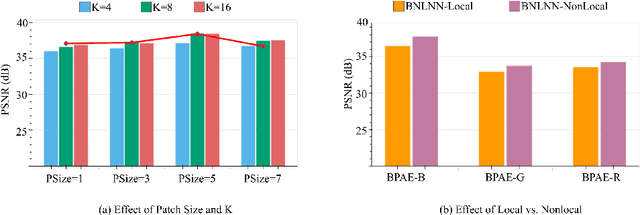
Confocal microscopy is essential for histopathologic cell visualization and quantification. Despite its significant role in biology, fluorescence confocal microscopy suffers from the presence of inherent noise during image acquisition. Non-local patch-wise Bayesian mean filtering (NLB) was until recently the state-of-the-art denoising approach. However, classic denoising methods have been outperformed by neural networks in recent years. In this work, we propose to exploit the strengths of NLB in the framework of Bayesian deep learning. We do so by designing a convolutional neural network and training it to learn parameters of a Gaussian model approximating the prior on noise-free patches given their nearest, similar yet non-local, neighbors. We then apply Bayesian reasoning to leverage the prior and information from the noisy patch in the process of approximating the noise-free patch. Specifically, we use the closed-form analytic \textit{maximum a posteriori} (MAP) estimate in the NLB algorithm to obtain the noise-free patch that maximizes the posterior distribution. The performance of our proposed method is evaluated on confocal microscopy images with real noise Poisson-Gaussian noise. Our experiments reveal the superiority of our approach against state-of-the-art unsupervised denoising techniques.
Degenerative Adversarial NeuroImage Nets for 3D Simulations: Application in Longitudinal MRI
Dec 03, 2019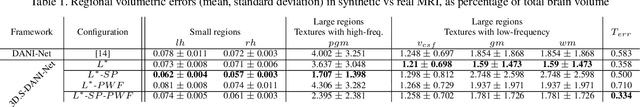
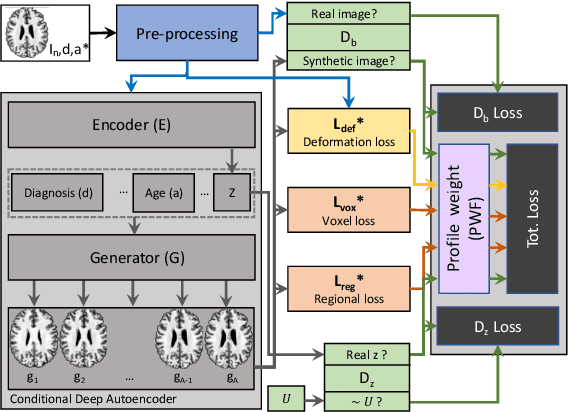
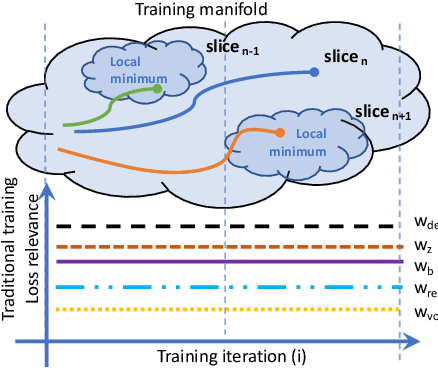
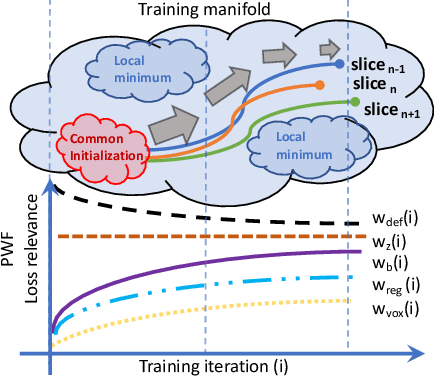
The recent success of deep learning together with the availability of large medical imaging datasets have enabled researchers to improve our understanding of complex chronic medical conditions such as neurodegenerative diseases. The possibility of predicting realistic and accurate images would be a breakthrough for many clinical healthcare applications. However, current image simulators designed to model neurodegenerative disease progression present limitations that preclude their utility in clinical practice. These limitations include personalization of disease progression and the ability to synthesize spatiotemporal images in high resolution. In particular, memory limitations prohibit full 3D image models, necessitating various techniques to discard spatiotemporal information, such as patch-based approaches. In this work, we introduce a novel technique to address this challenge, called Profile Weight Functions (PWF). We demonstrate its effectiveness integrated within our new deep learning framework, showing that it enables the extension to 3D of a recent state-of-the-art 2D approach. To our knowledge, we are the first to implement a personalized disease progression simulator able to predict accurate, personalised, high-resolution, 3D MRI. In particular, we trained a model of ageing and Alzheimer's disease progression using 9652 T1-weighted (longitudinal) MRI from the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset and validated on a separate test set of 1283 MRI (also from ADNI, random partition). We validated our model by analyzing its capability to synthesize MRI that produce accurate volumes of specific brain regions associated with neurodegeneration. Our experiments demonstrate the effectiveness of our solution to provide a 3D simulation that produces accurate and convincing synthetic MRI that emulate ageing and disease progression.
MaxUp: A Simple Way to Improve Generalization of Neural Network Training
Feb 20, 2020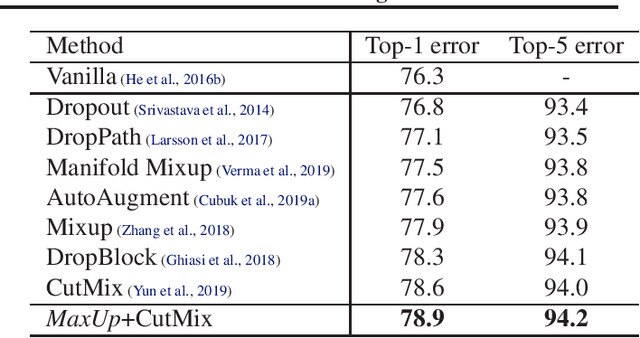
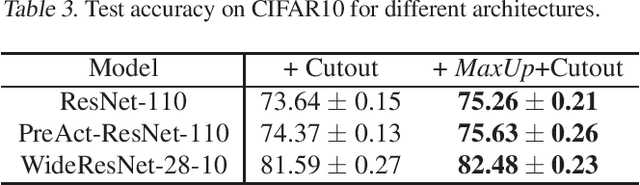
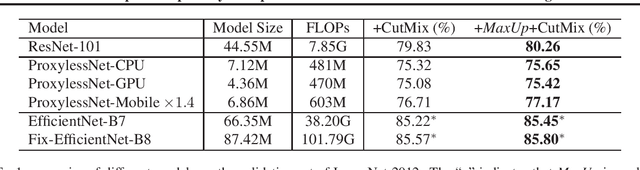
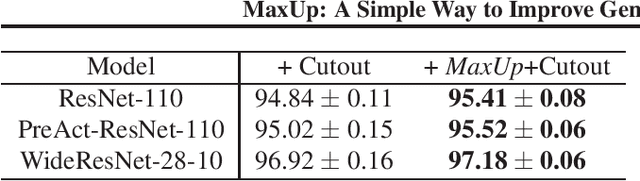

We propose \emph{MaxUp}, an embarrassingly simple, highly effective technique for improving the generalization performance of machine learning models, especially deep neural networks. The idea is to generate a set of augmented data with some random perturbations or transforms and minimize the maximum, or worst case loss over the augmented data. By doing so, we implicitly introduce a smoothness or robustness regularization against the random perturbations, and hence improve the generation performance. For example, in the case of Gaussian perturbation, \emph{MaxUp} is asymptotically equivalent to using the gradient norm of the loss as a penalty to encourage smoothness. We test \emph{MaxUp} on a range of tasks, including image classification, language modeling, and adversarial certification, on which \emph{MaxUp} consistently outperforms the existing best baseline methods, without introducing substantial computational overhead. In particular, we improve ImageNet classification from the state-of-the-art top-1 accuracy $85.5\%$ without extra data to $85.8\%$. Code will be released soon.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge