"Image": models, code, and papers
Assessing Robustness of Deep learning Methods in Dermatological Workflow
Jan 15, 2020
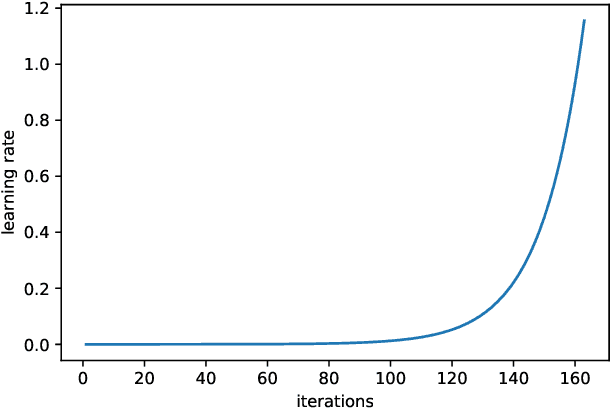
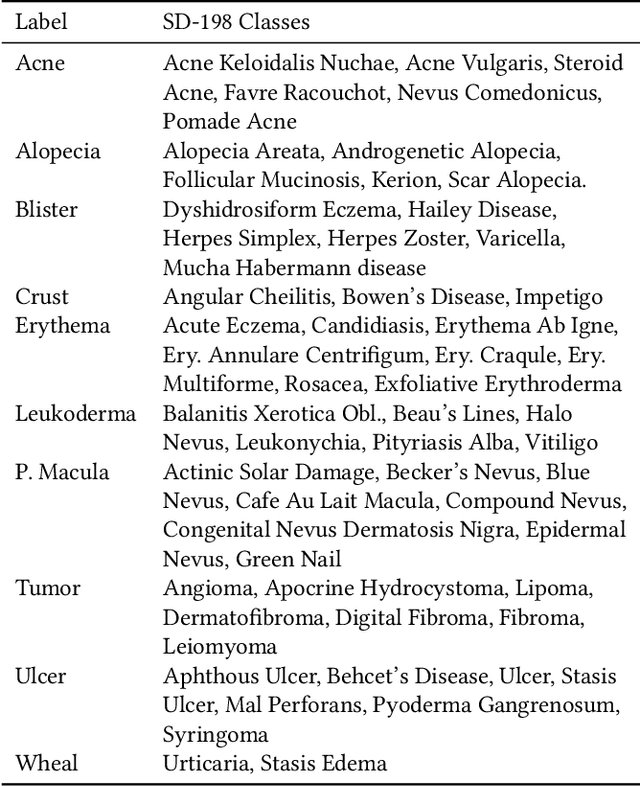
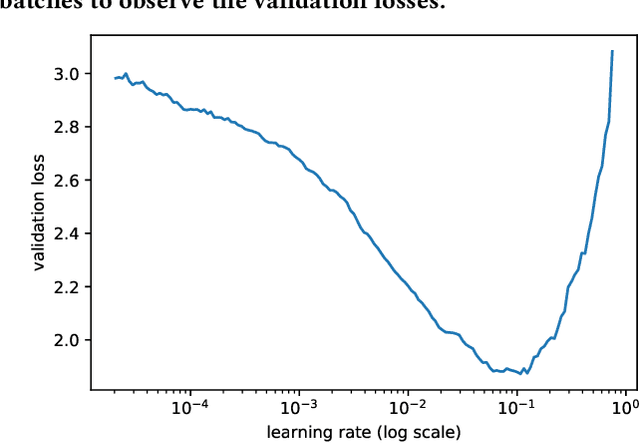
This paper aims to evaluate the suitability of current deep learning methods for clinical workflow especially by focusing on dermatology. Although deep learning methods have been attempted to get dermatologist level accuracy in several individual conditions, it has not been rigorously tested for common clinical complaints. Most projects involve data acquired in well-controlled laboratory conditions. This may not reflect regular clinical evaluation where corresponding image quality is not always ideal. We test the robustness of deep learning methods by simulating non-ideal characteristics on user submitted images of ten classes of diseases. Assessing via imitated conditions, we have found the overall accuracy to drop and individual predictions change significantly in many cases despite of robust training.
Leveraging Domain Knowledge to improve EM image segmentation with Lifted Multicuts
May 25, 2019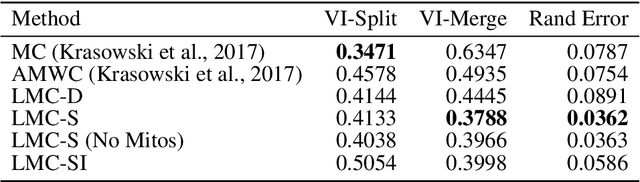
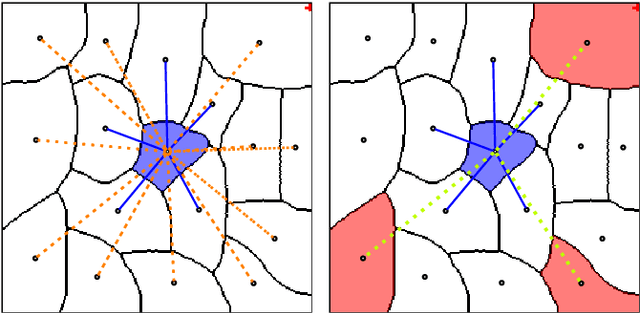
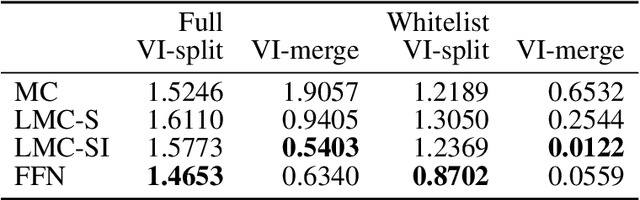
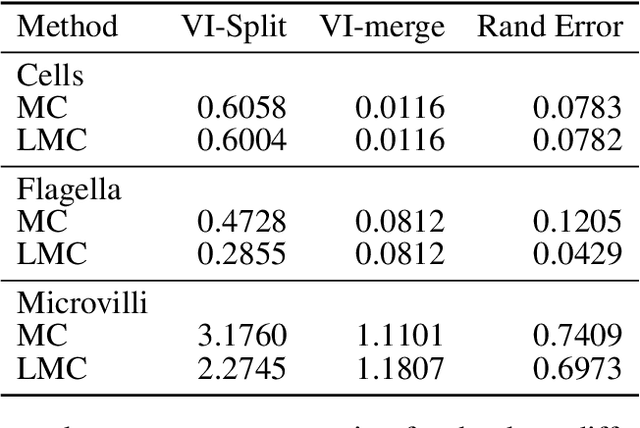
The throughput of electron microscopes has increased significantly in recent years, enabling detailed analysis of cell morphology and ultrastructure. Analysis of neural circuits at single-synapse resolution remains the flagship target of this technique, but applications to cell and developmental biology are also starting to emerge at scale. The amount of data acquired in such studies makes manual instance segmentation, a fundamental step in many analysis pipelines, impossible. While automatic segmentation approaches have improved significantly thanks to the adoption of convolutional neural networks, their accuracy still lags behind human annotations and requires additional manual proof-reading. A major hindrance to further improvements is the limited field of view of the segmentation networks preventing them from exploiting the expected cell morphology or other prior biological knowledge which humans use to inform their segmentation decisions. In this contribution, we show how such domain-specific information can be leveraged by expressing it as long-range interactions in a graph partitioning problem known as the lifted multicut problem. Using this formulation, we demonstrate significant improvement in segmentation accuracy for three challenging EM segmentation problems from neuroscience and cell biology.
S2A: Wasserstein GAN with Spatio-Spectral Laplacian Attention for Multi-Spectral Band Synthesis
Apr 08, 2020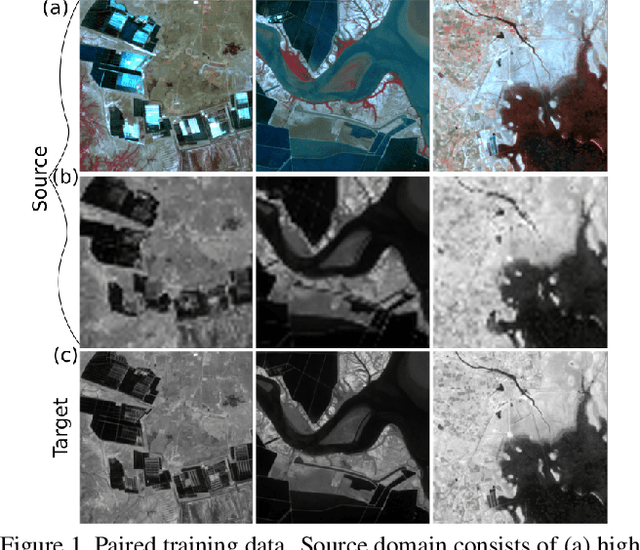
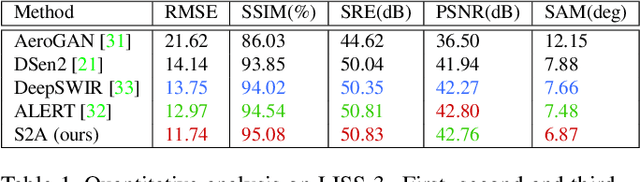
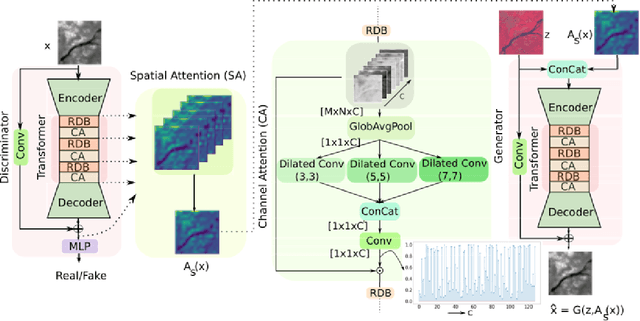
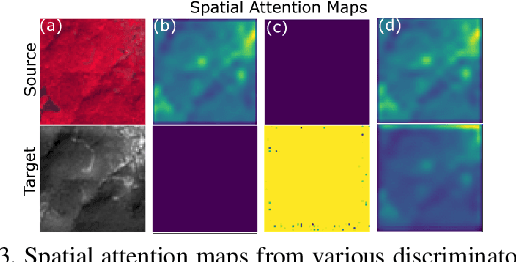
Intersection of adversarial learning and satellite image processing is an emerging field in remote sensing. In this study, we intend to address synthesis of high resolution multi-spectral satellite imagery using adversarial learning. Guided by the discovery of attention mechanism, we regulate the process of band synthesis through spatio-spectral Laplacian attention. Further, we use Wasserstein GAN with gradient penalty norm to improve training and stability of adversarial learning. In this regard, we introduce a new cost function for the discriminator based on spatial attention and domain adaptation loss. We critically analyze the qualitative and quantitative results compared with state-of-the-art methods using widely adopted evaluation metrics. Our experiments on datasets of three different sensors, namely LISS-3, LISS-4, and WorldView-2 show that attention learning performs favorably against state-of-the-art methods. Using the proposed method we provide an additional data product in consistent with existing high resolution bands. Furthermore, we synthesize over 4000 high resolution scenes covering various terrains to analyze scientific fidelity. At the end, we demonstrate plausible large scale real world applications of the synthesized band.
LinesToFacePhoto: Face Photo Generation from Lines with Conditional Self-Attention Generative Adversarial Network
Oct 20, 2019
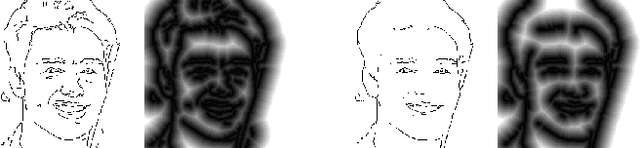
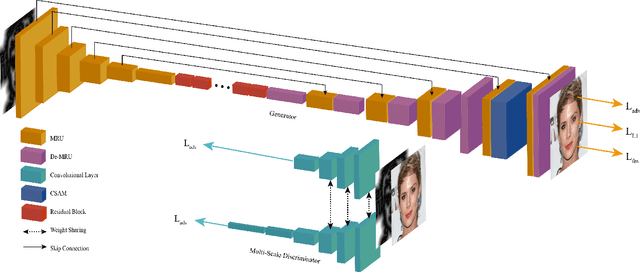
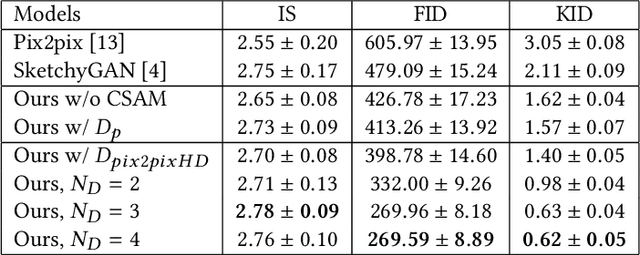

In this paper, we explore the task of generating photo-realistic face images from lines. Previous methods based on conditional generative adversarial networks (cGANs) have shown their power to generate visually plausible images when a conditional image and an output image share well-aligned structures. However, these models fail to synthesize face images with a whole set of well-defined structures, e.g. eyes, noses, mouths, etc., especially when the conditional line map lacks one or several parts. To address this problem, we propose a conditional self-attention generative adversarial network (CSAGAN). We introduce a conditional self-attention mechanism to cGANs to capture long-range dependencies between different regions in faces. We also build a multi-scale discriminator. The large-scale discriminator enforces the completeness of global structures and the small-scale discriminator encourages fine details, thereby enhancing the realism of generated face images. We evaluate the proposed model on the CelebA-HD dataset by two perceptual user studies and three quantitative metrics. The experiment results demonstrate that our method generates high-quality facial images while preserving facial structures. Our results outperform state-of-the-art methods both quantitatively and qualitatively.
Deform-GAN:An Unsupervised Learning Model for Deformable Registration
Feb 26, 2020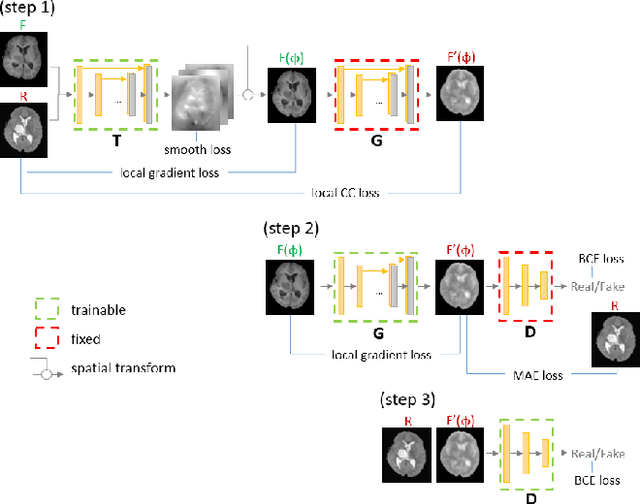
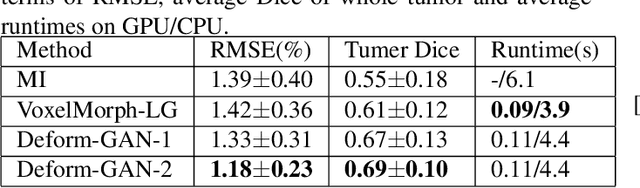
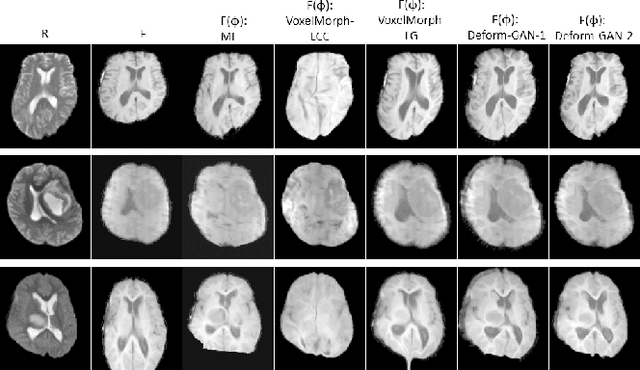
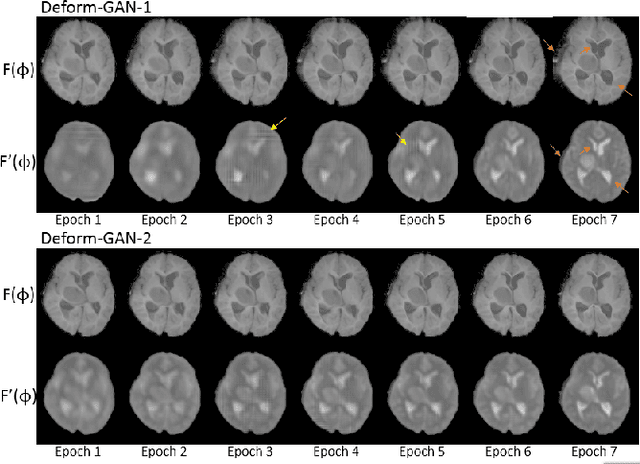
Deformable registration is one of the most challenging task in the field of medical image analysis, especially for the alignment between different sequences and modalities. In this paper, a non-rigid registration method is proposed for 3D medical images leveraging unsupervised learning. To the best of our knowledge, this is the first attempt to introduce gradient loss into deep-learning-based registration. The proposed gradient loss is robust across sequences and modals for large deformation. Besides, adversarial learning approach is used to transfer multi-modal similarity to mono-modal similarity and improve the precision. Neither ground-truth nor manual labeling is required during training. We evaluated our network on a 3D brain registration task comprehensively. The experiments demonstrate that the proposed method can cope with the data which has non-functional intensity relations, noise and blur. Our approach outperforms other methods especially in accuracy and speed.
Atlas: End-to-End 3D Scene Reconstruction from Posed Images
Mar 23, 2020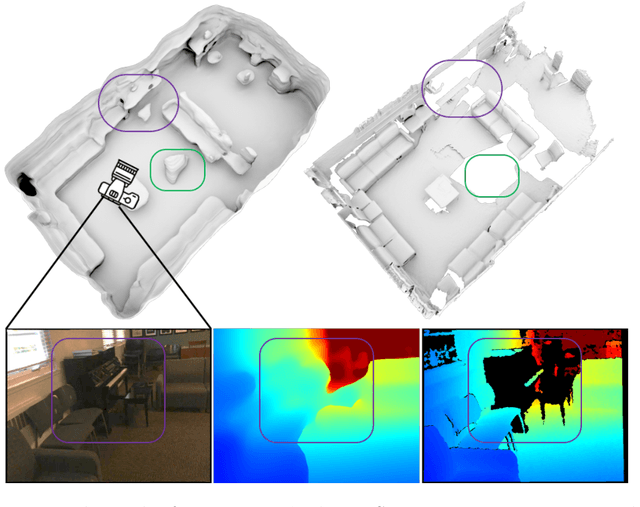
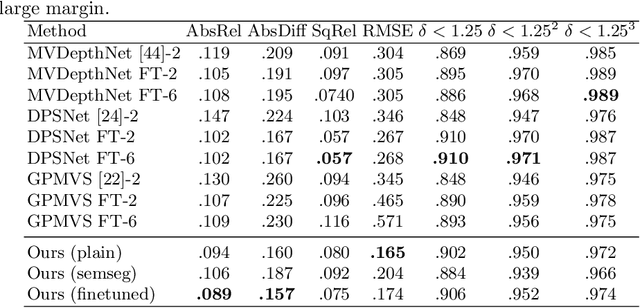
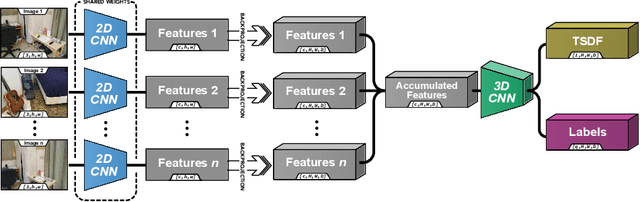
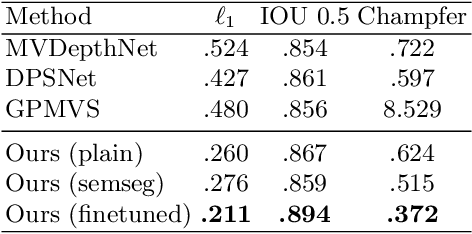
We present an end-to-end 3D reconstruction method for a scene by directly regressing a truncated signed distance function (TSDF) from a set of posed RGB images. Traditional approaches to 3D reconstruction rely on an intermediate representation of depth maps prior to estimating a full 3D model of a scene. We hypothesize that a direct regression to 3D is more effective. A 2D CNN extracts features from each image independently which are then back-projected and accumulated into a voxel volume using the camera intrinsics and extrinsics. After accumulation, a 3D CNN refines the accumulated features and predicts the TSDF values. Additionally, semantic segmentation of the 3D model is obtained without significant computation. This approach is evaluated on the Scannet dataset where we significantly outperform state-of-the-art baselines (deep multiview stereo followed by traditional TSDF fusion) both quantitatively and qualitatively. We compare our 3D semantic segmentation to prior methods that use a depth sensor since no previous work attempts the problem with only RGB input.
P2ExNet: Patch-based Prototype Explanation Network
May 05, 2020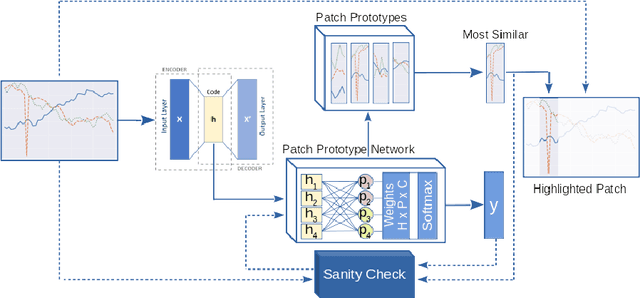
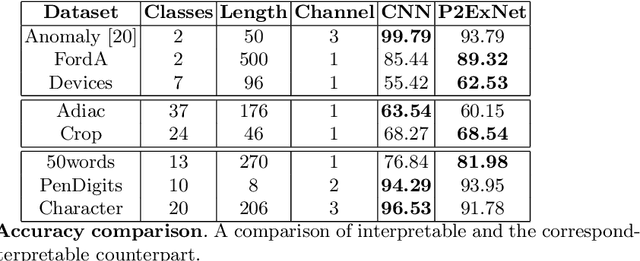
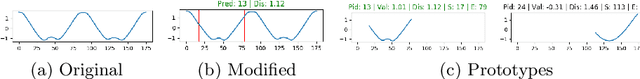
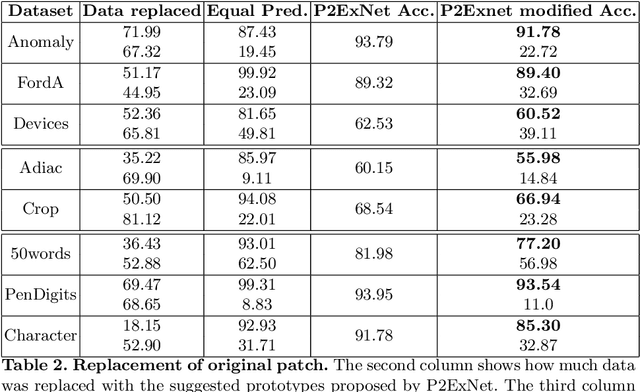
Deep learning methods have shown great success in several domains as they process a large amount of data efficiently, capable of solving complex classification, forecast, segmentation, and other tasks. However, they come with the inherent drawback of inexplicability limiting their applicability and trustworthiness. Although there exists work addressing this perspective, most of the existing approaches are limited to the image modality due to the intuitive and prominent concepts. Conversely, the concepts in the time-series domain are more complex and non-comprehensive but these and an explanation for the network decision are pivotal in critical domains like medical, financial, or industry. Addressing the need for an explainable approach, we propose a novel interpretable network scheme, designed to inherently use an explainable reasoning process inspired by the human cognition without the need of additional post-hoc explainability methods. Therefore, class-specific patches are used as they cover local concepts relevant to the classification to reveal similarities with samples of the same class. In addition, we introduce a novel loss concerning interpretability and accuracy that constraints P2ExNet to provide viable explanations of the data including relevant patches, their position, class similarities, and comparison methods without compromising accuracy. Analysis of the results on eight publicly available time-series datasets reveals that P2ExNet reaches comparable performance when compared to its counterparts while inherently providing understandable and traceable decisions.
Feature Augmentation Improves Anomalous Change Detection for Human Activity Identification in Synthetic Aperture Radar Imagery
Dec 07, 2019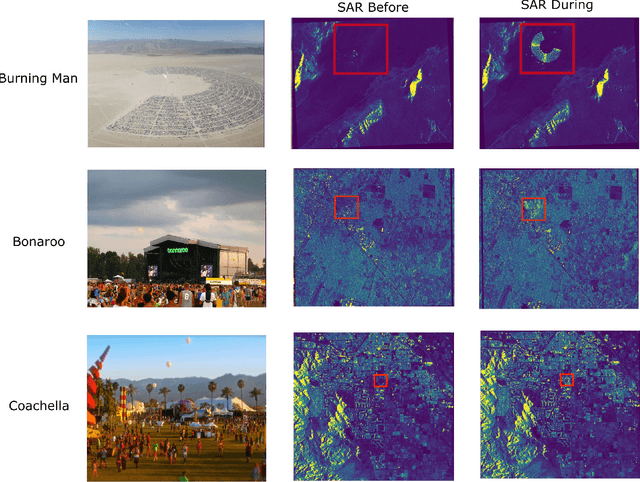
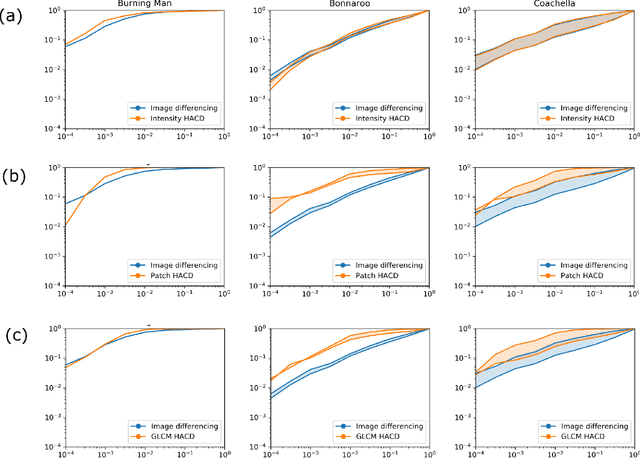
Anomalous change detection (ACD) methods separate common, uninteresting changes from rare, significant changes in co-registered images collected at different points in time. In this paper we evaluate methods to improve the performance of ACD in detecting human activity in SAR imagery using outdoor music festivals as a target. Our results show that the low dimensionality of SAR data leads to poor performance of ACD when compared to simpler methods such as image differencing, but augmenting the dimensionality of our input feature space by incorporating local spatial information leads to enhanced performance.
New version of Gram-Schmidt Process with inverse for Signal and Image Processing
Jul 16, 2016

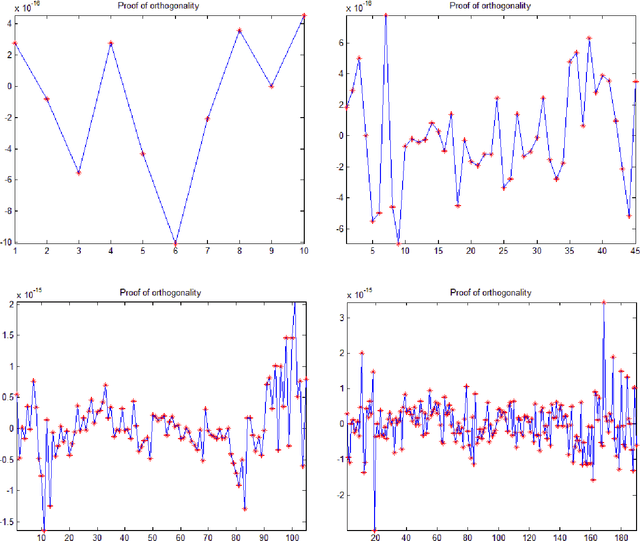
The Gram-Schmidt Process (GSP) is used to convert a non-orthogonal basis (a set of linearly independent vectors, matrices, etc) into an orthonormal basis (a set of orthogonal, unit-length vectors, bi or tri dimensional matrices). The process consists of taking each array and then subtracting the projections in common with the previous arrays. This paper introduces an enhanced version of the Gram-Schmidt Process (EGSP) with inverse, which is useful for Digital Signal and Image Processing, among others applications.
AlignShift: Bridging the Gap of Imaging Thickness in 3D Anisotropic Volumes
May 05, 2020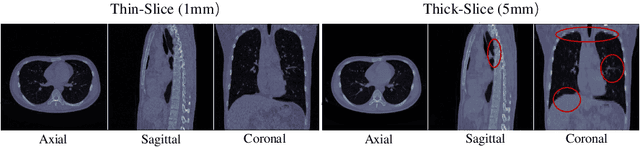
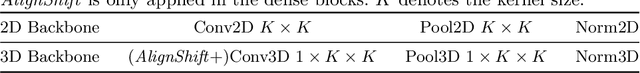
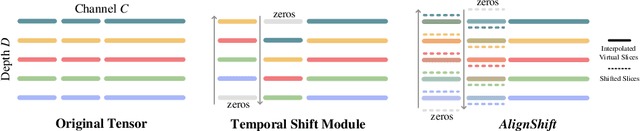
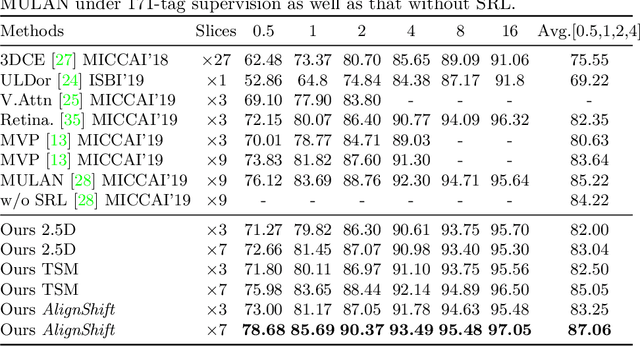
This paper addresses a fundamental challenge in 3D medical image processing: how to deal with imaging thickness. For anisotropic medical volumes, there is a significant performance gap between thin-slice (mostly 1mm) and thick-slice (mostly 5mm) volumes. Prior arts tend to use 3D approaches for the thin-slice and 2D approaches for the thick-slice, respectively. We aim at a unified approach for both thin- and thick-slice medical volumes. Inspired by recent advances in video analysis, we propose AlignShift, a novel parameter-free operator to convert theoretically any 2D pretrained network into thickness-aware 3D network. Remarkably, the converted networks behave like 3D for the thin-slice, nevertheless degenerate to 2D for the thick-slice adaptively. The unified thickness-aware representation learning is achieved by shifting and fusing aligned "virtual slices" as per the input imaging thickness. Extensive experiments on public large-scale DeepLesion benchmark, consisting of 32K lesions for universal lesion detection, validate the effectiveness of our method, which outperforms previous state of the art by considerable margins, without whistles and bells. More importantly, to our knowledge, this is the first method that bridges the performance gap between thin- and thick-slice volumes by a unified framework. To improve research reproducibility, our code in PyTorch is open source at https://github.com/M3DV/AlignShift.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge