"Image": models, code, and papers
Explaining Motion Relevance for Activity Recognition in Video Deep Learning Models
Mar 31, 2020
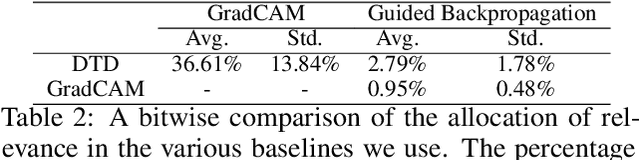
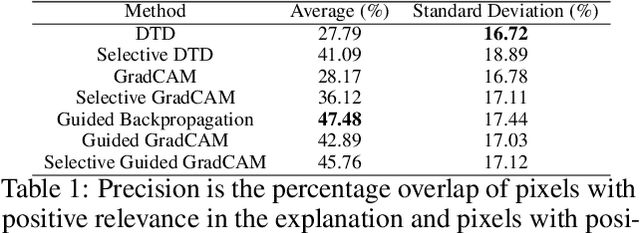
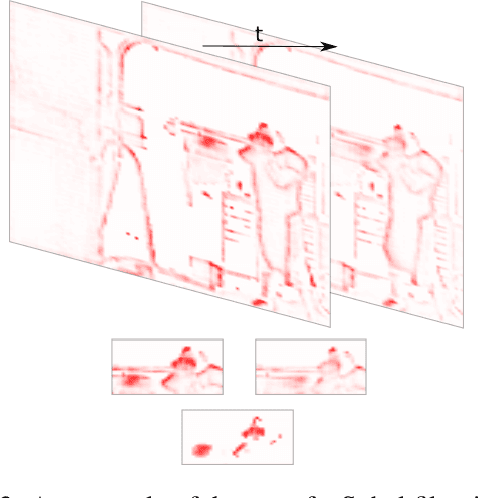

A small subset of explainability techniques developed initially for image recognition models has recently been applied for interpretability of 3D Convolutional Neural Network models in activity recognition tasks. Much like the models themselves, the techniques require little or no modification to be compatible with 3D inputs. However, these explanation techniques regard spatial and temporal information jointly. Therefore, using such explanation techniques, a user cannot explicitly distinguish the role of motion in a 3D model's decision. In fact, it has been shown that these models do not appropriately factor motion information into their decision. We propose a selective relevance method for adapting the 2D explanation techniques to provide motion-specific explanations, better aligning them with the human understanding of motion as conceptually separate from static spatial features. We demonstrate the utility of our method in conjunction with several widely-used 2D explanation methods, and show that it improves explanation selectivity for motion. Our results show that the selective relevance method can not only provide insight on the role played by motion in the model's decision -- in effect, revealing and quantifying the model's spatial bias -- but the method also simplifies the resulting explanations for human consumption.
Embedding Human Heuristics in Machine-Learning-Enabled Probe Microscopy
Jul 31, 2019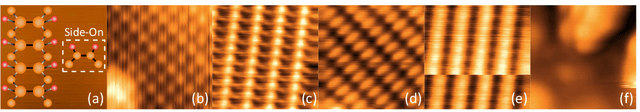
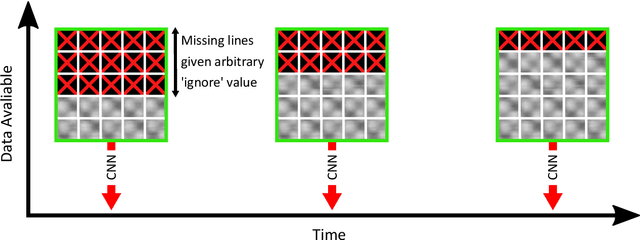
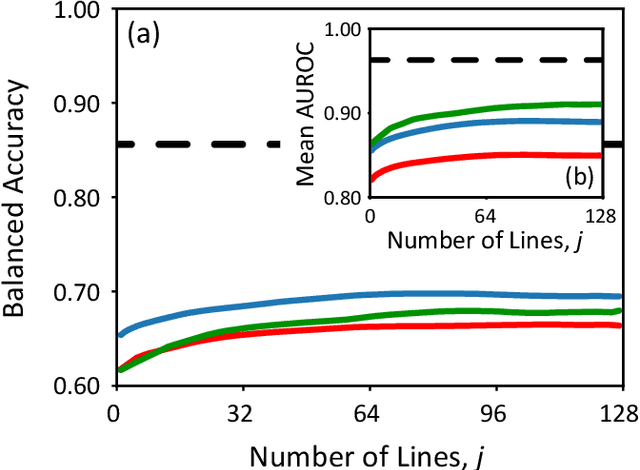
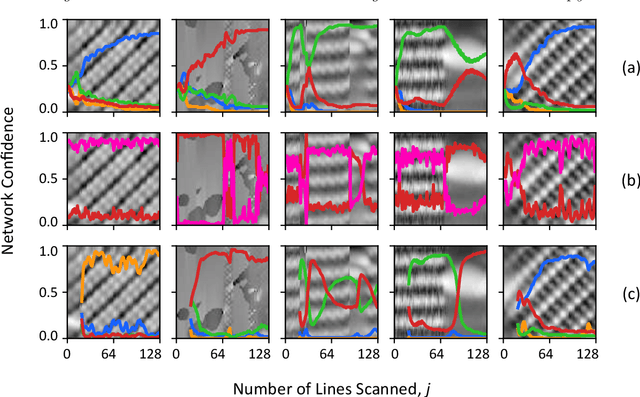
Scanning probe microscopists generally do not rely on complete images to assess the quality of data acquired during a scan. Instead, assessments of the state of the tip apex, which not only determines the resolution in any scanning probe technique but can also generate a wide array of frustrating artefacts, are carried out in real time on the basis of a few lines of an image (and, typically, their associated line profiles.) The very small number of machine learning approaches to probe microscopy published to date, however, involve classifications based on full images. Given that data acquisition is the most time-consuming task during routine tip conditioning, automated methods are thus currently extremely slow in comparison to the tried-and-trusted strategies and heuristics used routinely by probe microscopists. Here, we explore various strategies by which different STM image classes (arising from changes in the tip state) can be correctly identified from partial scans. By employing a secondary temporal network and a rolling window of a small group of individual scanlines, we find that tip assessment is possible with a small fraction of a complete image. We achieve this with little-to-no performance penalty -- or, indeed, markedly improved performance in some cases -- and introduce a protocol to detect the state of the tip apex in real time.
Inverting Gradients -- How easy is it to break privacy in federated learning?
Mar 31, 2020
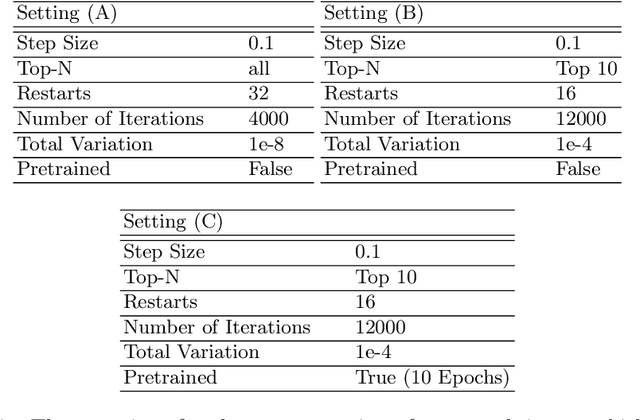
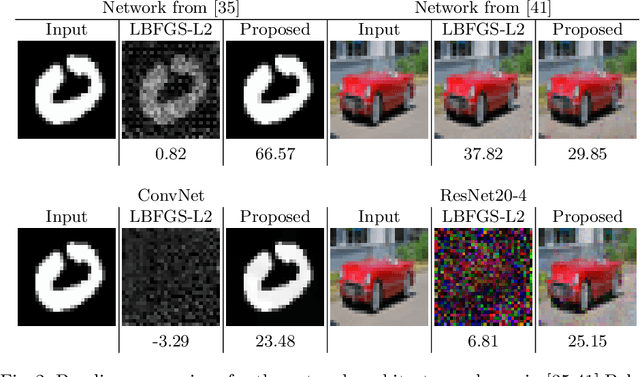

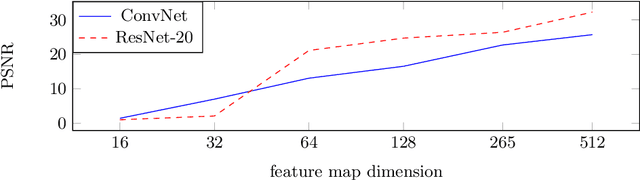
The idea of federated learning is to collaboratively train a neural network on a server. Each user receives the current weights of the network and in turns sends parameter updates (gradients) based on local data. This protocol has been designed not only to train neural networks data-efficiently, but also to provide privacy benefits for users, as their input data remains on device and only parameter gradients are shared. In this paper we show that sharing parameter gradients is by no means secure: By exploiting a cosine similarity loss along with optimization methods from adversarial attacks, we are able to faithfully reconstruct images at high resolution from the knowledge of their parameter gradients, and demonstrate that such a break of privacy is possible even for trained deep networks. Moreover, we analyze the effects of architecture as well as parameters on the difficulty of reconstructing the input image, prove that any input to a fully connected layer can be reconstructed analytically independent of the remaining architecture, and show numerically that even averaging gradients over several iterations or several images does not protect the user's privacy in federated learning applications in computer vision.
UI-Net: Interactive Artificial Neural Networks for Iterative Image Segmentation Based on a User Model
Sep 11, 2017
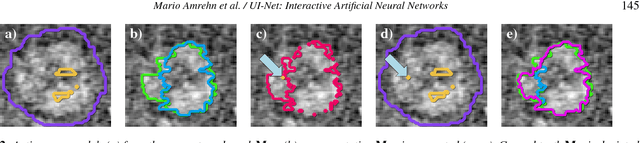
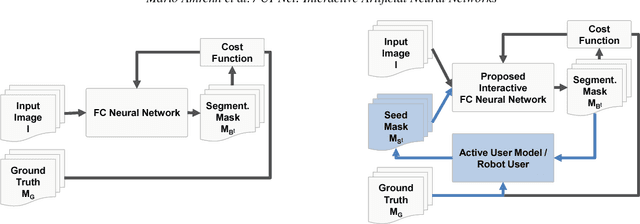
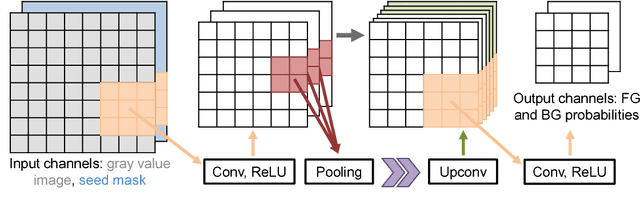
For complex segmentation tasks, fully automatic systems are inherently limited in their achievable accuracy for extracting relevant objects. Especially in cases where only few data sets need to be processed for a highly accurate result, semi-automatic segmentation techniques exhibit a clear benefit for the user. One area of application is medical image processing during an intervention for a single patient. We propose a learning-based cooperative segmentation approach which includes the computing entity as well as the user into the task. Our system builds upon a state-of-the-art fully convolutional artificial neural network (FCN) as well as an active user model for training. During the segmentation process, a user of the trained system can iteratively add additional hints in form of pictorial scribbles as seed points into the FCN system to achieve an interactive and precise segmentation result. The segmentation quality of interactive FCNs is evaluated. Iterative FCN approaches can yield superior results compared to networks without the user input channel component, due to a consistent improvement in segmentation quality after each interaction.
* This work is submitted to the 2017 Eurographics Workshop on Visual Computing for Biology and Medicine
Effective Learning of a GMRF Mixture Model
May 20, 2020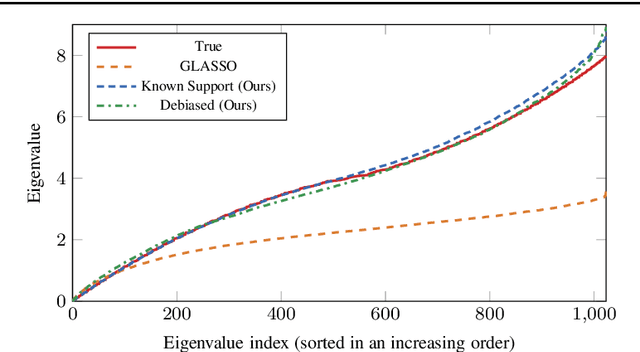
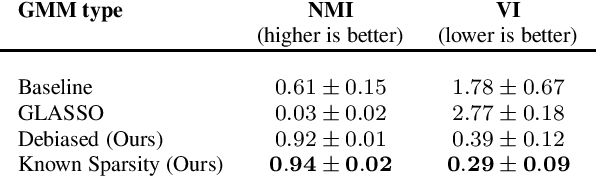

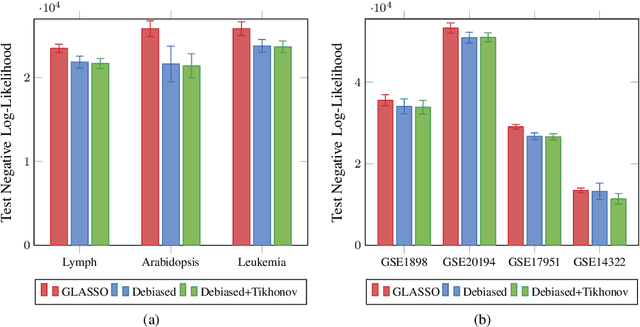
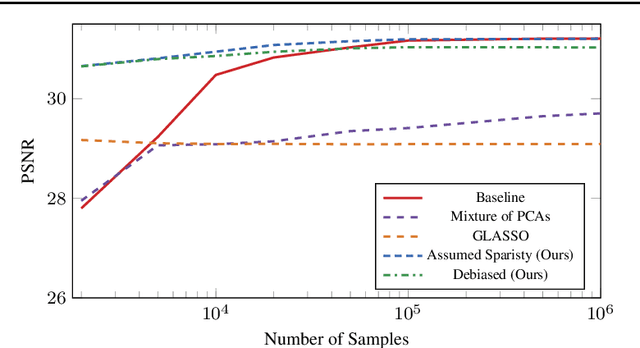
Learning a Gaussian Mixture Model (GMM) is hard when the number of parameters is too large given the amount of available data. As a remedy, we propose restricting the GMM to a Gaussian Markov Random Field Mixture Model (GMRF-MM), as well as a new method for estimating the latter's sparse precision (i.e., inverse covariance) matrices. When the sparsity pattern of each matrix is known, we propose an efficient optimization method for the Maximum Likelihood Estimate (MLE) of that matrix. When it is unknown, we utilize the popular Graphical LASSO (GLASSO) to estimate that pattern. However, we show that even for a single Gaussian, when GLASSO is tuned to successfully estimate the sparsity pattern, it does so at the price of a substantial bias of the values of the nonzero entries of the matrix, and we show that this problem only worsens in a mixture setting. To overcome this, we discard the non-zero values estimated by GLASSO, keep only its pattern estimate and use it within the proposed MLE method. This yields an effective two-step procedure that removes the bias. We show that our "debiasing" approach outperforms GLASSO in both the single-GMRF and the GMRF-MM cases. We also show that when learning priors for image patches, our method outperforms GLASSO even if we merely use an educated guess about the sparsity pattern, and that our GMRF-MM outperforms the baseline GMM on real and synthetic high-dimensional datasets. Our code is available at \url{https://github.com/shahaffind/GMRF-MM}.
Convolutional Neural Networks with Intermediate Loss for 3D Super-Resolution of CT and MRI Scans
Jan 05, 2020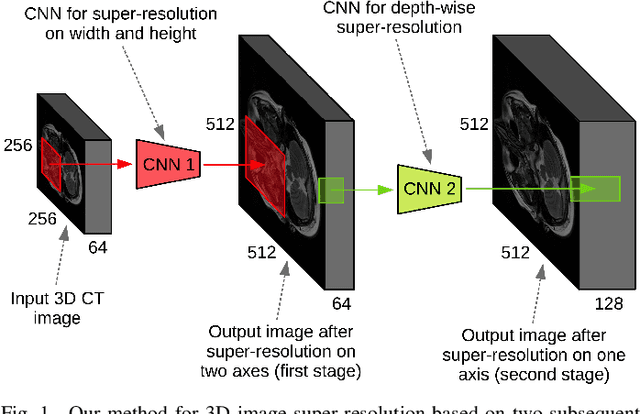
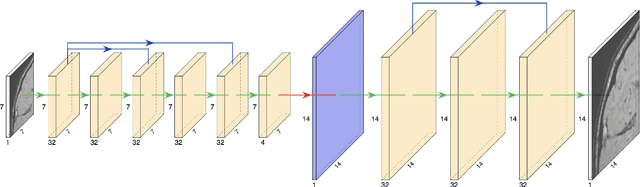
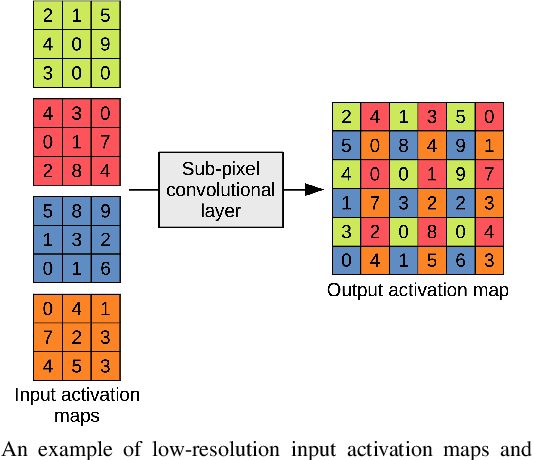
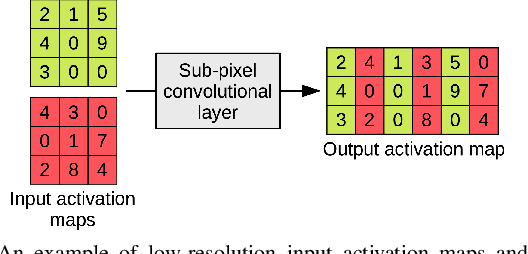
CT scanners that are commonly-used in hospitals nowadays produce low-resolution images, up to 512 pixels in size. One pixel in the image corresponds to a one millimeter piece of tissue. In order to accurately segment tumors and make treatment plans, doctors need CT scans of higher resolution. The same problem appears in MRI. In this paper, we propose an approach for the single-image super-resolution of 3D CT or MRI scans. Our method is based on deep convolutional neural networks (CNNs) composed of 10 convolutional layers and an intermediate upscaling layer that is placed after the first 6 convolutional layers. Our first CNN, which increases the resolution on two axes (width and height), is followed by a second CNN, which increases the resolution on the third axis (depth). Different from other methods, we compute the loss with respect to the ground-truth high-resolution output right after the upscaling layer, in addition to computing the loss after the last convolutional layer. The intermediate loss forces our network to produce a better output, closer to the ground-truth. A widely-used approach to obtain sharp results is to add Gaussian blur using a fixed standard deviation. In order to avoid overfitting to a fixed standard deviation, we apply Gaussian smoothing with various standard deviations, unlike other approaches. We evaluate our method in the context of 2D and 3D super-resolution of CT and MRI scans from two databases, comparing it to relevant related works from the literature and baselines based on various interpolation schemes, using 2x and 4x scaling factors. The empirical results show that our approach attains superior results to all other methods. Moreover, our human annotation study reveals that both doctors and regular annotators chose our method in favor of Lanczos interpolation in 97.55% cases for 2x upscaling factor and in 96.69% cases for 4x upscaling factor.
Using the quantization error from Self-Organized Map (SOM) output for detecting critical variability in large bodies of image time series in less than a minute
Oct 29, 2017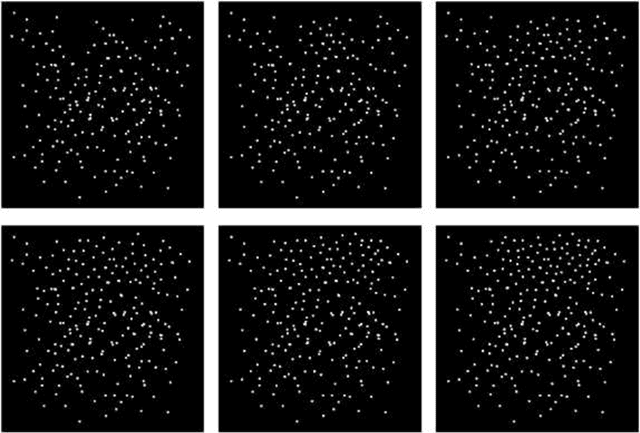
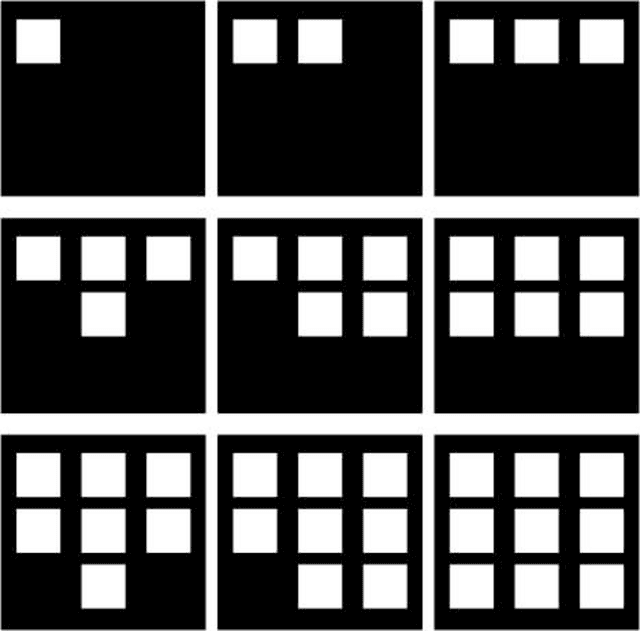
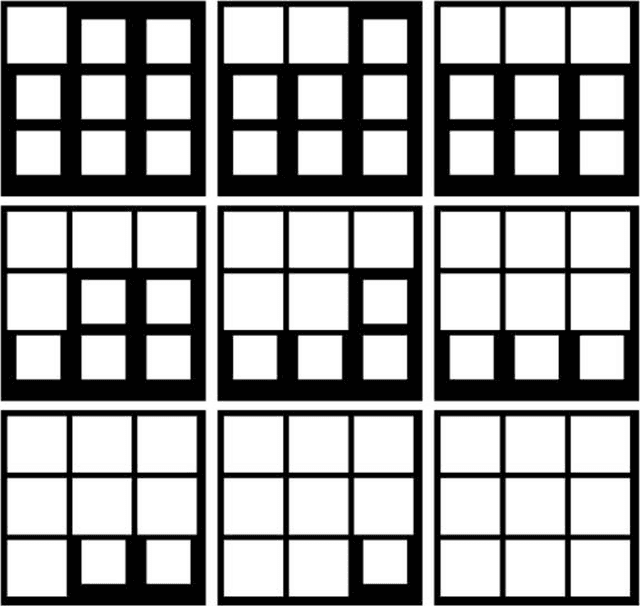



The quantization error (QE) from SOM applied on time series of spatial contrast images with variable relative amount of white and dark pixel contents, as in monochromatic medical images or satellite images, is proven a reliable indicator of potentially critical changes in image homogeneity. The QE is shown to increase linearly with the variability in spatial contrast contents across time when contrast intensity is kept constant.
The Mertens Unrolled Network (MU-Net): A High Dynamic Range Fusion Neural Network for Through the Windshield Driver Recognition
Feb 27, 2020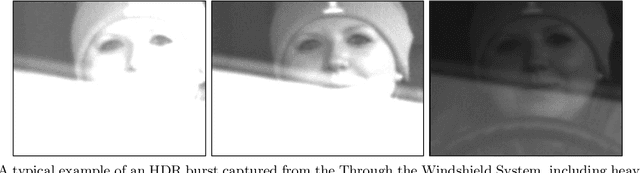
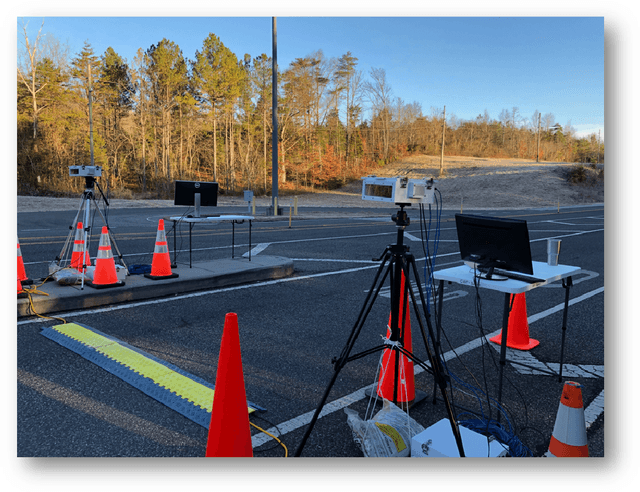
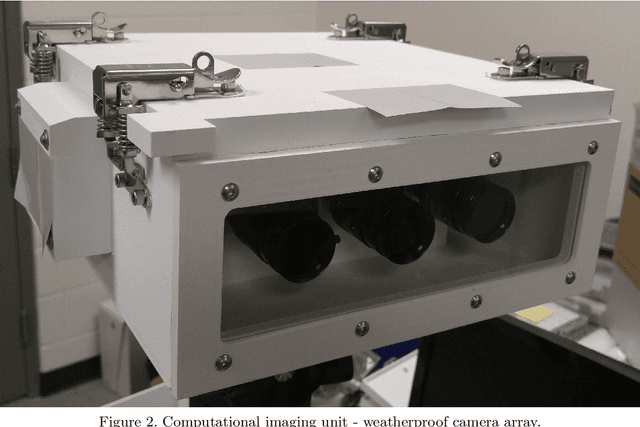
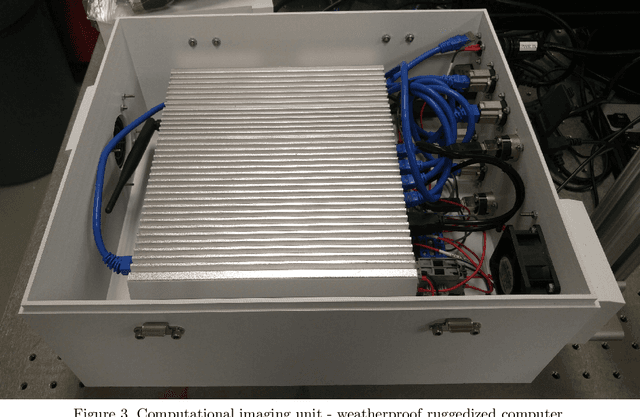



Face recognition of vehicle occupants through windshields in unconstrained environments poses a number of unique challenges ranging from glare, poor illumination, driver pose and motion blur. In this paper, we further develop the hardware and software components of a custom vehicle imaging system to better overcome these challenges. After the build out of a physical prototype system that performs High Dynamic Range (HDR) imaging, we collect a small dataset of through-windshield image captures of known drivers. We then re-formulate the classical Mertens-Kautz-Van Reeth HDR fusion algorithm as a pre-initialized neural network, which we name the Mertens Unrolled Network (MU-Net), for the purpose of fine-tuning the HDR output of through-windshield images. Reconstructed faces from this novel HDR method are then evaluated and compared against other traditional and experimental HDR methods in a pre-trained state-of-the-art (SOTA) facial recognition pipeline, verifying the efficacy of our approach.
Simple Multi-Resolution Representation Learning for Human Pose Estimation
Apr 14, 2020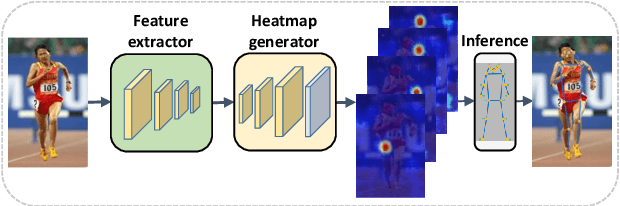
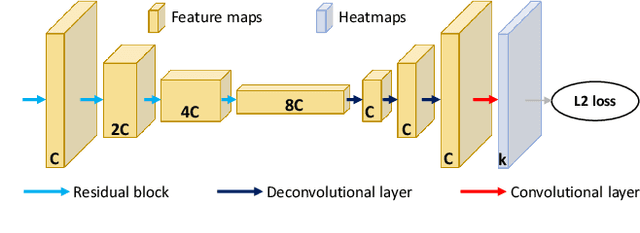
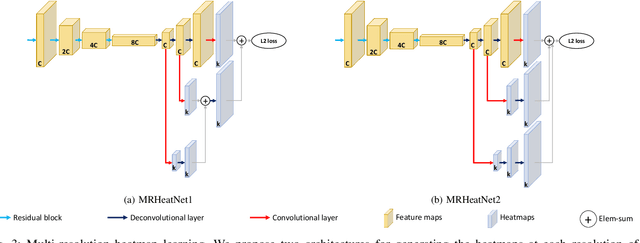
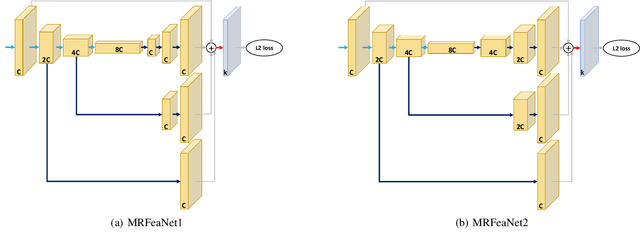
Human pose estimation - the process of recognizing human keypoints in a given image - is one of the most important tasks in computer vision and has a wide range of applications including movement diagnostics, surveillance, or self-driving vehicle. The accuracy of human keypoint prediction is increasingly improved thanks to the burgeoning development of deep learning. Most existing methods solved human pose estimation by generating heatmaps in which the ith heatmap indicates the location confidence of the ith keypoint. In this paper, we introduce novel network structures referred to as multiresolution representation learning for human keypoint prediction. At different resolutions in the learning process, our networks branch off and use extra layers to learn heatmap generation. We firstly consider the architectures for generating the multiresolution heatmaps after obtaining the lowest-resolution feature maps. Our second approach allows learning during the process of feature extraction in which the heatmaps are generated at each resolution of the feature extractor. The first and second approaches are referred to as multi-resolution heatmap learning and multi-resolution feature map learning respectively. Our architectures are simple yet effective, achieving good performance. We conducted experiments on two common benchmarks for human pose estimation: MS-COCO and MPII dataset.
Deep Learning Models for Digital Pathology
Oct 29, 2019
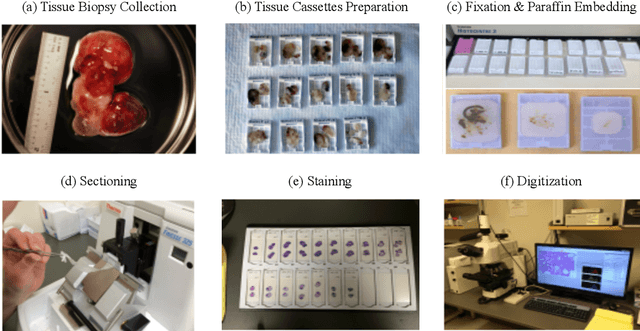

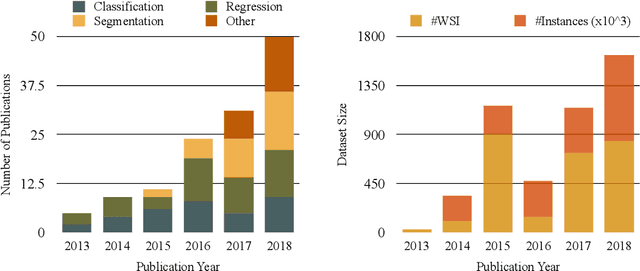
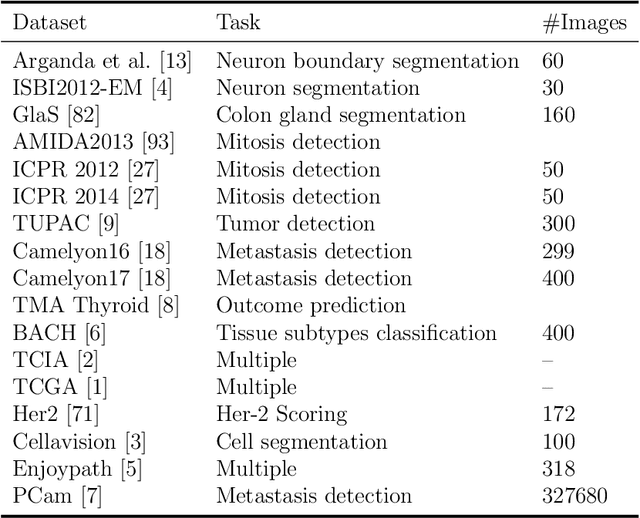
Histopathology images; microscopy images of stained tissue biopsies contain fundamental prognostic information that forms the foundation of pathological analysis and diagnostic medicine. However, diagnostics from histopathology images generally rely on a visual cognitive assessment of tissue slides which implies an inherent element of interpretation and hence subjectivity. Access to digitized histopathology images enabled the development of computational systems aiming at reducing manual intervention and automating parts of pathologists' workflow. Specifically, applications of deep learning to histopathology image analysis now offer opportunities for better quantitative modeling of disease appearance and hence possibly improved prediction of disease aggressiveness and patient outcome. However digitized histopathology tissue slides are unique in a variety of ways and come with their own set of computational challenges. In this survey, we summarize the different challenges facing computational systems for digital pathology and provide a review of state-of-the-art works that developed deep learning-based solutions for the predictive modeling of histopathology images from a detection, stain normalization, segmentation, and tissue classification perspective. We then discuss the challenges facing the validation and integration of such deep learning-based computational systems in clinical workflow and reflect on future opportunities for histopathology derived image measurements and better predictive modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge