"Image": models, code, and papers
BiNet: Degraded-Manuscript Binarization in Diverse Document Textures and Layouts using Deep Encoder-Decoder Networks
Nov 13, 2019
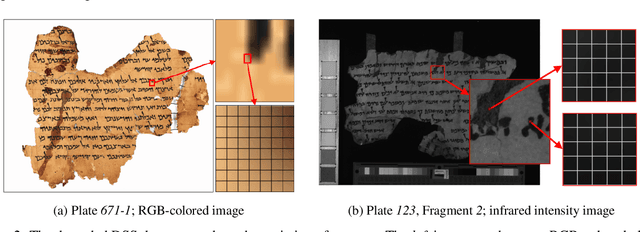
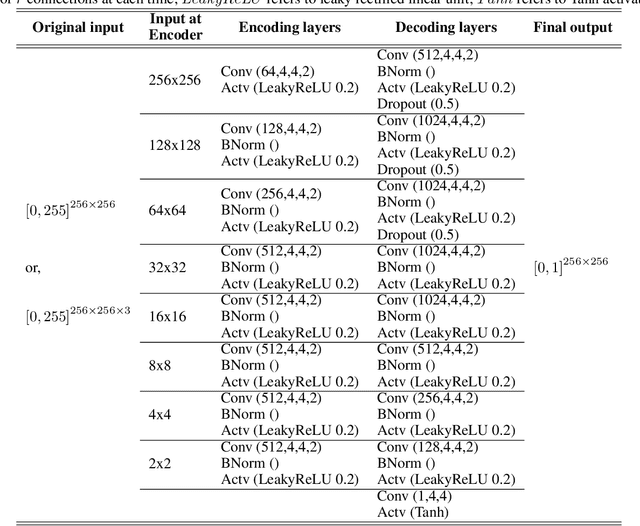

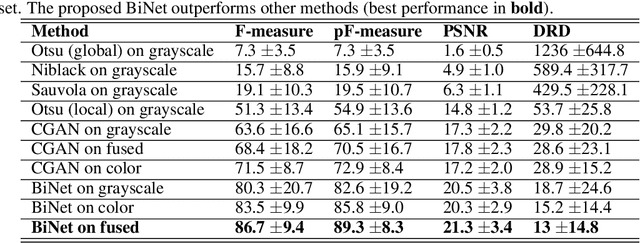
Handwritten document-image binarization is a semantic segmentation process to differentiate ink pixels from background pixels. It is one of the essential steps towards character recognition, writer identification, and script-style evolution analysis. The binarization task itself is challenging due to the vast diversity of writing styles, inks, and paper materials. It is even more difficult for historical manuscripts due to the aging and degradation of the documents over time. One of such manuscripts is the Dead Sea Scrolls (DSS) image collection, which poses extreme challenges for the existing binarization techniques. This article proposes a new binarization technique for the DSS images using the deep encoder-decoder networks. Although the artificial neural network proposed here is primarily designed to binarize the DSS images, it can be trained on different manuscript collections as well. Additionally, the use of transfer learning makes the network already utilizable for a wide range of handwritten documents, making it a unique multi-purpose tool for binarization. Qualitative results and several quantitative comparisons using both historical manuscripts and datasets from handwritten document image binarization competition (H-DIBCO and DIBCO) exhibit the robustness and the effectiveness of the system. The best performing network architecture proposed here is a variant of the U-Net encoder-decoders.
Visualization for Histopathology Images using Graph Convolutional Neural Networks
Jun 16, 2020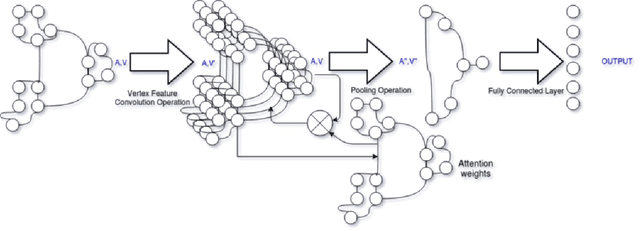
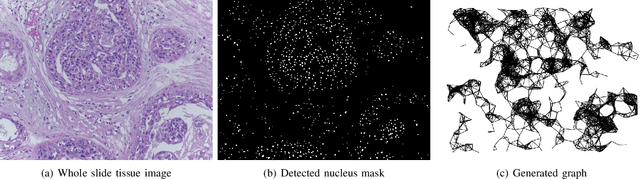
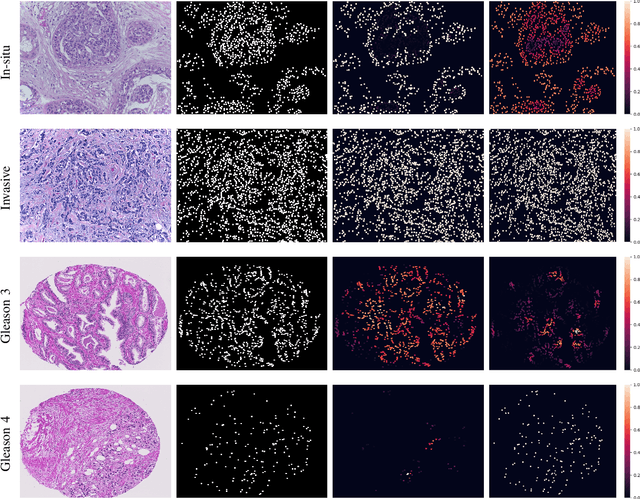
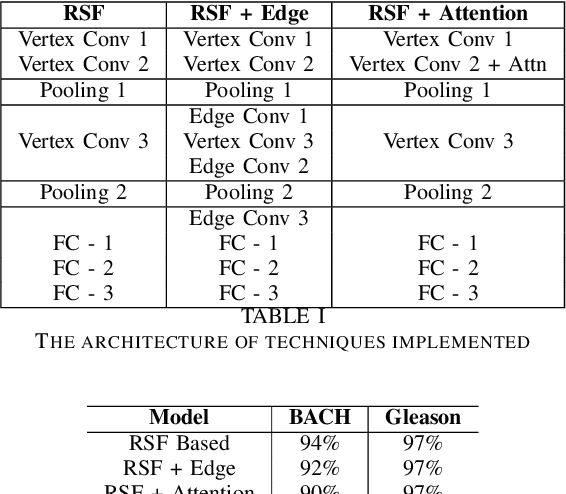
With the increase in the use of deep learning for computer-aided diagnosis in medical images, the criticism of the black-box nature of the deep learning models is also on the rise. The medical community needs interpretable models for both due diligence and advancing the understanding of disease and treatment mechanisms. In histology, in particular, while there is rich detail available at the cellular level and that of spatial relationships between cells, it is difficult to modify convolutional neural networks to point out the relevant visual features. We adopt an approach to model histology tissue as a graph of nuclei and develop a graph convolutional network framework based on attention mechanism and node occlusion for disease diagnosis. The proposed method highlights the relative contribution of each cell nucleus in the whole-slide image. Our visualization of such networks trained to distinguish between invasive and in-situ breast cancers, and Gleason 3 and 4 prostate cancers generate interpretable visual maps that correspond well with our understanding of the structures that are important to experts for their diagnosis.
ManifoldNorm: Extending normalizations on Riemannian Manifolds
Apr 04, 2020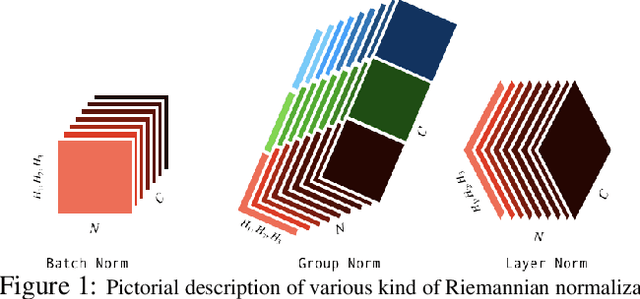
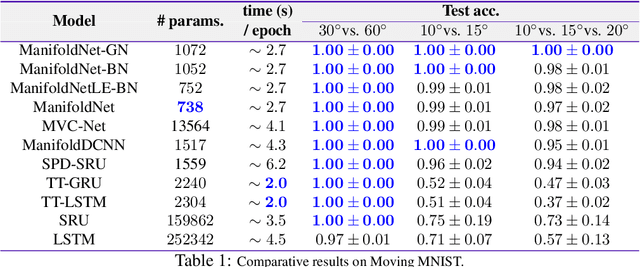
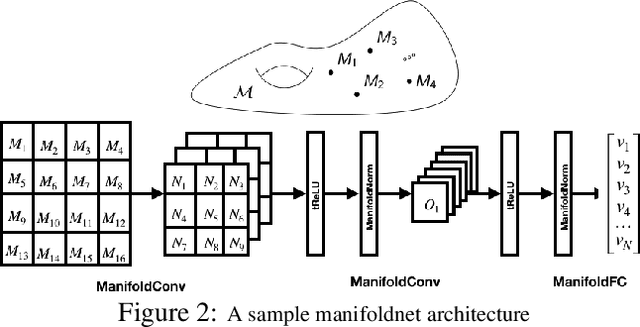
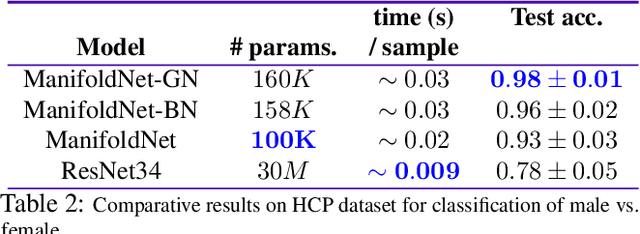
Many measurements in computer vision and machine learning manifest as non-Euclidean data samples. Several researchers recently extended a number of deep neural network architectures for manifold valued data samples. Researchers have proposed models for manifold valued spatial data which are common in medical image processing including processing of diffusion tensor imaging (DTI) where images are fields of $3\times 3$ symmetric positive definite matrices or representation in terms of orientation distribution field (ODF) where the identification is in terms of field on hypersphere. There are other sequential models for manifold valued data that recently researchers have shown to be effective for group difference analysis in study for neuro-degenerative diseases. Although, several of these methods are effective to deal with manifold valued data, the bottleneck includes the instability in optimization for deeper networks. In order to deal with these instabilities, researchers have proposed residual connections for manifold valued data. One of the other remedies to deal with the instabilities including gradient explosion is to use normalization techniques including {\it batch norm} and {\it group norm} etc.. But, so far there is no normalization techniques applicable for manifold valued data. In this work, we propose a general normalization techniques for manifold valued data. We show that our proposed manifold normalization technique have special cases including popular batch norm and group norm techniques. On the experimental side, we focus on two types of manifold valued data including manifold of symmetric positive definite matrices and hypersphere. We show the performance gain in one synthetic experiment for moving MNIST dataset and one real brain image dataset where the representation is in terms of orientation distribution field (ODF).
Adversarial Graph Representation Adaptation for Cross-Domain Facial Expression Recognition
Aug 04, 2020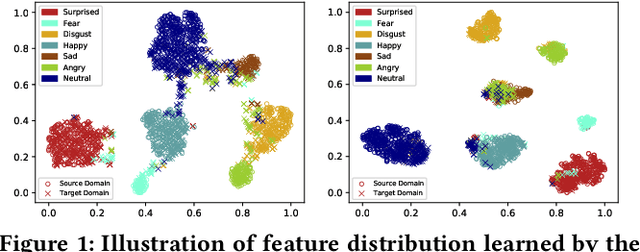
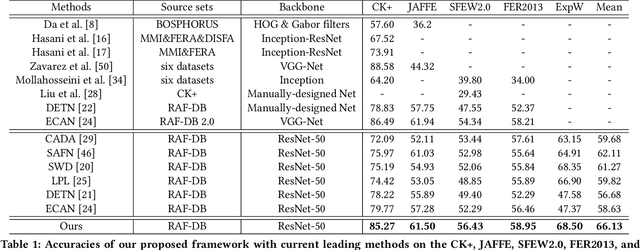
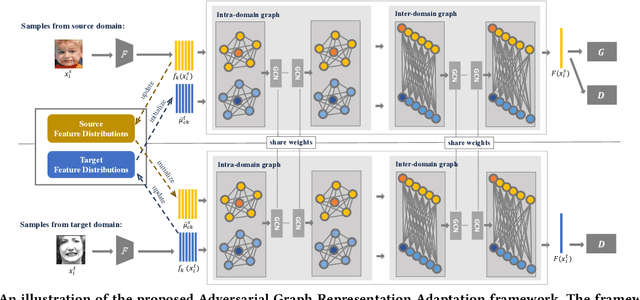
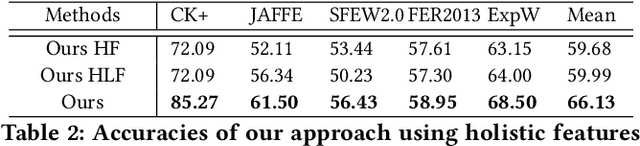
Data inconsistency and bias are inevitable among different facial expression recognition (FER) datasets due to subjective annotating process and different collecting conditions. Recent works resort to adversarial mechanisms that learn domain-invariant features to mitigate domain shift. However, most of these works focus on holistic feature adaptation, and they ignore local features that are more transferable across different datasets. Moreover, local features carry more detailed and discriminative content for expression recognition, and thus integrating local features may enable fine-grained adaptation. In this work, we propose a novel Adversarial Graph Representation Adaptation (AGRA) framework that unifies graph representation propagation with adversarial learning for cross-domain holistic-local feature co-adaptation. To achieve this, we first build a graph to correlate holistic and local regions within each domain and another graph to correlate these regions across different domains. Then, we learn the per-class statistical distribution of each domain and extract holistic-local features from the input image to initialize the corresponding graph nodes. Finally, we introduce two stacked graph convolution networks to propagate holistic-local feature within each domain to explore their interaction and across different domains for holistic-local feature co-adaptation. In this way, the AGRA framework can adaptively learn fine-grained domain-invariant features and thus facilitate cross-domain expression recognition. We conduct extensive and fair experiments on several popular benchmarks and show that the proposed AGRA framework achieves superior performance over previous state-of-the-art methods.
Real-World Multi-Domain Data Applications for Generalizations to Clinical Settings
Jul 24, 2020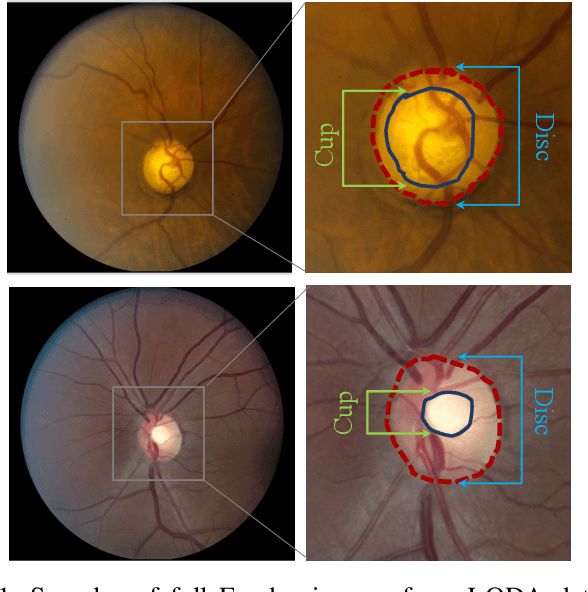
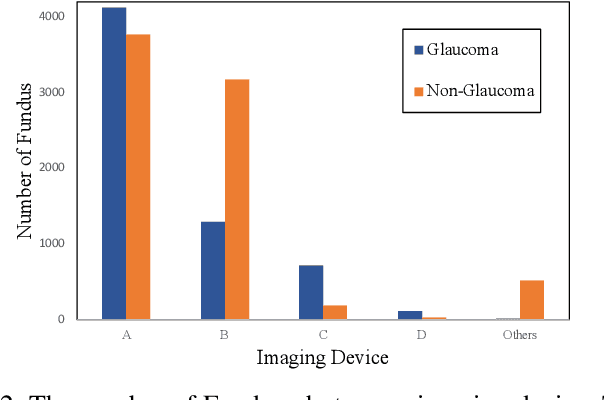
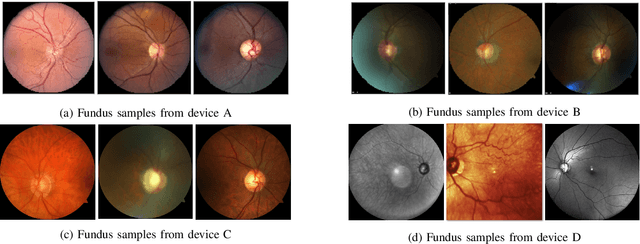
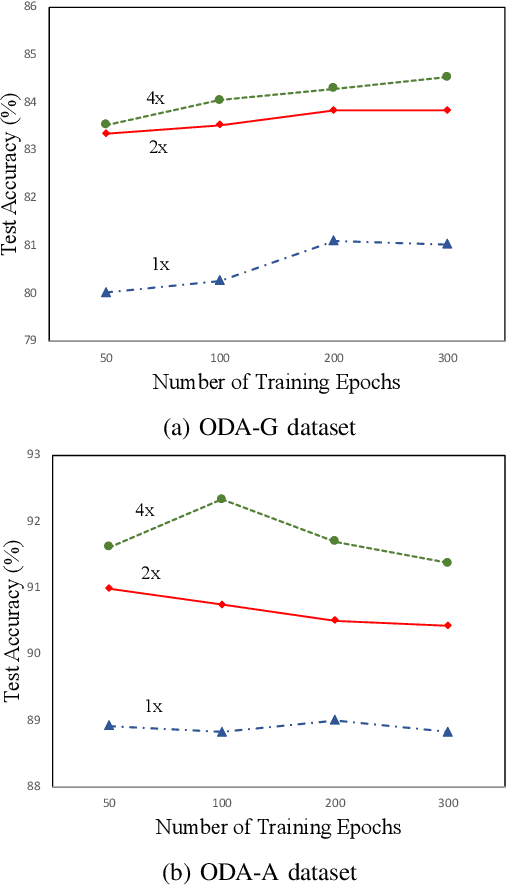
With promising results of machine learning based models in computer vision, applications on medical imaging data have been increasing exponentially. However, generalizations to complex real-world clinical data is a persistent problem. Deep learning models perform well when trained on standardized datasets from artificial settings, such as clinical trials. However, real-world data is different and translations are yielding varying results. The complexity of real-world applications in healthcare could emanate from a mixture of different data distributions across multiple device domains alongside the inevitable noise sourced from varying image resolutions, human errors, and the lack of manual gradings. In addition, healthcare applications not only suffer from the scarcity of labeled data, but also face limited access to unlabeled data due to HIPAA regulations, patient privacy, ambiguity in data ownership, and challenges in collecting data from different sources. These limitations pose additional challenges to applying deep learning algorithms in healthcare and clinical translations. In this paper, we utilize self-supervised representation learning methods, formulated effectively in transfer learning settings, to address limited data availability. Our experiments verify the importance of diverse real-world data for generalization to clinical settings. We show that by employing a self-supervised approach with transfer learning on a multi-domain real-world dataset, we can achieve 16% relative improvement on a standardized dataset over supervised baselines.
Transfer Learning from Partial Annotations for Whole Brain Segmentation
Aug 28, 2019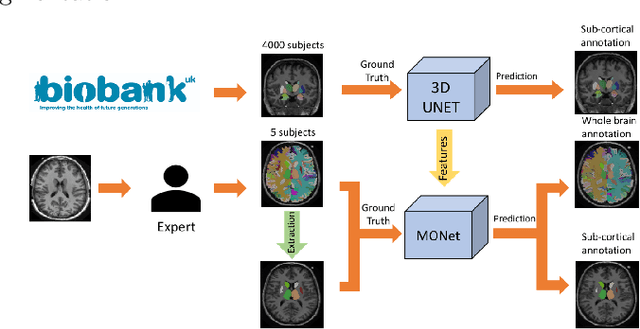

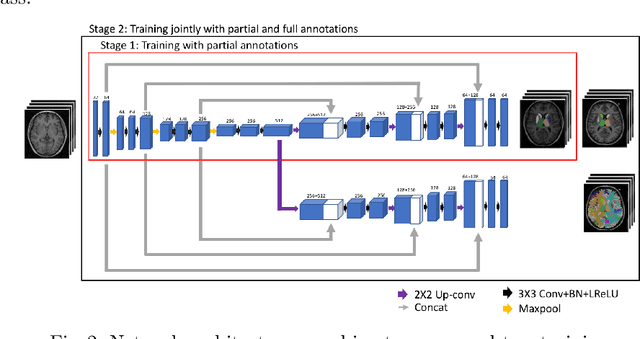
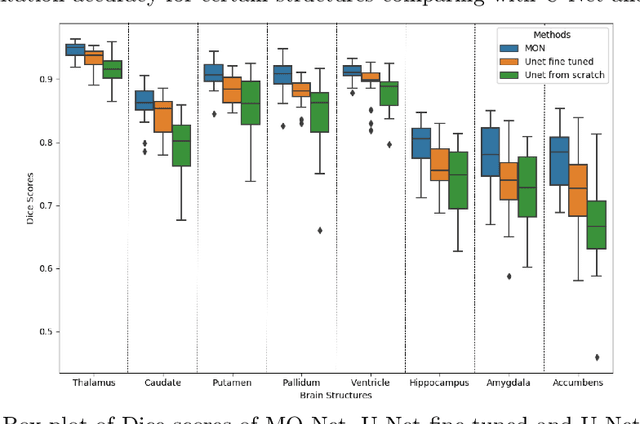
Brain MR image segmentation is a key task in neuroimaging studies. It is commonly conducted using standard computational tools, such as FSL, SPM, multi-atlas segmentation etc, which are often registration-based and suffer from expensive computation cost. Recently, there is an increased interest using deep neural networks for brain image segmentation, which have demonstrated advantages in both speed and performance. However, neural networks-based approaches normally require a large amount of manual annotations for optimising the massive amount of network parameters. For 3D networks used in volumetric image segmentation, this has become a particular challenge, as a 3D network consists of many more parameters compared to its 2D counterpart. Manual annotation of 3D brain images is extremely time-consuming and requires extensive involvement of trained experts. To address the challenge with limited manual annotations, here we propose a novel multi-task learning framework for brain image segmentation, which utilises a large amount of automatically generated partial annotations together with a small set of manually created full annotations for network training. Our method yields a high performance comparable to state-of-the-art methods for whole brain segmentation.
Dynamic Graph Correlation Learning for Disease Diagnosis with Incomplete Labels
Feb 28, 2020
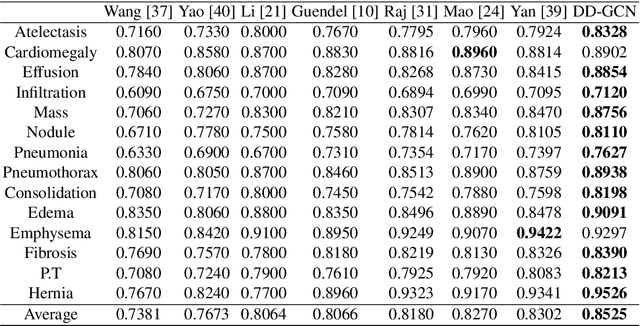
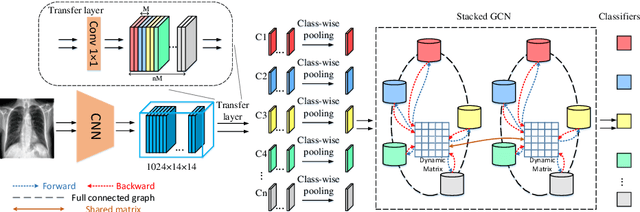
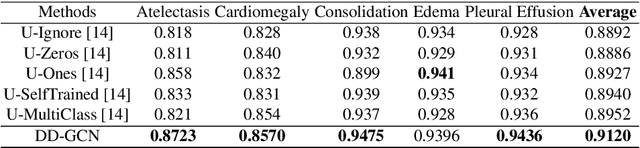
Disease diagnosis on chest X-ray images is a challenging multi-label classification task. Previous works generally classify the diseases independently on the input image without considering any correlation among diseases. However, such correlation actually exists, for example, Pleural Effusion is more likely to appear when Pneumothorax is present. In this work, we propose a Disease Diagnosis Graph Convolutional Network (DD-GCN) that presents a novel view of investigating the inter-dependency among different diseases by using a dynamic learnable adjacency matrix in graph structure to improve the diagnosis accuracy. To learn more natural and reliable correlation relationship, we feed each node with the image-level individual feature map corresponding to each type of disease. To our knowledge, our method is the first to build a graph over the feature maps with a dynamic adjacency matrix for correlation learning. To further deal with a practical issue of incomplete labels, DD-GCN also utilizes an adaptive loss and a curriculum learning strategy to train the model on incomplete labels. Experimental results on two popular chest X-ray (CXR) datasets show that our prediction accuracy outperforms state-of-the-arts, and the learned graph adjacency matrix establishes the correlation representations of different diseases, which is consistent with expert experience. In addition, we apply an ablation study to demonstrate the effectiveness of each component in DD-GCN.
Single Image Super-Resolution Using Multi-Scale Convolutional Neural Network
May 15, 2017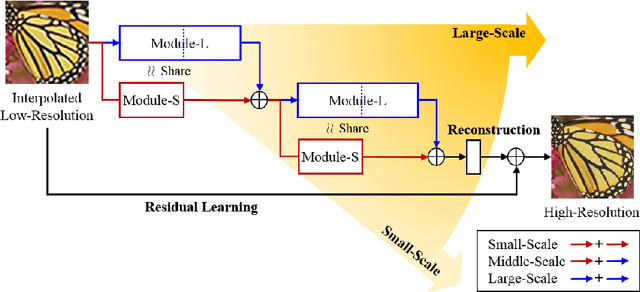


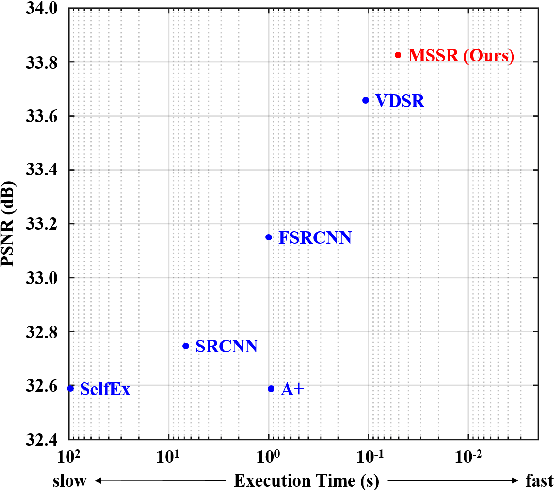
Methods based on convolutional neural network (CNN) have demonstrated tremendous improvements on single image super-resolution. However, the previous methods mainly restore images from one single area in the low resolution (LR) input, which limits the flexibility of models to infer various scales of details for high resolution (HR) output. Moreover, most of them train a specific model for each up-scale factor. In this paper, we propose a multi-scale super resolution (MSSR) network. Our network consists of multi-scale paths to make the HR inference, which can learn to synthesize features from different scales. This property helps reconstruct various kinds of regions in HR images. In addition, only one single model is needed for multiple up-scale factors, which is more efficient without loss of restoration quality. Experiments on four public datasets demonstrate that the proposed method achieved state-of-the-art performance with fast speed.
A Neural Dirichlet Process Mixture Model for Task-Free Continual Learning
Jan 14, 2020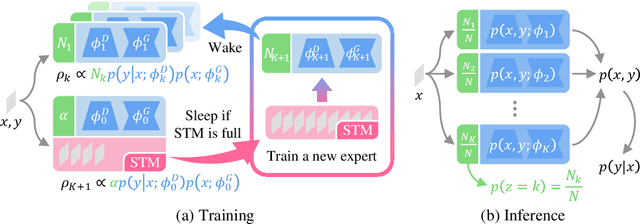
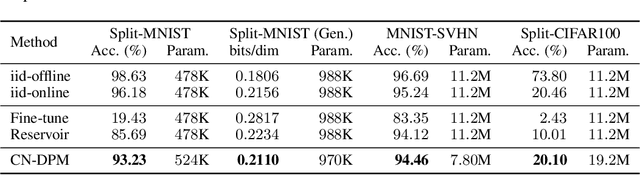
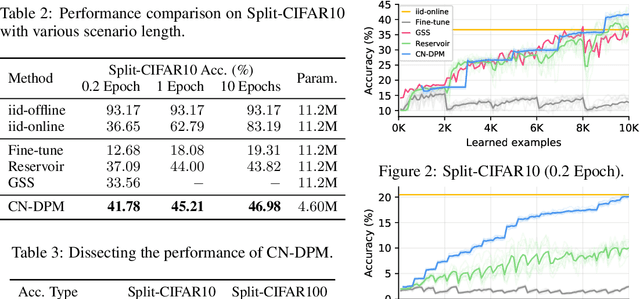
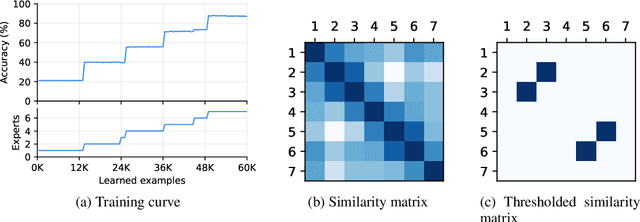
Despite the growing interest in continual learning, most of its contemporary works have been studied in a rather restricted setting where tasks are clearly distinguishable, and task boundaries are known during training. However, if our goal is to develop an algorithm that learns as humans do, this setting is far from realistic, and it is essential to develop a methodology that works in a task-free manner. Meanwhile, among several branches of continual learning, expansion-based methods have the advantage of eliminating catastrophic forgetting by allocating new resources to learn new data. In this work, we propose an expansion-based approach for task-free continual learning. Our model, named Continual Neural Dirichlet Process Mixture (CN-DPM), consists of a set of neural network experts that are in charge of a subset of the data. CN-DPM expands the number of experts in a principled way under the Bayesian nonparametric framework. With extensive experiments, we show that our model successfully performs task-free continual learning for both discriminative and generative tasks such as image classification and image generation.
Convolutional Networks with MuxOut Layers as Multi-rate Systems for Image Upscaling
May 22, 2017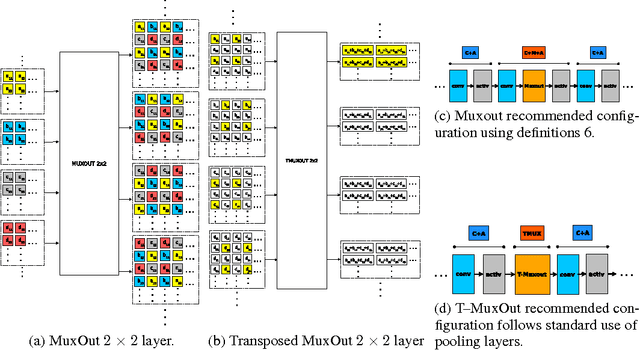
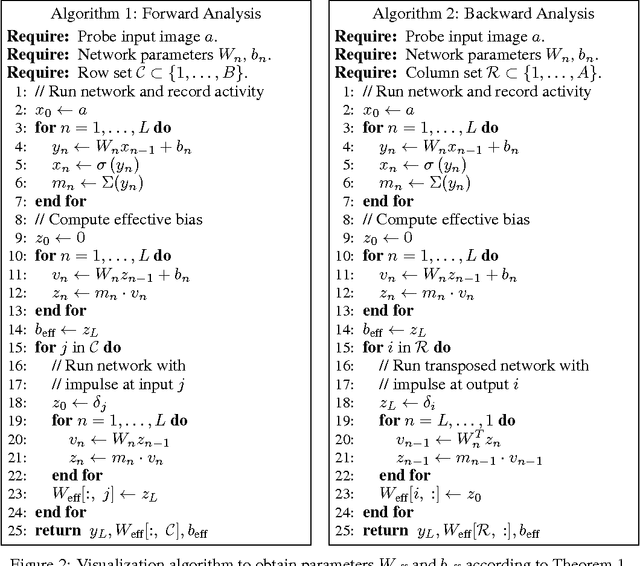

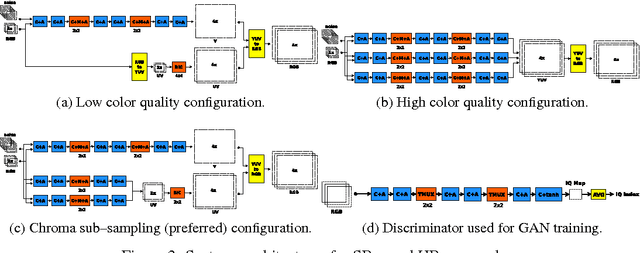
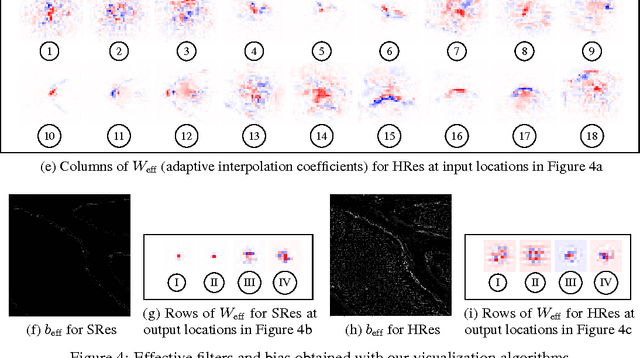
We interpret convolutional networks as adaptive filters and combine them with so-called MuxOut layers to efficiently upscale low resolution images. We formalize this interpretation by deriving a linear and space-variant structure of a convolutional network when its activations are fixed. We introduce general purpose algorithms to analyze a network and show its overall filter effect for each given location. We use this analysis to evaluate two types of image upscalers: deterministic upscalers that target the recovery of details from original content; and second, a new generation of upscalers that can sample the distribution of upscale aliases (images that share the same downscale version) that look like real content.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge