"Image": models, code, and papers
Transfer Learning from Partial Annotations for Whole Brain Segmentation
Aug 28, 2019

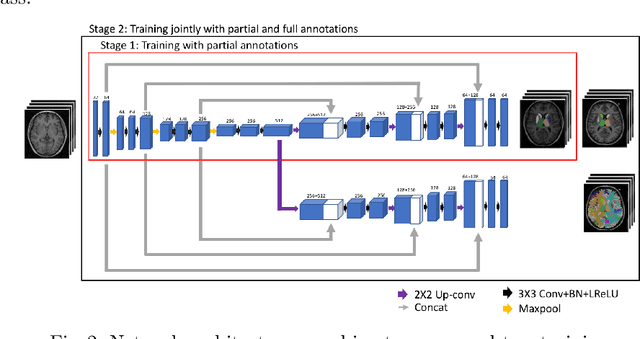
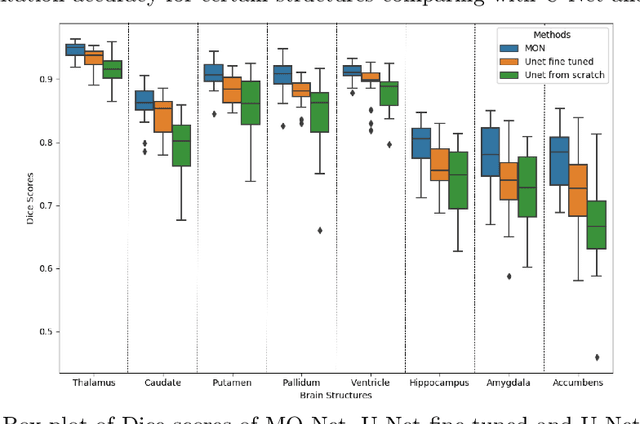
Brain MR image segmentation is a key task in neuroimaging studies. It is commonly conducted using standard computational tools, such as FSL, SPM, multi-atlas segmentation etc, which are often registration-based and suffer from expensive computation cost. Recently, there is an increased interest using deep neural networks for brain image segmentation, which have demonstrated advantages in both speed and performance. However, neural networks-based approaches normally require a large amount of manual annotations for optimising the massive amount of network parameters. For 3D networks used in volumetric image segmentation, this has become a particular challenge, as a 3D network consists of many more parameters compared to its 2D counterpart. Manual annotation of 3D brain images is extremely time-consuming and requires extensive involvement of trained experts. To address the challenge with limited manual annotations, here we propose a novel multi-task learning framework for brain image segmentation, which utilises a large amount of automatically generated partial annotations together with a small set of manually created full annotations for network training. Our method yields a high performance comparable to state-of-the-art methods for whole brain segmentation.
Deep Learning for Quantile Regression: DeepQuantreg
Jul 14, 2020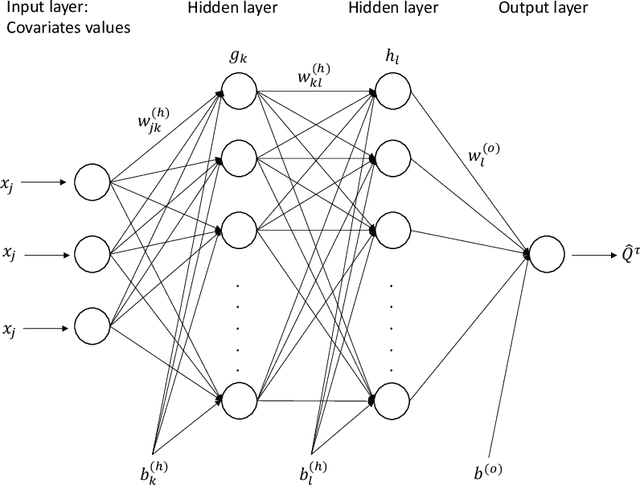
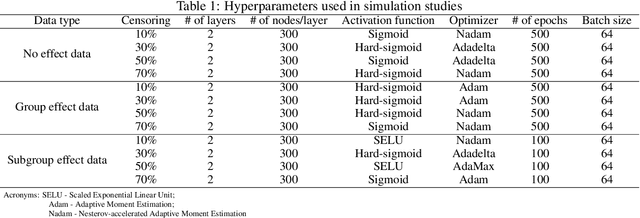
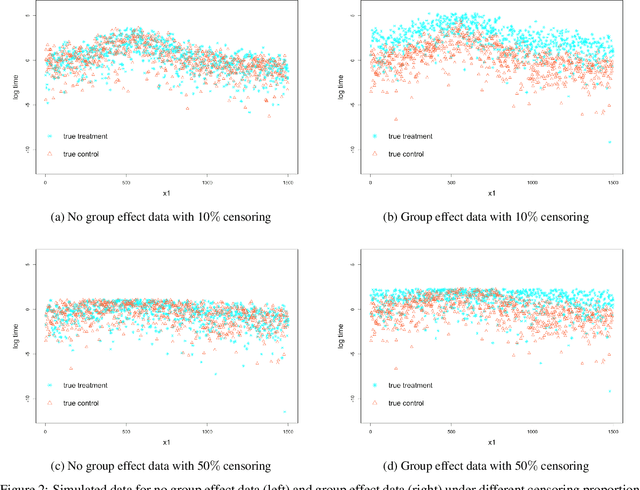
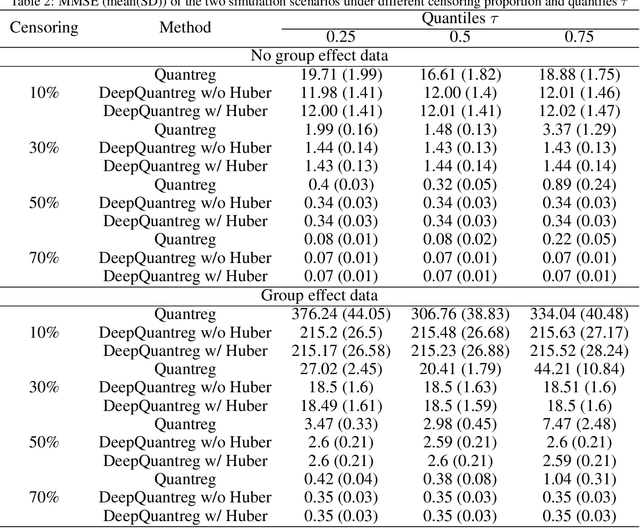
The computational prediction algorithm of neural network, or deep learning, has drawn much attention recently in statistics as well as in image recognition and natural language processing. Particularly in statistical application for censored survival data, the loss function used for optimization has been mainly based on the partial likelihood from Cox's model and its variations to utilize existing neural network library such as Keras, which was built upon the open source library of TensorFlow. This paper presents a novel application of the neural network to the quantile regression for survival data with right censoring, which is adjusted by the inverse of the estimated censoring distribution in the check function. The main purpose of this work is to show that the deep learning method could be flexible enough to predict nonlinear patterns more accurately compared to the traditional method even in low-dimensional data, emphasizing on practicality of the method for censored survival data. Simulation studies were performed to generate nonlinear censored survival data and compare the deep learning method with the traditional quantile regression method in terms of prediction accuracy. The proposed method is illustrated with a publicly available breast cancer data set with gene signatures. The source code is freely available at https://github.com/yicjia/DeepQuantreg.
RGB-D Salient Object Detection with Cross-Modality Modulation and Selection
Jul 14, 2020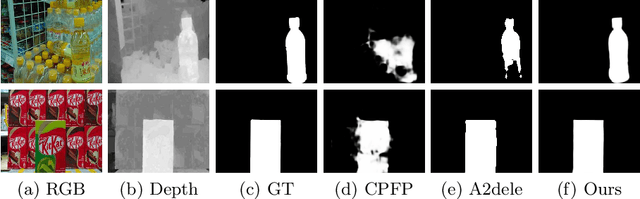
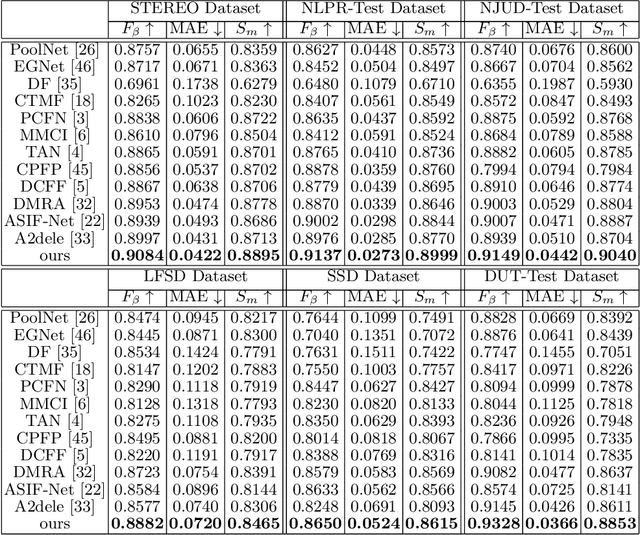
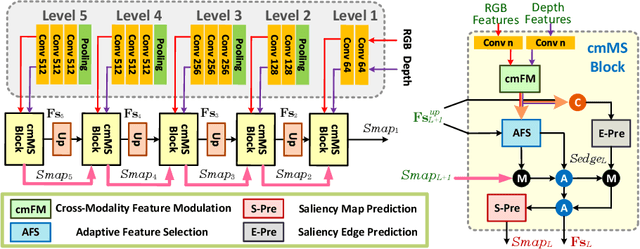
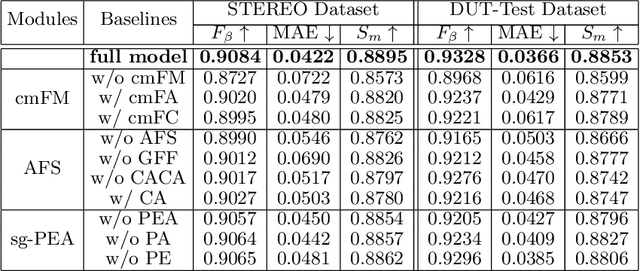
We present an effective method to progressively integrate and refine the cross-modality complementarities for RGB-D salient object detection (SOD). The proposed network mainly solves two challenging issues: 1) how to effectively integrate the complementary information from RGB image and its corresponding depth map, and 2) how to adaptively select more saliency-related features. First, we propose a cross-modality feature modulation (cmFM) module to enhance feature representations by taking the depth features as prior, which models the complementary relations of RGB-D data. Second, we propose an adaptive feature selection (AFS) module to select saliency-related features and suppress the inferior ones. The AFS module exploits multi-modality spatial feature fusion with the self-modality and cross-modality interdependencies of channel features are considered. Third, we employ a saliency-guided position-edge attention (sg-PEA) module to encourage our network to focus more on saliency-related regions. The above modules as a whole, called cmMS block, facilitates the refinement of saliency features in a coarse-to-fine fashion. Coupled with a bottom-up inference, the refined saliency features enable accurate and edge-preserving SOD. Extensive experiments demonstrate that our network outperforms state-of-the-art saliency detectors on six popular RGB-D SOD benchmarks.
The efficiency of deep learning algorithms for detecting anatomical reference points on radiological images of the head profile
May 25, 2020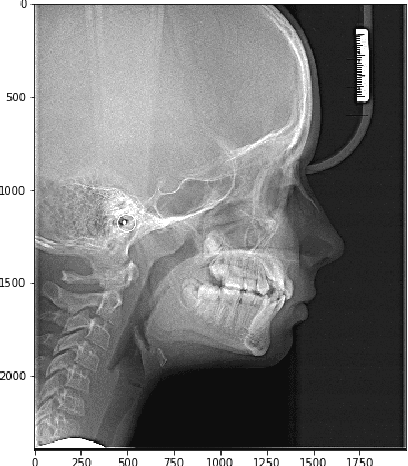
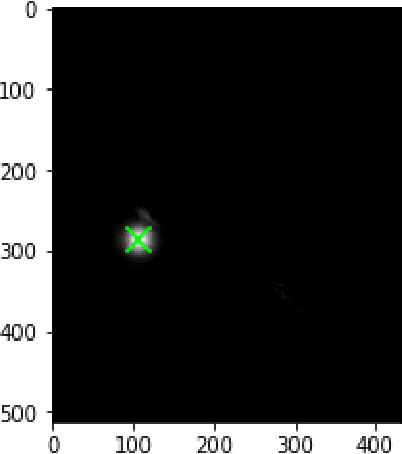
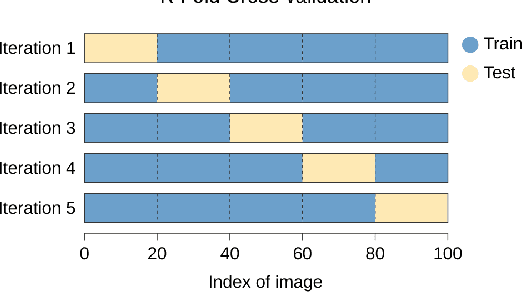
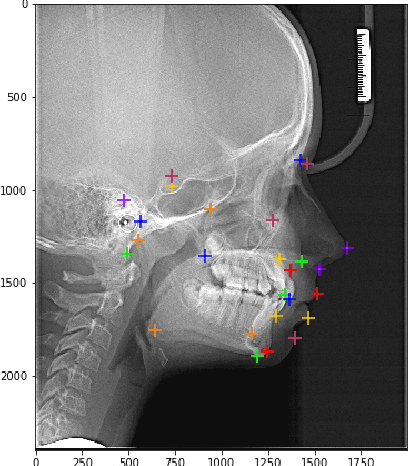
In this article we investigate the efficiency of deep learning algorithms in solving the task of detecting anatomical reference points on radiological images of the head in lateral projection using a fully convolutional neural network and a fully convolutional neural network with an extended architecture for biomedical image segmentation - U-Net. A comparison is made for the results of detection anatomical reference points for each of the selected neural network architectures and their comparison with the results obtained when orthodontists detected anatomical reference points. Based on the obtained results, it was concluded that a U-Net neural network allows performing the detection of anatomical reference points more accurately than a fully convolutional neural network. The results of the detection of anatomical reference points by the U-Net neural network are closer to the average results of the detection of reference points by a group of orthodontists.
Head2Head++: Deep Facial Attributes Re-Targeting
Jun 17, 2020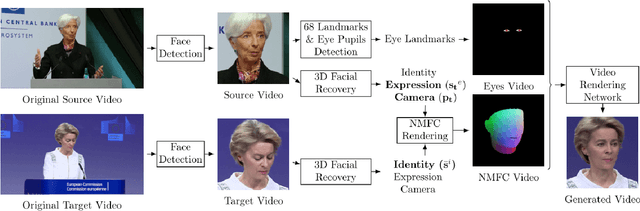
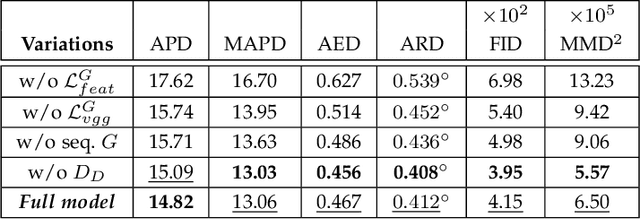

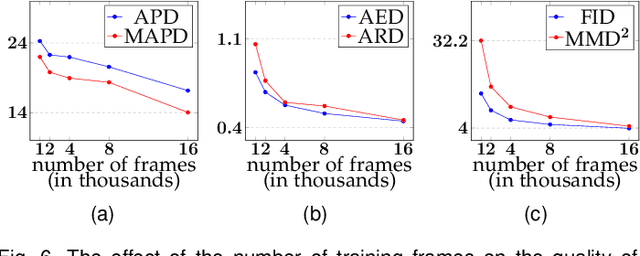
Facial video re-targeting is a challenging problem aiming to modify the facial attributes of a target subject in a seamless manner by a driving monocular sequence. We leverage the 3D geometry of faces and Generative Adversarial Networks (GANs) to design a novel deep learning architecture for the task of facial and head reenactment. Our method is different to purely 3D model-based approaches, or recent image-based methods that use Deep Convolutional Neural Networks (DCNNs) to generate individual frames. We manage to capture the complex non-rigid facial motion from the driving monocular performances and synthesise temporally consistent videos, with the aid of a sequential Generator and an ad-hoc Dynamics Discriminator network. We conduct a comprehensive set of quantitative and qualitative tests and demonstrate experimentally that our proposed method can successfully transfer facial expressions, head pose and eye gaze from a source video to a target subject, in a photo-realistic and faithful fashion, better than other state-of-the-art methods. Most importantly, our system performs end-to-end reenactment in nearly real-time speed (18 fps).
The Semantic Mutex Watershed for Efficient Bottom-Up Semantic Instance Segmentation
Dec 29, 2019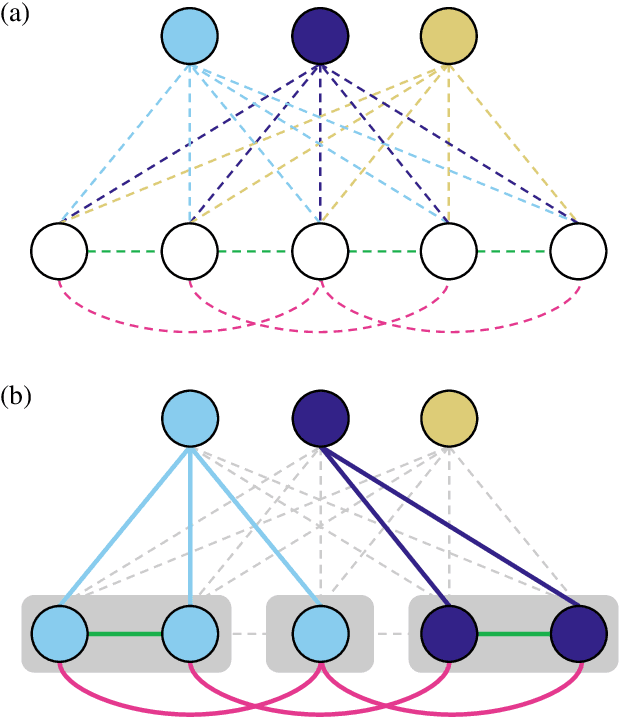
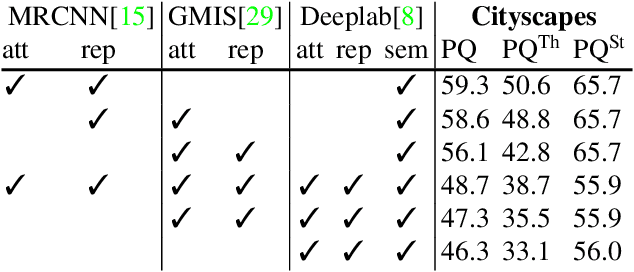
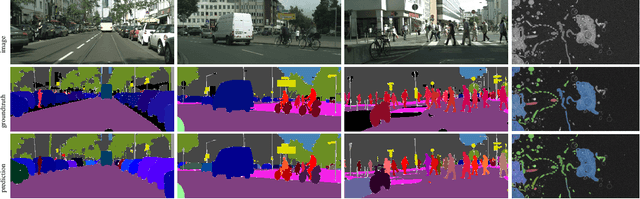


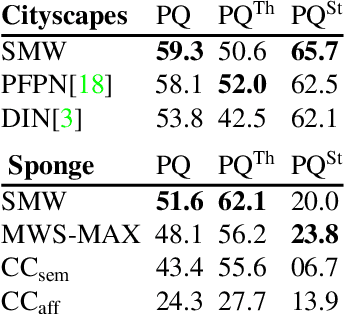
Semantic instance segmentation is the task of simultaneously partitioning an image into distinct segments while associating each pixel with a class label. In commonly used pipelines, segmentation and label assignment are solved separately since joint optimization is computationally expensive. We propose a greedy algorithm for joint graph partitioning and labeling derived from the efficient Mutex Watershed partitioning algorithm. It optimizes an objective function closely related to the Symmetric Multiway Cut objective and empirically shows efficient scaling behavior. Due to the algorithm's efficiency it can operate directly on pixels without prior over-segmentation of the image into superpixels. We evaluate the performance on the Cityscapes dataset (2D urban scenes) and on a 3D microscopy volume. In urban scenes, the proposed algorithm combined with current deep neural networks outperforms the strong baseline of `Panoptic Feature Pyramid Networks' by Kirillov et al. (2019). In the 3D electron microscopy images, we show explicitly that our joint formulation outperforms a separate optimization of the partitioning and labeling problems.
AnalogNet: Convolutional Neural Network Inference on Analog Focal Plane Sensor Processors
Jun 02, 2020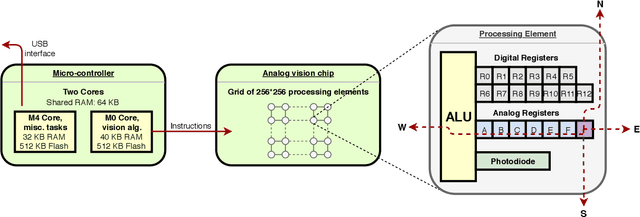
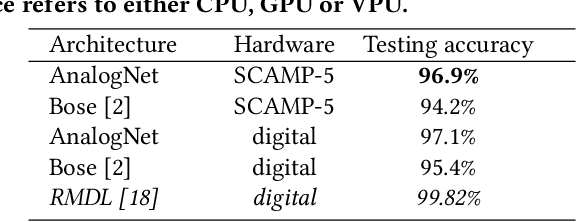
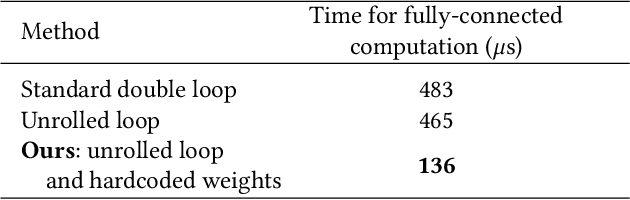


We present a high-speed, energy-efficient Convolutional Neural Network (CNN) architecture utilising the capabilities of a unique class of devices known as analog Focal Plane Sensor Processors (FPSP), in which the sensor and the processor are embedded together on the same silicon chip. Unlike traditional vision systems, where the sensor array sends collected data to a separate processor for processing, FPSPs allow data to be processed on the imaging device itself. This unique architecture enables ultra-fast image processing and high energy efficiency, at the expense of limited processing resources and approximate computations. In this work, we show how to convert standard CNNs to FPSP code, and demonstrate a method of training networks to increase their robustness to analog computation errors. Our proposed architecture, coined AnalogNet, reaches a testing accuracy of 96.9% on the MNIST handwritten digits recognition task, at a speed of 2260 FPS, for a cost of 0.7 mJ per frame.
Deep Learning-based Denoising of Mammographic Images using Physics-driven Data Augmentation
Dec 11, 2019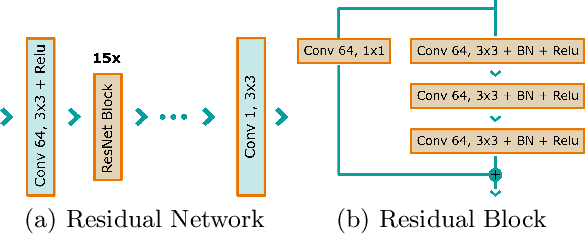
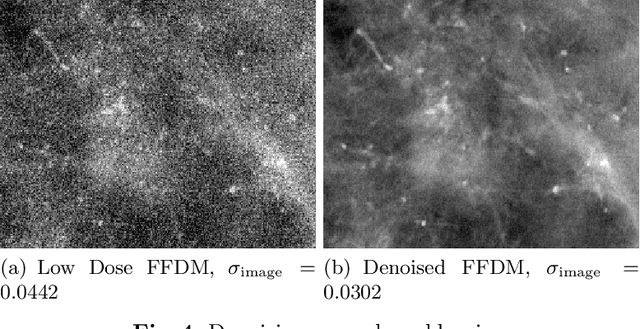


Mammography is using low-energy X-rays to screen the human breast and is utilized by radiologists to detect breast cancer. Typically radiologists require a mammogram with impeccable image quality for an accurate diagnosis. In this study, we propose a deep learning method based on Convolutional Neural Networks (CNNs) for mammogram denoising to improve the image quality. We first enhance the noise level and employ Anscombe Transformation (AT) to transform Poisson noise to white Gaussian noise. With this data augmentation, a deep residual network is trained to learn the noise map of the noisy images. We show, that the proposed method can remove not only simulated but also real noise. Furthermore, we also compare our results with state-of-the-art denoising methods, such as BM3D and DNCNN. In an early investigation, we achieved qualitatively better mammogram denoising results.
Flows Succeed Where GANs Fail: Lessons from Low-Dimensional Data
Jun 17, 2020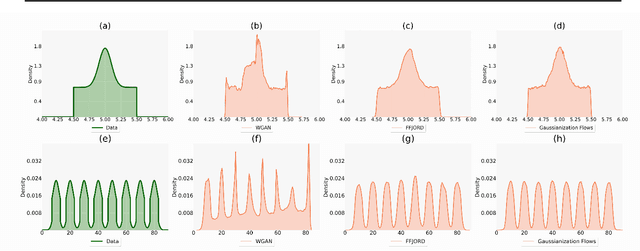
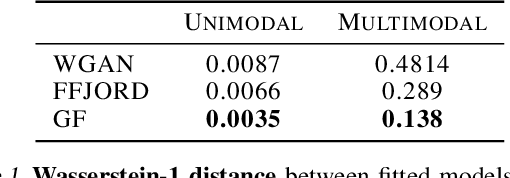
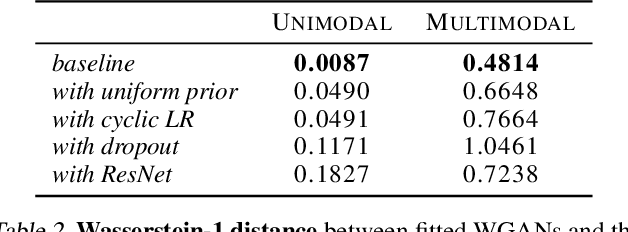
Normalizing flows and generative adversarial networks (GANs) are both approaches to density estimation that use deep neural networks to transform samples from an uninformative prior distribution to an approximation of the data distribution. There is great interest in both for general-purpose statistical modeling, but the two approaches have seldom been compared to each other for modeling non-image data. The difficulty of computing likelihoods with GANs, which are implicit models, makes conducting such a comparison challenging. We work around this difficulty by considering several low-dimensional synthetic datasets. An extensive grid search over GAN architectures, hyperparameters, and training procedures suggests that no GAN is capable of modeling our simple low-dimensional data well, a task we view as a prerequisite for an approach to be considered suitable for general-purpose statistical modeling. Several normalizing flows, on the other hand, excelled at these tasks, even substantially outperforming WGAN in terms of Wasserstein distance---the metric that WGAN alone targets. Overall, normalizing flows appear to be more reliable tools for statistical inference than GANs.
BiNet: Degraded-Manuscript Binarization in Diverse Document Textures and Layouts using Deep Encoder-Decoder Networks
Nov 13, 2019
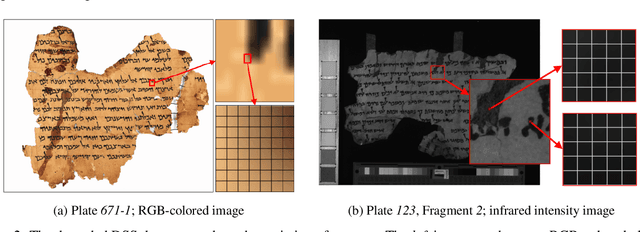
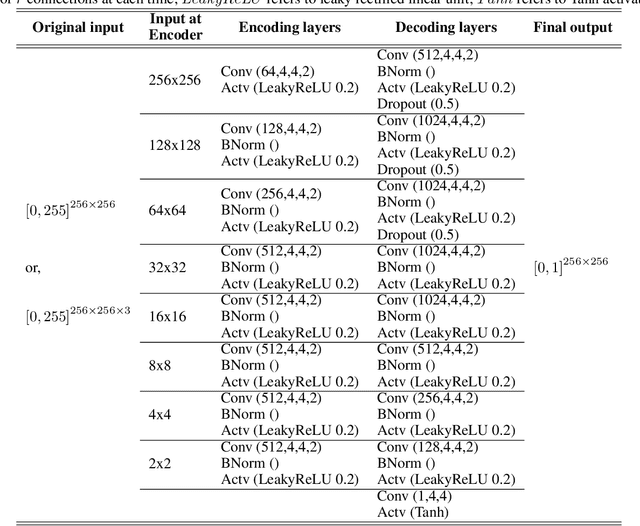

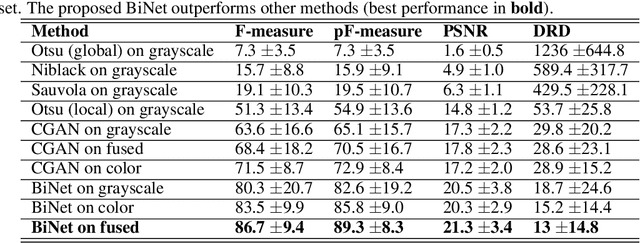
Handwritten document-image binarization is a semantic segmentation process to differentiate ink pixels from background pixels. It is one of the essential steps towards character recognition, writer identification, and script-style evolution analysis. The binarization task itself is challenging due to the vast diversity of writing styles, inks, and paper materials. It is even more difficult for historical manuscripts due to the aging and degradation of the documents over time. One of such manuscripts is the Dead Sea Scrolls (DSS) image collection, which poses extreme challenges for the existing binarization techniques. This article proposes a new binarization technique for the DSS images using the deep encoder-decoder networks. Although the artificial neural network proposed here is primarily designed to binarize the DSS images, it can be trained on different manuscript collections as well. Additionally, the use of transfer learning makes the network already utilizable for a wide range of handwritten documents, making it a unique multi-purpose tool for binarization. Qualitative results and several quantitative comparisons using both historical manuscripts and datasets from handwritten document image binarization competition (H-DIBCO and DIBCO) exhibit the robustness and the effectiveness of the system. The best performing network architecture proposed here is a variant of the U-Net encoder-decoders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge