"Image": models, code, and papers
DeepBiRD: An Automatic Bibliographic Reference Detection Approach
Dec 16, 2019
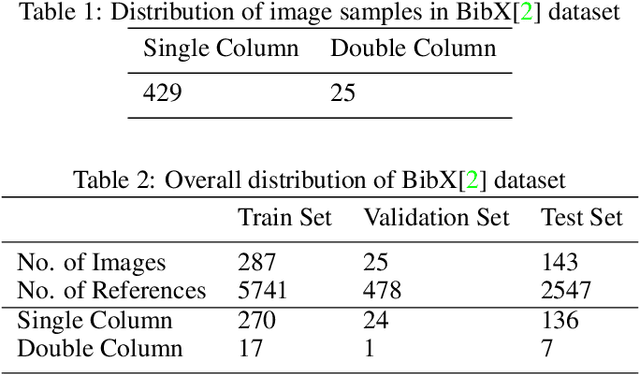

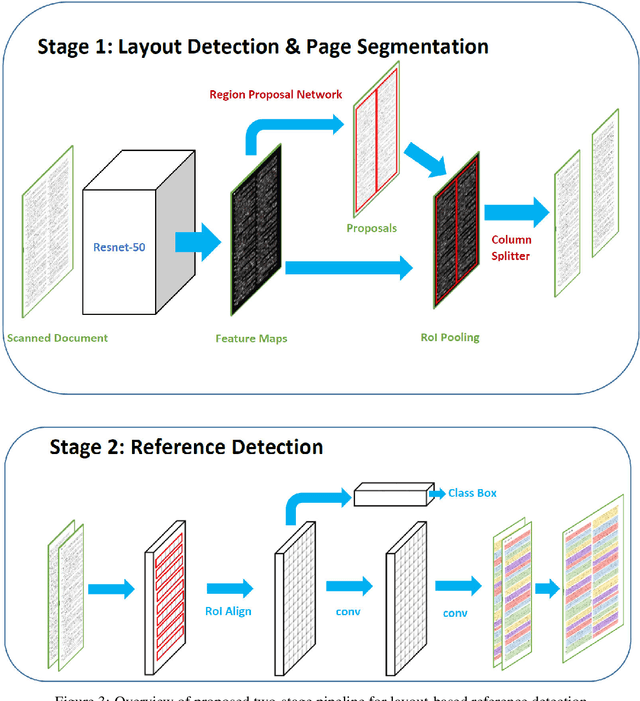
The contribution of this paper is two fold. First, it presents a novel approach called DeepBiRD which is inspired from human visual perception and exploits layout features to identify individual references in a scientific publication. Second, we present a new dataset for image-based reference detection with 2401 scans containing 12244 references, all manually annotated for individual reference. Our proposed approach consists of two stages, firstly it identifies whether given document image is single column or multi-column. Using this information, document image is then splitted into individual columns. Secondly it performs layout driven reference detection using Mask R-CNN in a given scientific publication. DeepBiRD was evaluated on two different datasets to demonstrate the generalization of this approach. The proposed system achieved an F-measure of 0.96 on our dataset. DeepBiRD detected 2.5 times more references than current state-of-the-art approach on their own dataset. Therefore, suggesting that DeepBiRD is significantly superior in performance, generalizable and independent of any domain or referencing style.
MARTA GANs: Unsupervised Representation Learning for Remote Sensing Image Classification
Nov 21, 2017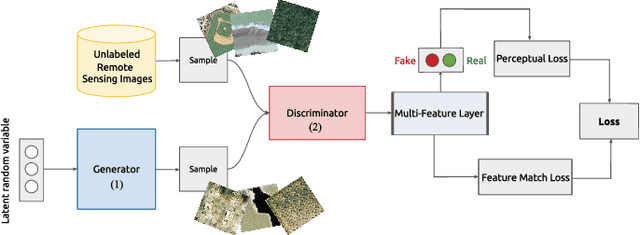
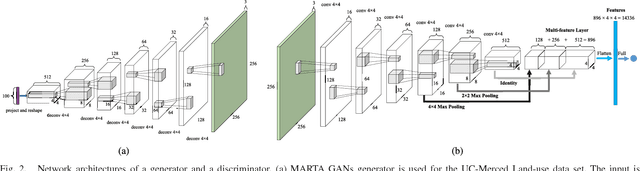

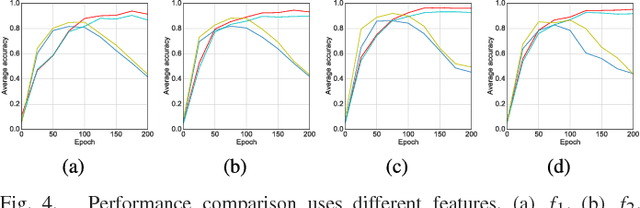
With the development of deep learning, supervised learning has frequently been adopted to classify remotely sensed images using convolutional networks (CNNs). However, due to the limited amount of labeled data available, supervised learning is often difficult to carry out. Therefore, we proposed an unsupervised model called multiple-layer feature-matching generative adversarial networks (MARTA GANs) to learn a representation using only unlabeled data. MARTA GANs consists of both a generative model $G$ and a discriminative model $D$. We treat $D$ as a feature extractor. To fit the complex properties of remote sensing data, we use a fusion layer to merge the mid-level and global features. $G$ can produce numerous images that are similar to the training data; therefore, $D$ can learn better representations of remotely sensed images using the training data provided by $G$. The classification results on two widely used remote sensing image databases show that the proposed method significantly improves the classification performance compared with other state-of-the-art methods.
* IEEE GRSL
Scilab and SIP for Image Processing
Mar 18, 2012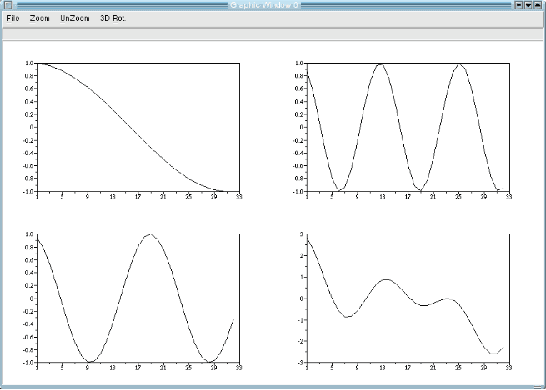
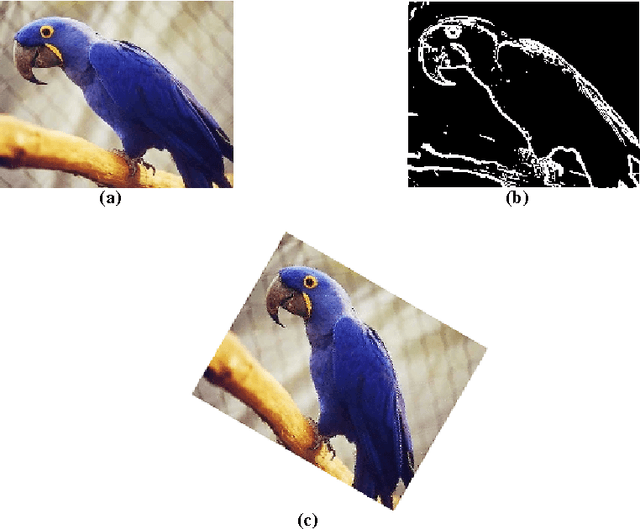
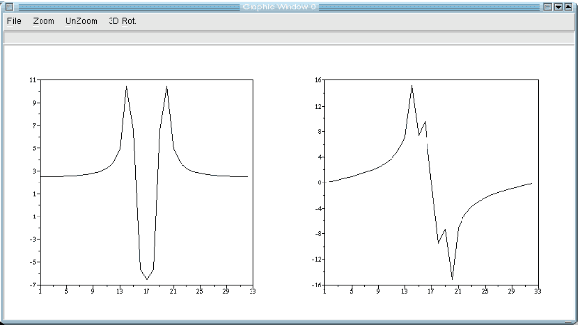
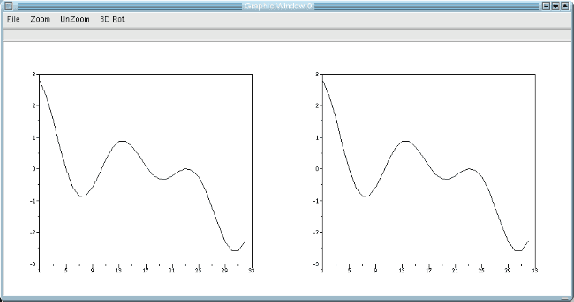

This paper is an overview of Image Processing and Analysis using Scilab, a free prototyping environment for numerical calculations similar to Matlab. We demonstrate the capabilities of SIP -- the Scilab Image Processing Toolbox -- which extends Scilab with many functions to read and write images in over 100 major file formats, including PNG, JPEG, BMP, and TIFF. It also provides routines for image filtering, edge detection, blurring, segmentation, shape analysis, and image recognition. Basic directions to install Scilab and SIP are given, and also a mini-tutorial on Scilab. Three practical examples of image analysis are presented, in increasing degrees of complexity, showing how advanced image analysis techniques seems uncomplicated in this environment.
Human eye inspired log-polar pre-processing for neural networks
Nov 04, 2019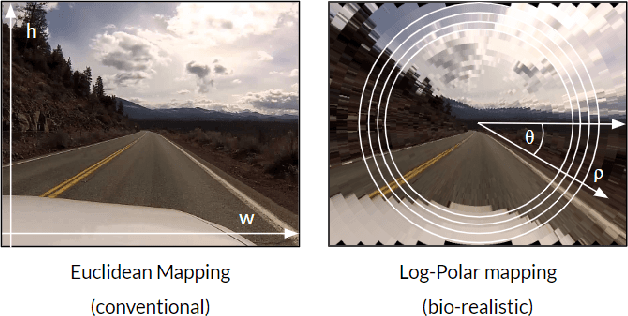
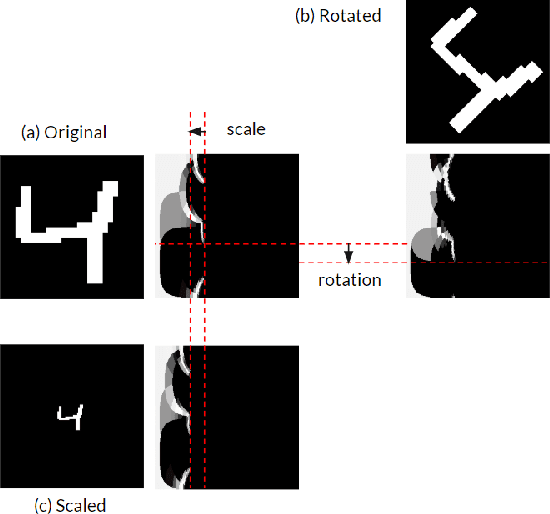
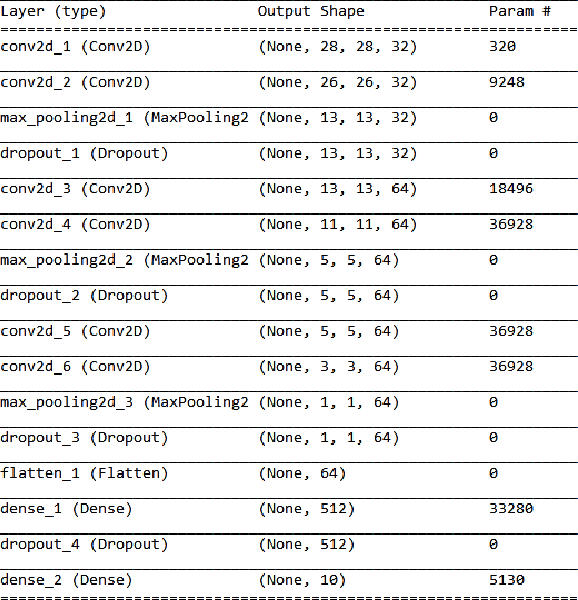

In this paper we draw inspiration from the human visual system, and present a bio-inspired pre-processing stage for neural networks. We implement this by applying a log-polar transformation as a pre-processing step, and to demonstrate, we have used a naive convolutional neural network (CNN). We demonstrate that a bio-inspired pre-processing stage can achieve rotation and scale robustness in CNNs. A key point in this paper is that the CNN does not need to be trained to identify rotation or scaling permutations; rather it is the log-polar pre-processing step that converts the image into a format that allows the CNN to handle rotation and scaling permutations. In addition we demonstrate how adding a log-polar transformation as a pre-processing step can reduce the image size to ~20\% of the Euclidean image size, without significantly compromising classification accuracy of the CNN. The pre-processing stage presented in this paper is modelled after the retina and therefore is only tested against an image dataset. Note: This paper has been submitted for SAUPEC/RobMech/PRASA 2020.
REFINED (REpresentation of Features as Images with NEighborhood Dependencies): A novel feature representation for Convolutional Neural Networks
Dec 11, 2019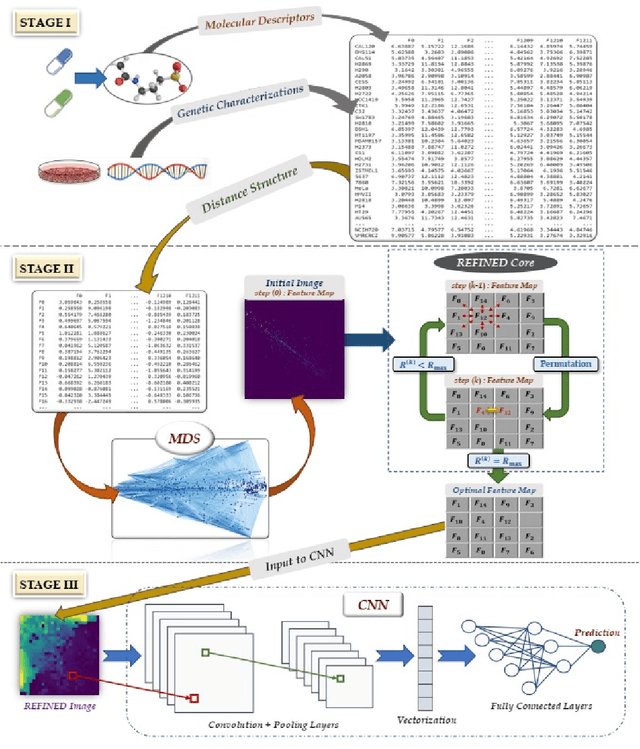
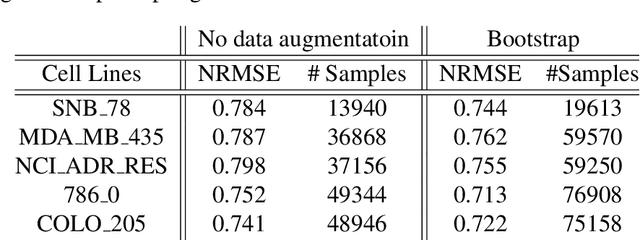
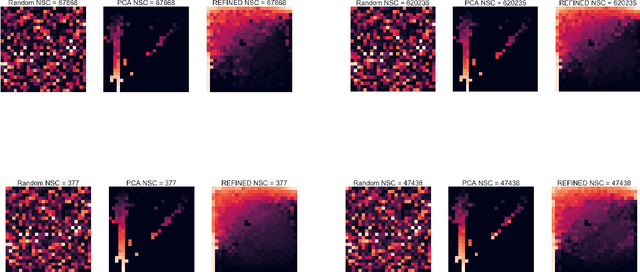
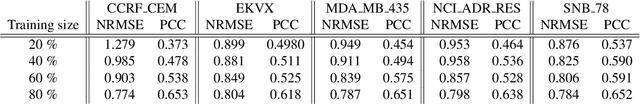
Deep learning with Convolutional Neural Networks has shown great promise in various areas of image-based classification and enhancement but is often unsuitable for predictive modeling involving non-image based features or features without spatial correlations. We present a novel approach for representation of high dimensional feature vector in a compact image form, termed REFINED (REpresentation of Features as Images with NEighborhood Dependencies), that is conducible for convolutional neural network based deep learning. We consider the correlations between features to generate a compact representation of the features in the form of a two-dimensional image using minimization of pairwise distances similar to multi-dimensional scaling. We hypothesize that this approach enables embedded feature selection and integrated with Convolutional Neural Network based Deep Learning can produce more accurate predictions as compared to Artificial Neural Networks, Random Forests and Support Vector Regression. We illustrate the superior predictive performance of the proposed representation, as compared to existing approaches, using synthetic datasets, cell line efficacy prediction based on drug chemical descriptors for NCI60 dataset and drug sensitivity prediction based on transcriptomic data and chemical descriptors using GDSC dataset. Results illustrated on both synthetic and biological datasets shows the higher prediction accuracy of the proposed framework as compared to existing methodologies while maintaining desirable properties in terms of bias and feature extraction.
Frame-To-Frame Consistent Semantic Segmentation
Aug 27, 2020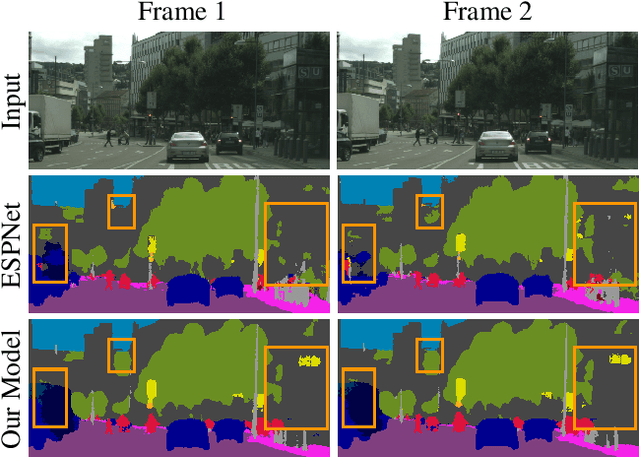
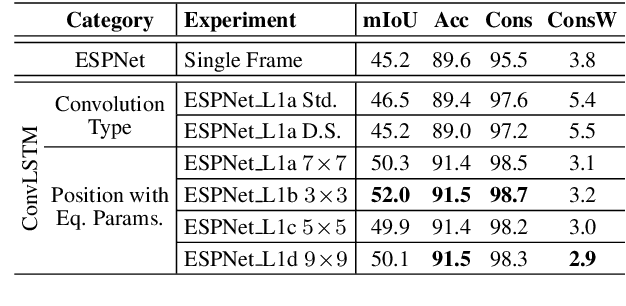
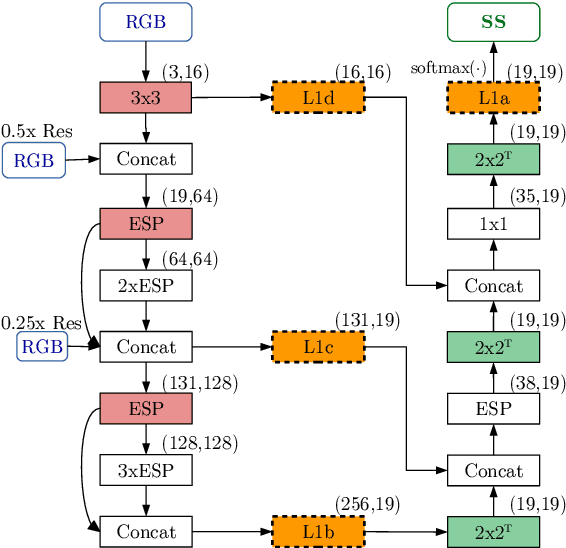
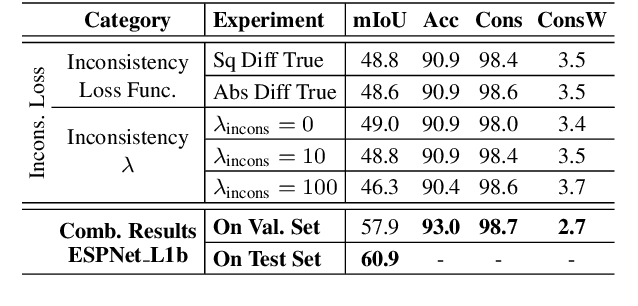
In this work, we aim for temporally consistent semantic segmentation throughout frames in a video. Many semantic segmentation algorithms process images individually which leads to an inconsistent scene interpretation due to illumination changes, occlusions and other variations over time. To achieve a temporally consistent prediction, we train a convolutional neural network (CNN) which propagates features through consecutive frames in a video using a convolutional long short term memory (ConvLSTM) cell. Besides the temporal feature propagation, we penalize inconsistencies in our loss function. We show in our experiments that the performance improves when utilizing video information compared to single frame prediction. The mean intersection over union (mIoU) metric on the Cityscapes validation set increases from 45.2 % for the single frames to 57.9 % for video data after implementing the ConvLSTM to propagate features trough time on the ESPNet. Most importantly, inconsistency decreases from 4.5 % to 1.3 % which is a reduction by 71.1 %. Our results indicate that the added temporal information produces a frame-to-frame consistent and more accurate image understanding compared to single frame processing. Code and videos are available at https://github.com/mrebol/f2f-consistent-semantic-segmentation
Assessing the validity of saliency maps for abnormality localization in medical imaging
May 29, 2020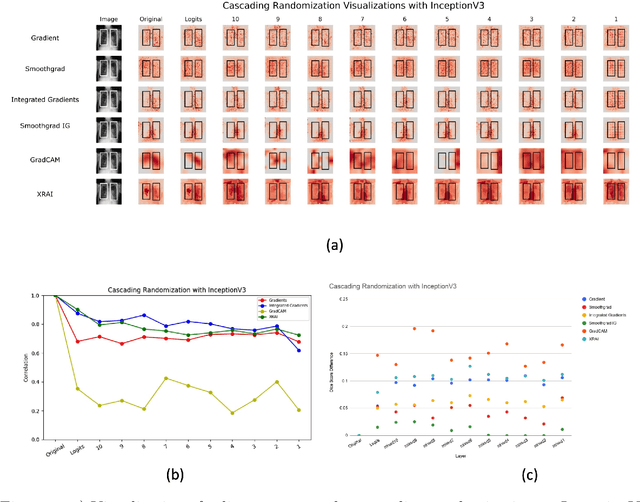
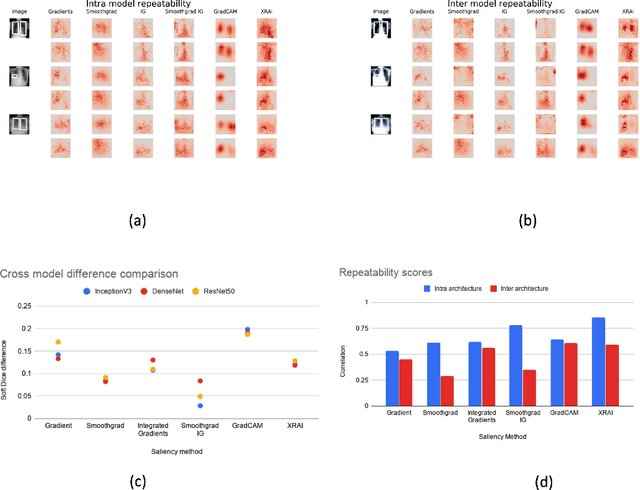
Saliency maps have become a widely used method to assess which areas of the input image are most pertinent to the prediction of a trained neural network. However, in the context of medical imaging, there is no study to our knowledge that has examined the efficacy of these techniques and quantified them using overlap with ground truth bounding boxes. In this work, we explored the credibility of the various existing saliency map methods on the RSNA Pneumonia dataset. We found that GradCAM was the most sensitive to model parameter and label randomization, and was highly agnostic to model architecture.
Transductive Zero-Shot Learning for 3D Point Cloud Classification
Dec 16, 2019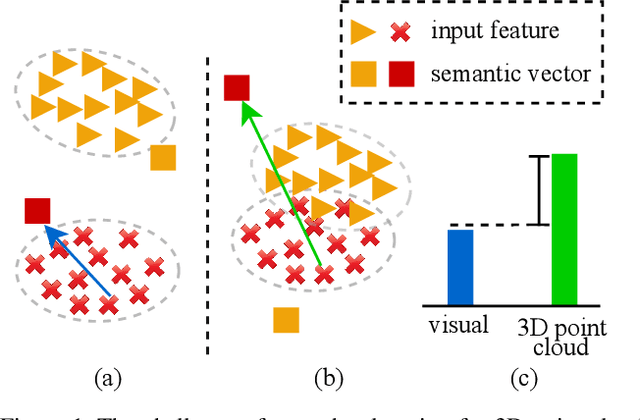
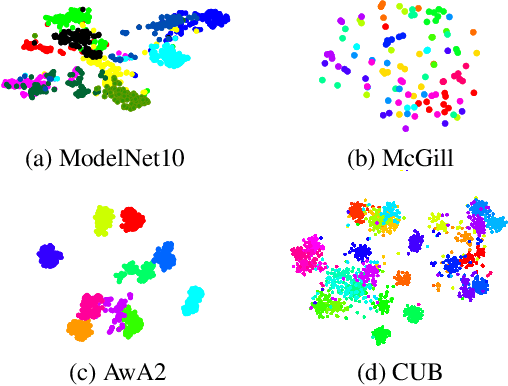
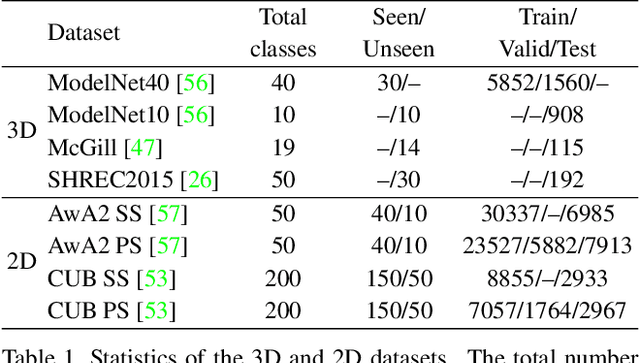

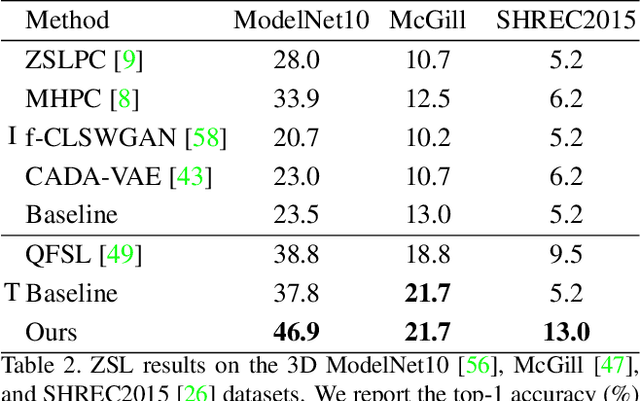
Zero-shot learning, the task of learning to recognize new classes not seen during training, has received considerable attention in the case of 2D image classification. However despite the increasing ubiquity of 3D sensors, the corresponding 3D point cloud classification problem has not been meaningfully explored and introduces new challenges. This paper extends, for the first time, transductive Zero-Shot Learning (ZSL) and Generalized Zero-Shot Learning (GZSL) approaches to the domain of 3D point cloud classification. To this end, a novel triplet loss is developed that takes advantage of unlabeled test data. While designed for the task of 3D point cloud classification, the method is also shown to be applicable to the more common use-case of 2D image classification. An extensive set of experiments is carried out, establishing state-of-the-art for ZSL and GZSL in the 3D point cloud domain, as well as demonstrating the applicability of the approach to the image domain.
FA-GANs: Facial Attractiveness Enhancement with Generative Adversarial Networks on Frontal Faces
May 17, 2020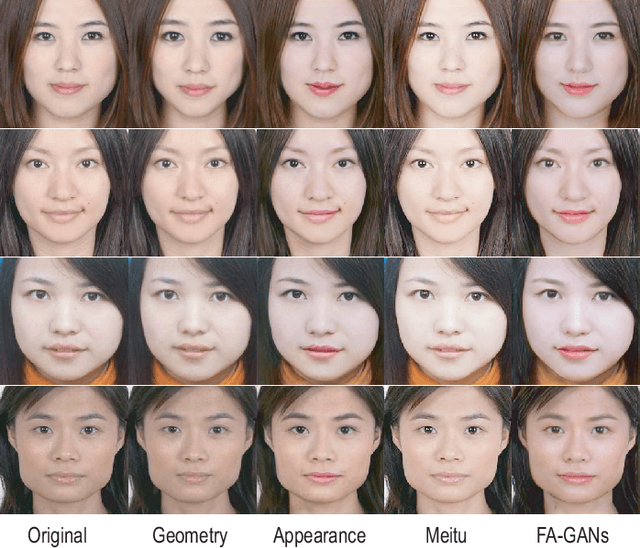
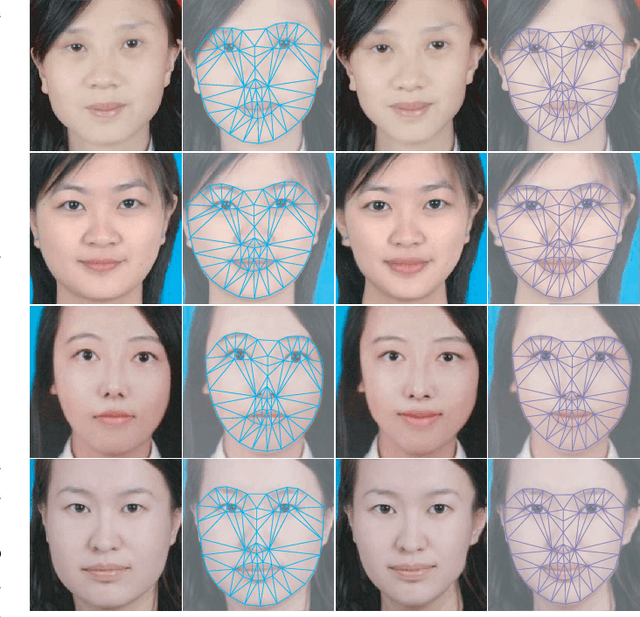
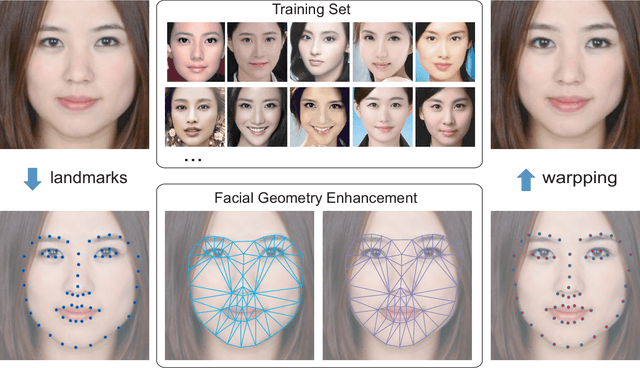
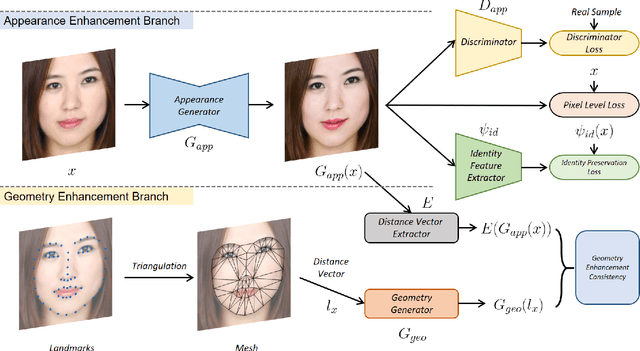


Facial attractiveness enhancement has been an interesting application in Computer Vision and Graphics over these years. It aims to generate a more attractive face via manipulations on image and geometry structure while preserving face identity. In this paper, we propose the first Generative Adversarial Networks (GANs) for enhancing facial attractiveness in both geometry and appearance aspects, which we call "FA-GANs". FA-GANs contain two branches and enhance facial attractiveness in two perspectives: facial geometry and facial appearance. Each branch consists of individual GANs with the appearance branch adjusting the facial image and the geometry branch adjusting the facial landmarks in appearance and geometry aspects, respectively. Unlike the traditional facial manipulations learning from paired faces, which are infeasible to collect before and after enhancement of the same individual, we achieve this by learning the features of attractiveness faces through unsupervised adversarial learning. The proposed FA-GANs are able to extract attractiveness features and impose them on the enhancement results. To better enhance faces, both the geometry and appearance networks are considered to refine the facial attractiveness by adjusting the geometry layout of faces and the appearance of faces independently. To the best of our knowledge, we are the first to enhance the facial attractiveness with GANs in both geometry and appearance aspects. The experimental results suggest that our FA-GANs can generate compelling perceptual results in both geometry structure and facial appearance and outperform current state-of-the-art methods.
DeepAtrophy: Teaching a Neural Network to Differentiate Progressive Changes from Noise on Longitudinal MRI in Alzheimer's Disease
Oct 24, 2020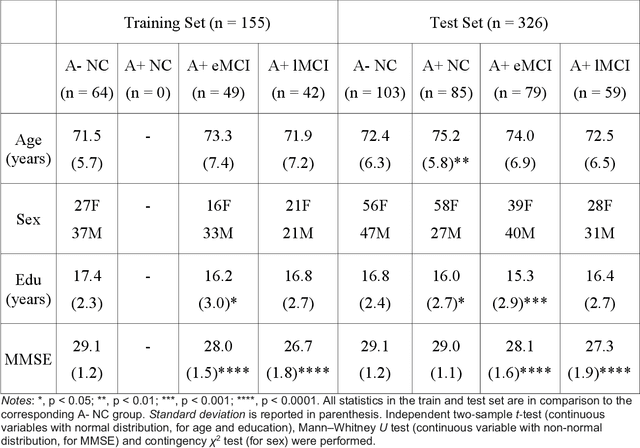
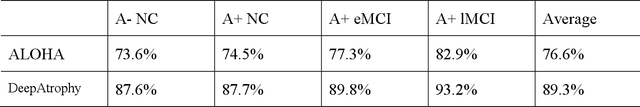
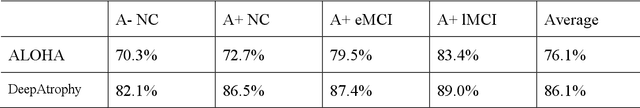
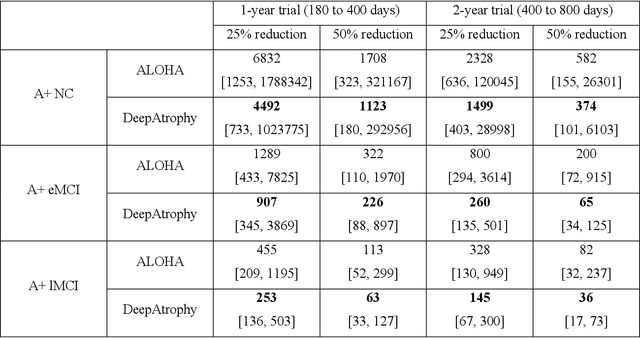
Volume change measures derived from longitudinal MRI (e.g. hippocampal atrophy) are a well-studied biomarker of disease progression in Alzheimer's Disease (AD) and are used in clinical trials to track the therapeutic efficacy of disease-modifying treatments. However, longitudinal MRI change measures can be confounded by non-biological factors, such as different degrees of head motion and susceptibility artifact between pairs of MRI scans. We hypothesize that deep learning methods applied directly to pairs of longitudinal MRI scans can be trained to differentiate between biological changes and non-biological factors better than conventional approaches based on deformable image registration. To achieve this, we make a simplifying assumption that biological factors are associated with time (i.e. the hippocampus shrinks overtime in the aging population) whereas non-biological factors are independent of time. We then formulate deep learning networks to infer the temporal order of same-subject MRI scans input to the network in arbitrary order; as well as to infer ratios between interscan intervals for two pairs of same-subject MRI scans. In the test dataset, these networks perform better in tasks of temporal ordering (89.3%) and interscan interval inference (86.1%) than a state-of-the-art deformation-based morphometry method ALOHA (76.6% and 76.1% respectively) (Das et al., 2012). Furthermore, we derive a disease progression score from the network that is able to detect a group difference between 58 preclinical AD and 75 beta-amyloid-negative cognitively normal individuals within one year, compared to two years for ALOHA. This suggests that deep learning can be trained to differentiate MRI changes due to biological factors (tissue loss) from changes due to non-biological factors, leading to novel biomarkers that are more sensitive to longitudinal changes at the earliest stages of AD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge