"Image": models, code, and papers
Supervised Segmentation of Retinal Vessel Structures Using ANN
Jan 15, 2020
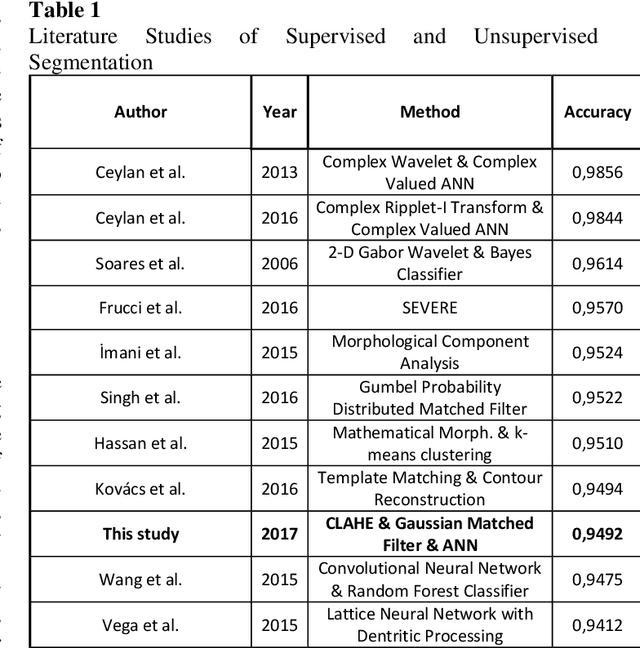
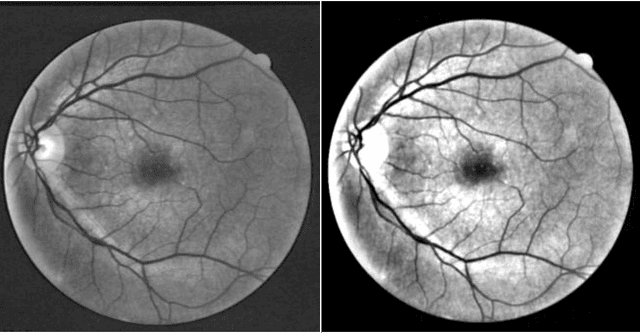
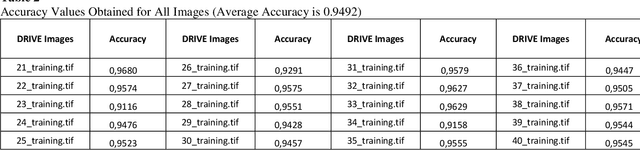
In this study, a supervised retina blood vessel segmentation process was performed on the green channel of the RGB image using artificial neural network (ANN). The green channel is preferred because the retinal vessel structures can be distinguished most clearly from the green channel of the RGB image. The study was performed using 20 images in the DRIVE data set which is one of the most common retina data sets known. The images went through some preprocessing stages like contrastlimited adaptive histogram equalization (CLAHE), color intensity adjustment, morphological operations and median and Gaussian filtering to obtain a good segmentation. Retinal vessel structures were highlighted with top-hat and bot-hat morphological operations and converted to binary image by using global thresholding. Then, the network was trained by the binary version of the images specified as training images in the dataset and the targets are the images segmented manually by a specialist. The average segmentation accuracy for 20 images was found as 0.9492.
StereoGAN: Bridging Synthetic-to-Real Domain Gap by Joint Optimization of Domain Translation and Stereo Matching
May 05, 2020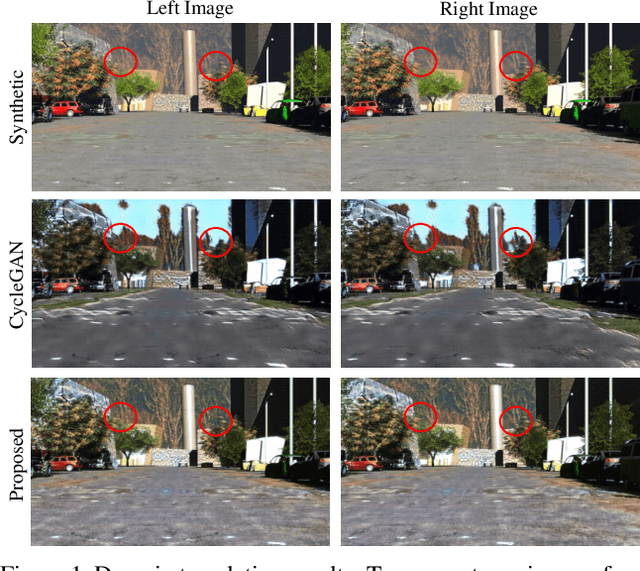
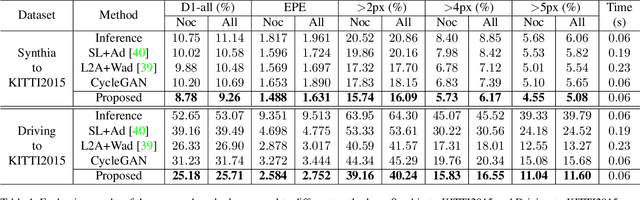

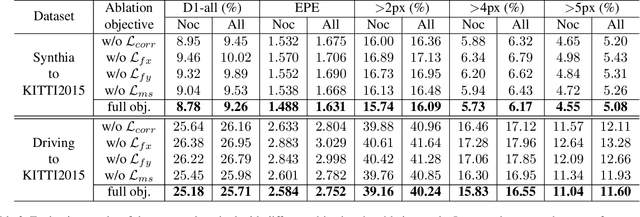
Large-scale synthetic datasets are beneficial to stereo matching but usually introduce known domain bias. Although unsupervised image-to-image translation networks represented by CycleGAN show great potential in dealing with domain gap, it is non-trivial to generalize this method to stereo matching due to the problem of pixel distortion and stereo mismatch after translation. In this paper, we propose an end-to-end training framework with domain translation and stereo matching networks to tackle this challenge. First, joint optimization between domain translation and stereo matching networks in our end-to-end framework makes the former facilitate the latter one to the maximum extent. Second, this framework introduces two novel losses, i.e., bidirectional multi-scale feature re-projection loss and correlation consistency loss, to help translate all synthetic stereo images into realistic ones as well as maintain epipolar constraints. The effective combination of above two contributions leads to impressive stereo-consistent translation and disparity estimation accuracy. In addition, a mode seeking regularization term is added to endow the synthetic-to-real translation results with higher fine-grained diversity. Extensive experiments demonstrate the effectiveness of the proposed framework on bridging the synthetic-to-real domain gap on stereo matching.
Pixel-wise Conditioning of Generative Adversarial Networks
Nov 02, 2019
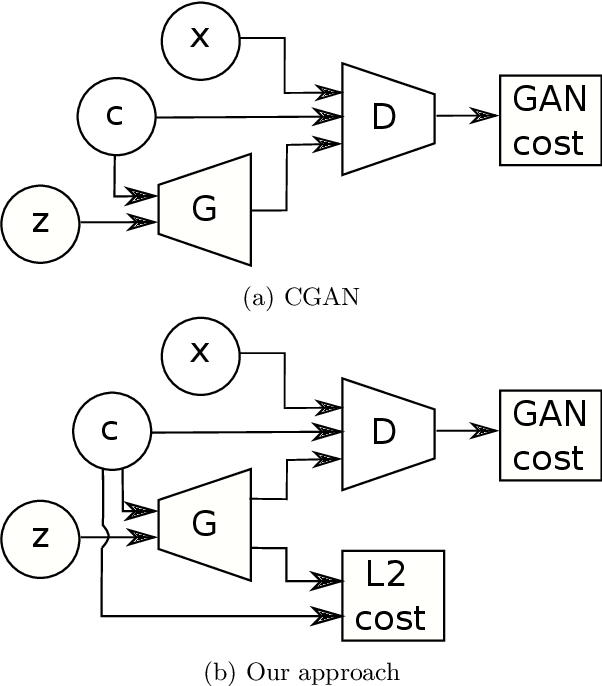
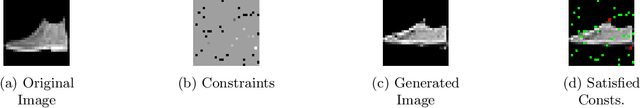
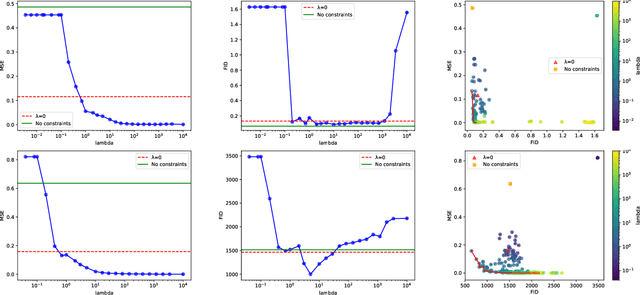
Generative Adversarial Networks (GANs) have proven successful for unsupervised image generation. Several works extended GANs to image inpainting by conditioning the generation with parts of the image one wants to reconstruct. However, these methods have limitations in settings where only a small subset of the image pixels is known beforehand. In this paper, we study the effectiveness of conditioning GANs by adding an explicit regularization term to enforce pixel-wise conditions when very few pixel values are provided. In addition, we also investigate the influence of this regularization term on the quality of the generated images and the satisfaction of the conditions. Conducted experiments on MNIST and FashionMNIST show evidence that this regularization term allows for controlling the trade-off between quality of the generated images and constraint satisfaction.
Towards Effective Codebookless Model for Image Classification
Jul 20, 2015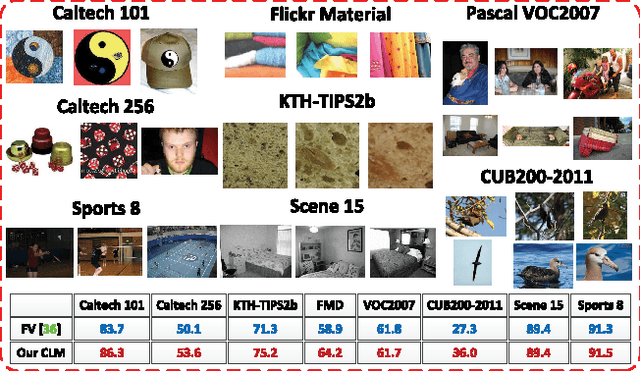
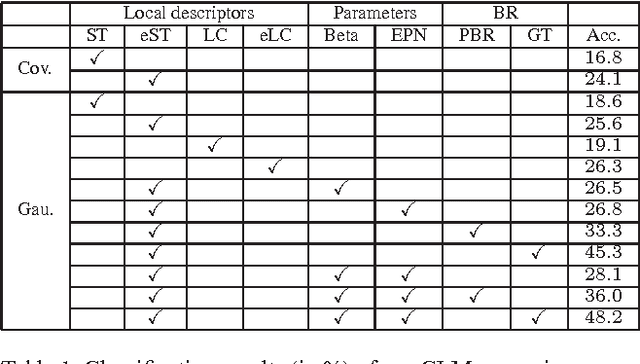
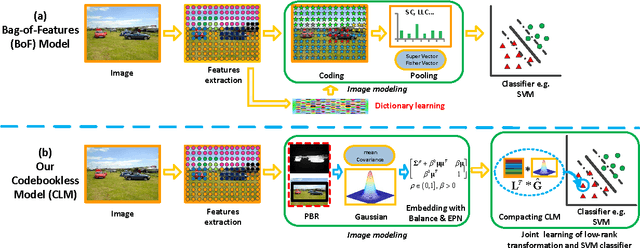
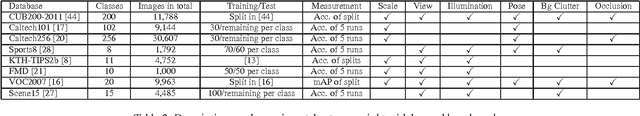
The bag-of-features (BoF) model for image classification has been thoroughly studied over the last decade. Different from the widely used BoF methods which modeled images with a pre-trained codebook, the alternative codebook free image modeling method, which we call Codebookless Model (CLM), attracted little attention. In this paper, we present an effective CLM that represents an image with a single Gaussian for classification. By embedding Gaussian manifold into a vector space, we show that the simple incorporation of our CLM into a linear classifier achieves very competitive accuracy compared with state-of-the-art BoF methods (e.g., Fisher Vector). Since our CLM lies in a high dimensional Riemannian manifold, we further propose a joint learning method of low-rank transformation with support vector machine (SVM) classifier on the Gaussian manifold, in order to reduce computational and storage cost. To study and alleviate the side effect of background clutter on our CLM, we also present a simple yet effective partial background removal method based on saliency detection. Experiments are extensively conducted on eight widely used databases to demonstrate the effectiveness and efficiency of our CLM method.
Bounds for Learning Lossless Source Coding
Sep 18, 2020
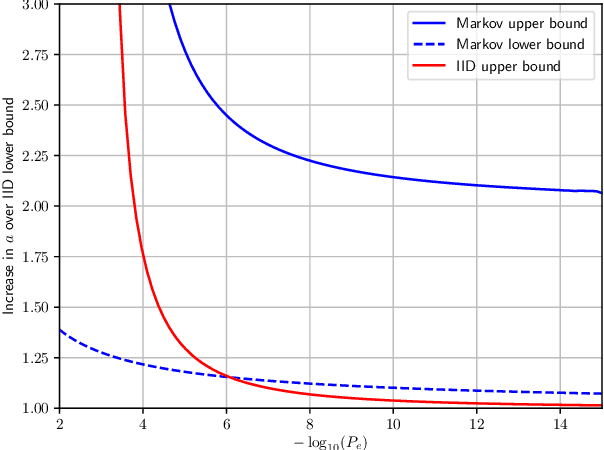
This paper asks a basic question: how much training is required to beat a universal source coder? Traditionally, there have been two types of source coders: fixed, optimum coders such as Huffman coders; and universal source coders, such as Lempel-Ziv The paper considers a third type of source coders: learned coders. These are coders that are trained on data of a particular type, and then used to encode new data of that type. This is a type of coder that has recently become very popular for (lossy) image and video coding. The paper consider two criteria for performance of learned coders: the average performance over training data, and a guaranteed performance over all training except for some error probability $P_e$. In both cases the coders are evaluated with respect to redundancy. The paper considers the IID binary case and binary Markov chains. In both cases it is shown that the amount of training data required is very moderate: to code sequences of length $l$ the amount of training data required to beat a universal source coder is $m=K\frac{l}{\log l}$, where the constant in front depends the case considered.
DeepLesion: Automated Deep Mining, Categorization and Detection of Significant Radiology Image Findings using Large-Scale Clinical Lesion Annotations
Oct 10, 2017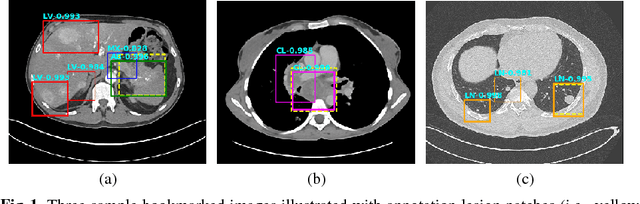
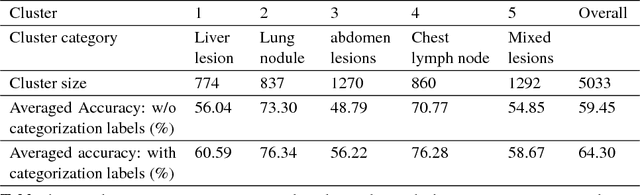
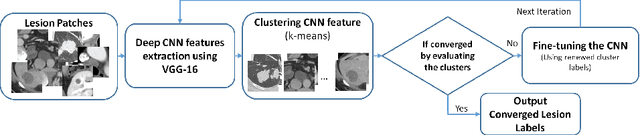
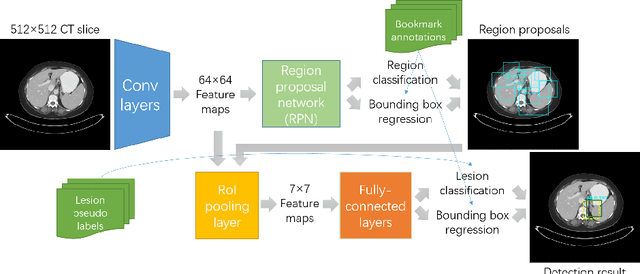
Extracting, harvesting and building large-scale annotated radiological image datasets is a greatly important yet challenging problem. It is also the bottleneck to designing more effective data-hungry computing paradigms (e.g., deep learning) for medical image analysis. Yet, vast amounts of clinical annotations (usually associated with disease image findings and marked using arrows, lines, lesion diameters, segmentation, etc.) have been collected over several decades and stored in hospitals' Picture Archiving and Communication Systems. In this paper, we mine and harvest one major type of clinical annotation data - lesion diameters annotated on bookmarked images - to learn an effective multi-class lesion detector via unsupervised and supervised deep Convolutional Neural Networks (CNN). Our dataset is composed of 33,688 bookmarked radiology images from 10,825 studies of 4,477 unique patients. For every bookmarked image, a bounding box is created to cover the target lesion based on its measured diameters. We categorize the collection of lesions using an unsupervised deep mining scheme to generate clustered pseudo lesion labels. Next, we adopt a regional-CNN method to detect lesions of multiple categories, regardless of missing annotations (normally only one lesion is annotated, despite the presence of multiple co-existing findings). Our integrated mining, categorization and detection framework is validated with promising empirical results, as a scalable, universal or multi-purpose CAD paradigm built upon abundant retrospective medical data. Furthermore, we demonstrate that detection accuracy can be significantly improved by incorporating pseudo lesion labels (e.g., Liver lesion/tumor, Lung nodule/tumor, Abdomen lesions, Chest lymph node and others). This dataset will be made publicly available (under the open science initiative).
Learning Diverse Latent Representations for Improving the Resilience to Adversarial Attacks
Jul 12, 2020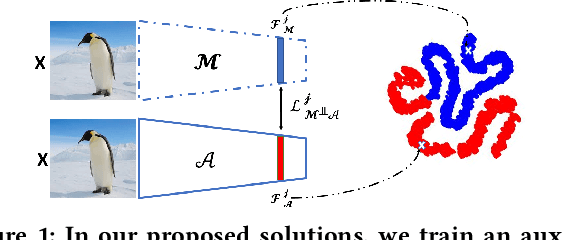
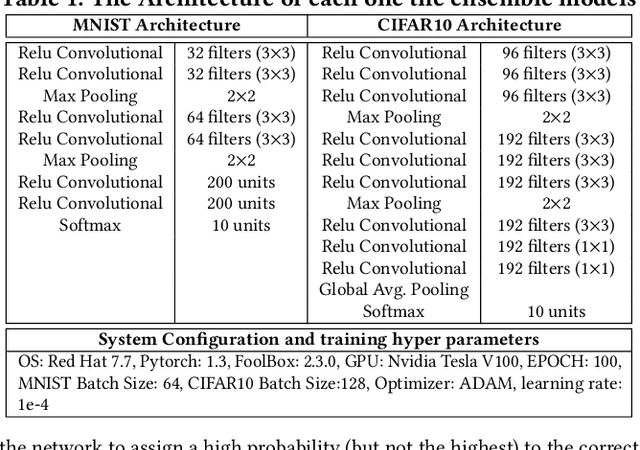
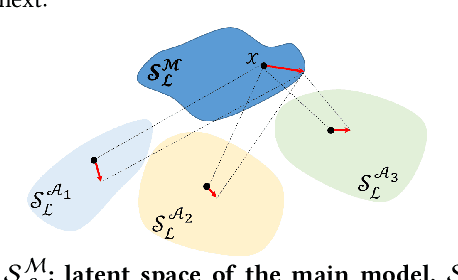
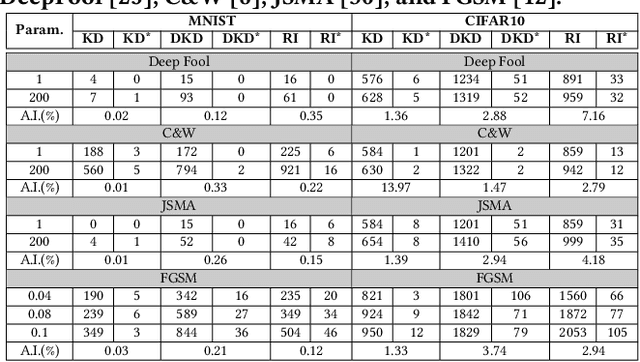
This paper proposes an ensemble learning model that is resistant to adversarial learning attacks. To build resilience, we proposed a training process where each member learns a radically different latent space. Member models are added one at a time to the ensemble. Each model is trained on data set to improve accuracy, while the loss function is regulated by a reverse knowledge distillation, forcing the new member to learn new features and map to a latent space safely distanced from those of existing members. We have evaluated the reliability and performance of the proposed solution on image classification tasks using CIFAR10 and MNIST datasets and show improved performance compared to the state of the art defense methods
Retrain or not retrain? -- efficient pruning methods of deep CNN networks
Feb 12, 2020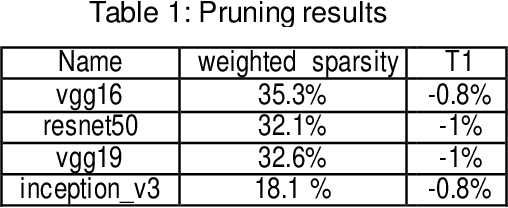
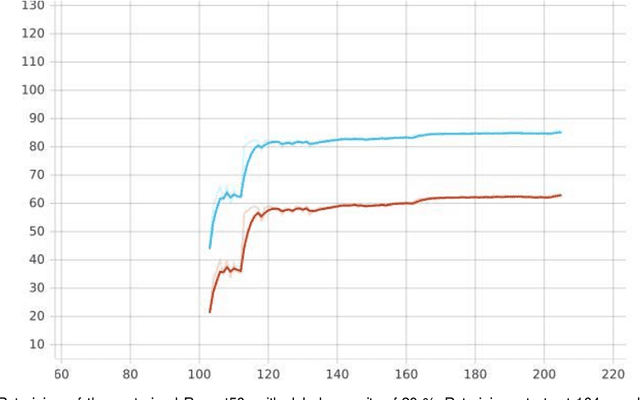
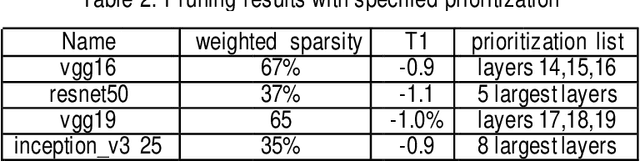
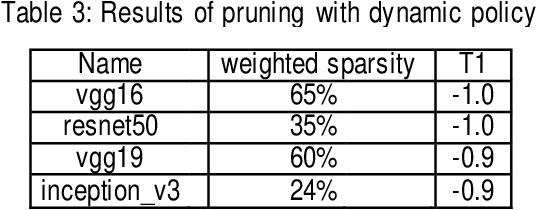
Convolutional neural networks (CNN) play a major role in image processing tasks like image classification, object detection, semantic segmentation. Very often CNN networks have from several to hundred stacked layers with several megabytes of weights. One of the possible methods to reduce complexity and memory footprint is pruning. Pruning is a process of removing weights which connect neurons from two adjacent layers in the network. The process of finding near optimal solution with specified drop in accuracy can be more sophisticated when DL model has higher number of convolutional layers. In the paper few approaches based on retraining and no retraining are described and compared together.
Models Genesis
Apr 09, 2020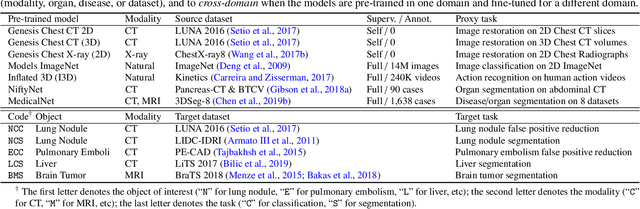
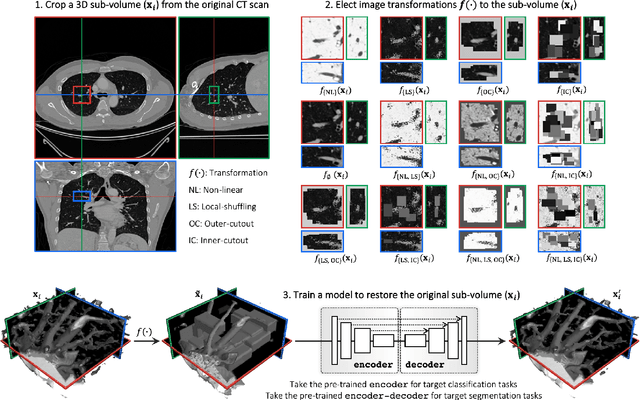
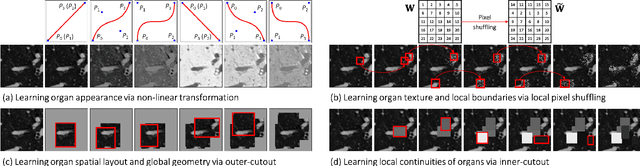
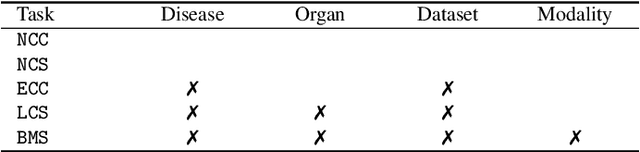
Transfer learning from natural image to medical image has been established as one of the most practical paradigms in deep learning for medical image analysis. To fit this paradigm, however, 3D imaging tasks in the most prominent imaging modalities (e.g., CT and MRI) have to be reformulated and solved in 2D, losing rich 3D anatomical information, thereby inevitably compromising its performance. To overcome this limitation, we have built a set of models, called Generic Autodidactic Models, nicknamed Models Genesis, because they are created ex nihilo (with no manual labeling), self-taught (learnt by self-supervision), and generic (served as source models for generating application-specific target models). Our extensive experiments demonstrate that our Models Genesis significantly outperform learning from scratch in all five target 3D applications covering both segmentation and classification. More importantly, learning a model from scratch simply in 3D may not necessarily yield performance better than transfer learning from ImageNet in 2D, but our Models Genesis consistently top any 2D/2.5D approaches including fine-tuning the models pre-trained from ImageNet as well as fine-tuning the 2D versions of our Models Genesis, confirming the importance of 3D anatomical information and significance of Models Genesis for 3D medical imaging. This performance is attributed to our unified self-supervised learning framework, built on a simple yet powerful observation: the sophisticated and recurrent anatomy in medical images can serve as strong yet free supervision signals for deep models to learn common anatomical representation automatically via self-supervision. As open science, all codes and pre-trained Models Genesis are available at https://github.com/MrGiovanni/ModelsGenesis
Neural method for Explicit Mapping of Quasi-curvature Locally Linear Embedding in image retrieval
Mar 11, 2017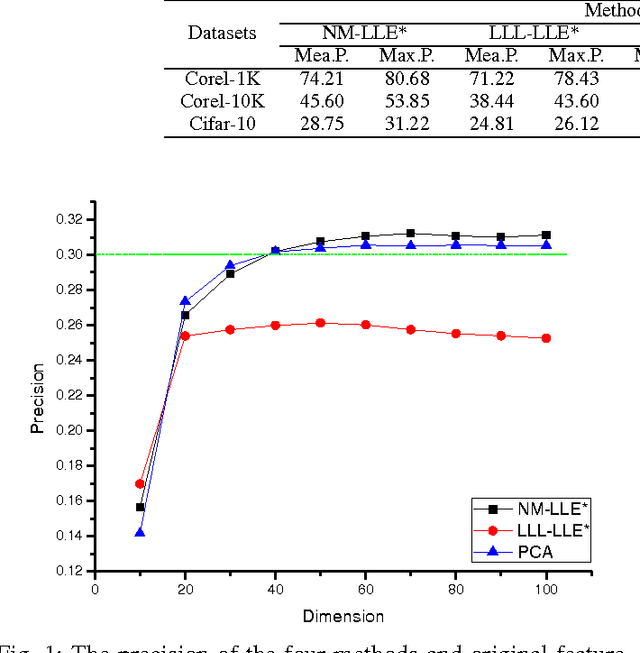
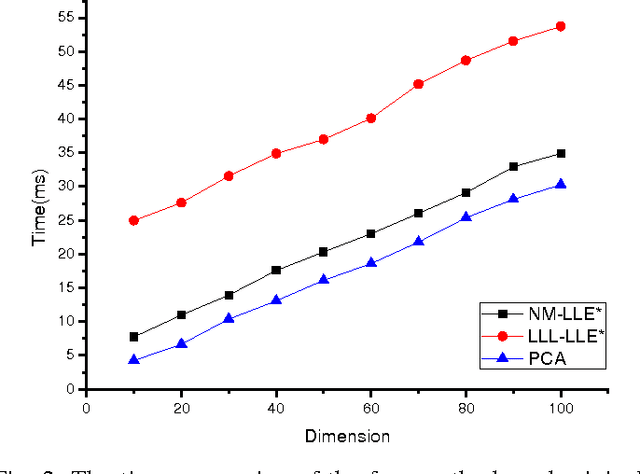
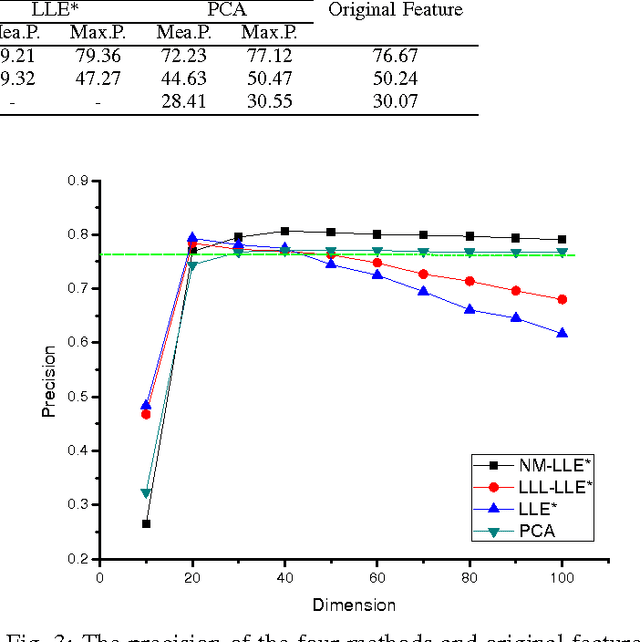
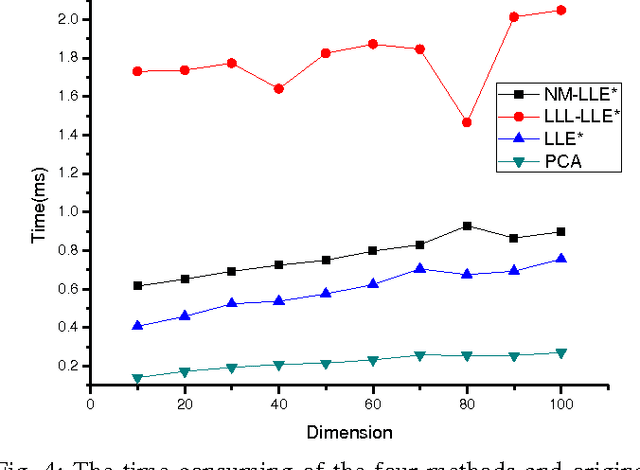
This paper proposed a new explicit nonlinear dimensionality reduction using neural networks for image retrieval tasks. We first proposed a Quasi-curvature Locally Linear Embedding (QLLE) for training set. QLLE guarantees the linear criterion in neighborhood of each sample. Then, a neural method (NM) is proposed for out-of-sample problem. Combining QLLE and NM, we provide a explicit nonlinear dimensionality reduction approach for efficient image retrieval. The experimental results in three benchmark datasets illustrate that our method can get better performance than other state-of-the-art out-of-sample methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge