"Image": models, code, and papers
Unsupervised Learning of Visual Features by Contrasting Cluster Assignments
Jun 18, 2020
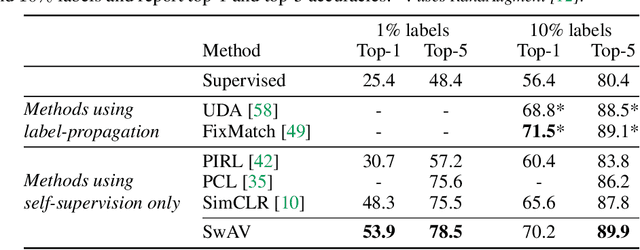
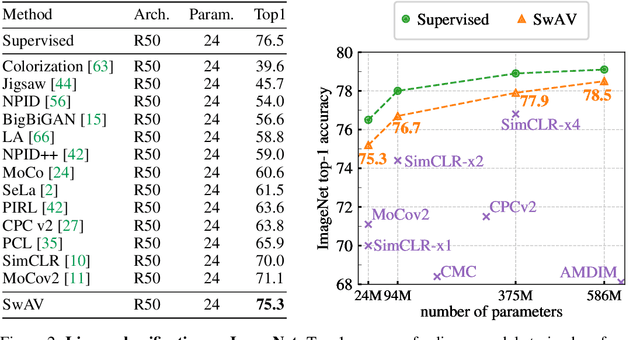

Unsupervised image representations have significantly reduced the gap with supervised pretraining, notably with the recent achievements of contrastive learning methods. These contrastive methods typically work online and rely on a large number of explicit pairwise feature comparisons, which is computationally challenging. In this paper, we propose an online algorithm, SwAV, that takes advantage of contrastive methods without requiring to compute pairwise comparisons. Specifically, our method simultaneously clusters the data while enforcing consistency between cluster assignments produced for different augmentations (or views) of the same image, instead of comparing features directly as in contrastive learning. Simply put, we use a swapped prediction mechanism where we predict the cluster assignment of a view from the representation of another view. Our method can be trained with large and small batches and can scale to unlimited amounts of data. Compared to previous contrastive methods, our method is more memory efficient since it does not require a large memory bank or a special momentum network. In addition, we also propose a new data augmentation strategy, multi-crop, that uses a mix of views with different resolutions in place of two full-resolution views, without increasing the memory or compute requirements much. We validate our findings by achieving 75.3% top-1 accuracy on ImageNet with ResNet-50, as well as surpassing supervised pretraining on all the considered transfer tasks.
Deep N-ary Error Correcting Output Codes
Sep 22, 2020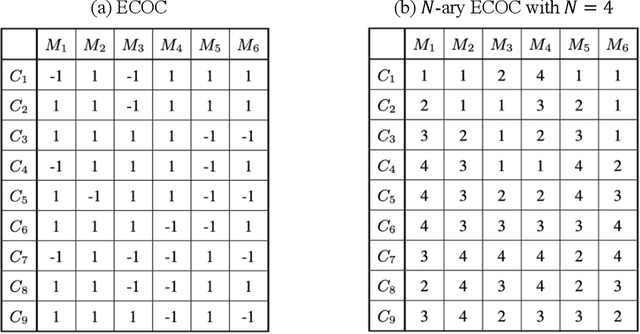
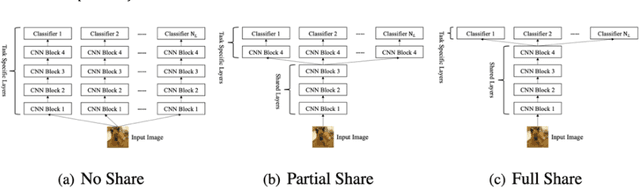
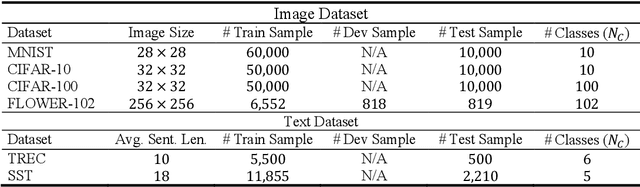
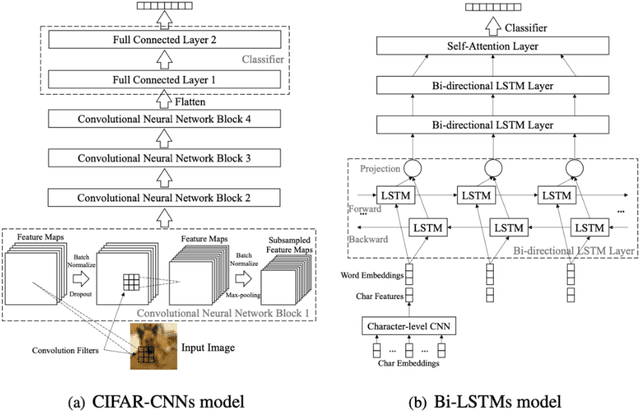
Ensemble learning consistently improves the performance of multi-class classification through aggregating a series of base classifiers. To this end, data-independent ensemble methods like Error Correcting Output Codes (ECOC) attract increasing attention due to its easiness of implementation and parallelization. Specifically, traditional ECOCs and its general extension N-ary ECOC decompose the original multi-class classification problem into a series of independent simpler classification subproblems. Unfortunately, integrating ECOCs, especially N-ary ECOC with deep neural networks, termed as deep N-ary ECOC, is not straightforward and yet fully exploited in the literature, due to the high expense of training base learners. To facilitate the training of N-ary ECOC with deep learning base learners, we further propose three different variants of parameter sharing architectures for deep N-ary ECOC. To verify the generalization ability of deep N-ary ECOC, we conduct experiments by varying the backbone with different deep neural network architectures for both image and text classification tasks. Furthermore, extensive ablation studies on deep N-ary ECOC show its superior performance over other deep data-independent ensemble methods.
Investigation of REFINED CNN ensemble learning for anti-cancer drug sensitivity prediction
Sep 09, 2020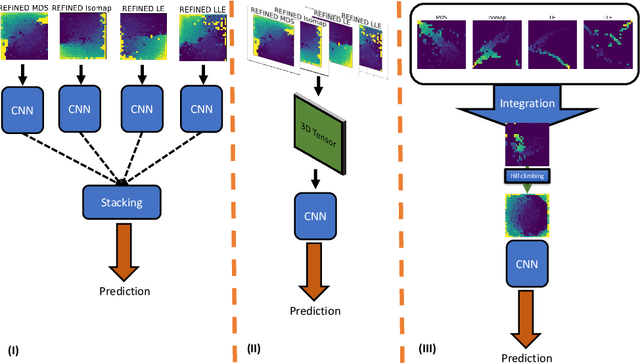
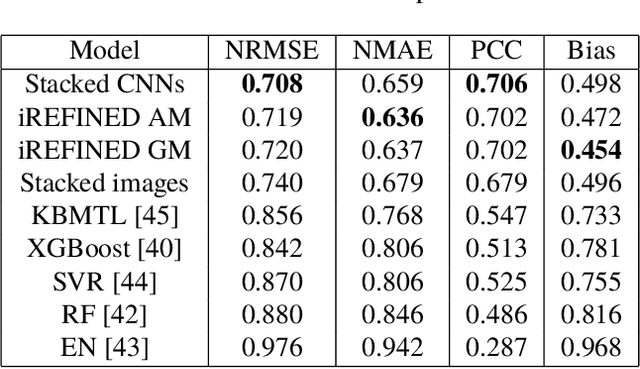
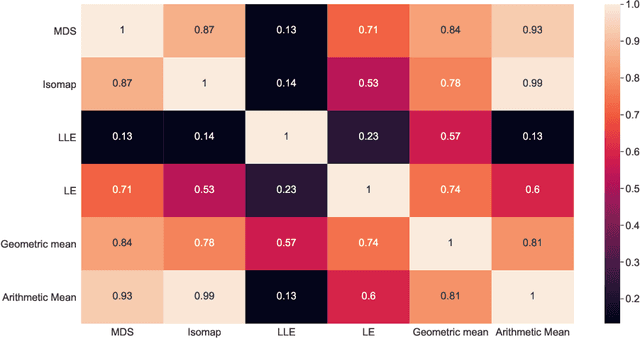
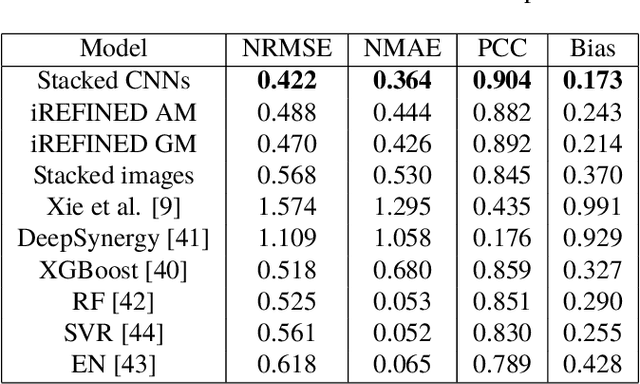
Anti-cancer drug sensitivity prediction using deep learning models for individual cell line is a significant challenge in personalized medicine. REFINED (REpresentation of Features as Images with NEighborhood Dependencies) CNN (Convolutional Neural Network) based models have shown promising results in drug sensitivity prediction. The primary idea behind REFINED CNN is representing high dimensional vectors as compact images with spatial correlations that can benefit from convolutional neural network architectures. However, the mapping from a vector to a compact 2D image is not unique due to variations in considered distance measures and neighborhoods. In this article, we consider predictions based on ensembles built from such mappings that can improve upon the best single REFINED CNN model prediction. Results illustrated using NCI60 and NCIALMANAC databases shows that the ensemble approaches can provide significant performance improvement as compared to individual models. We further illustrate that a single mapping created from the amalgamation of the different mappings can provide performance similar to stacking ensemble but with significantly lower computational complexity.
Improved Trainable Calibration Method for Neural Networks on Medical Imaging Classification
Sep 09, 2020
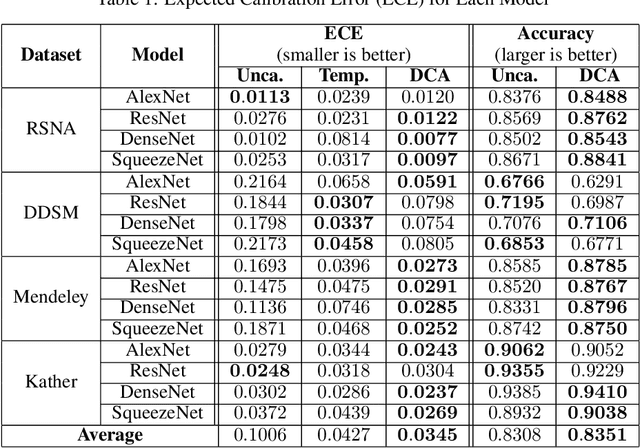
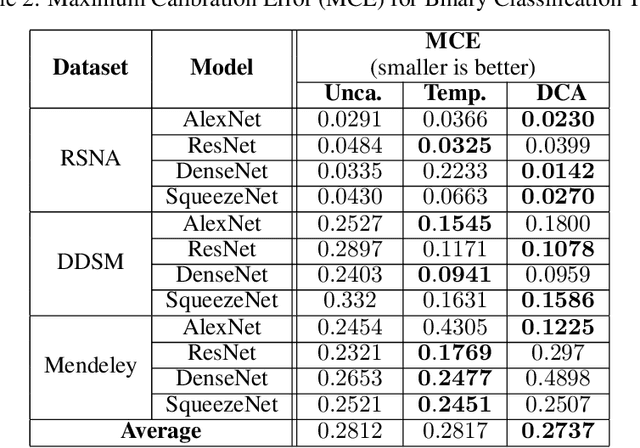
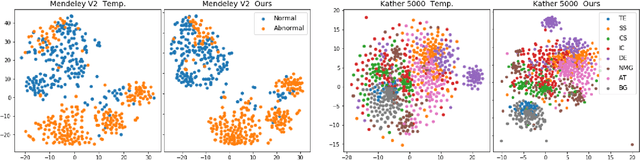
Recent works have shown that deep neural networks can achieve super-human performance in a wide range of image classification tasks in the medical imaging domain. However, these works have primarily focused on classification accuracy, ignoring the important role of uncertainty quantification. Empirically, neural networks are often miscalibrated and overconfident in their predictions. This miscalibration could be problematic in any automatic decision-making system, but we focus on the medical field in which neural network miscalibration has the potential to lead to significant treatment errors. We propose a novel calibration approach that maintains the overall classification accuracy while significantly improving model calibration. The proposed approach is based on expected calibration error, which is a common metric for quantifying miscalibration. Our approach can be easily integrated into any classification task as an auxiliary loss term, thus not requiring an explicit training round for calibration. We show that our approach reduces calibration error significantly across various architectures and datasets.
A Point Set Generation Network for 3D Object Reconstruction from a Single Image
Dec 07, 2016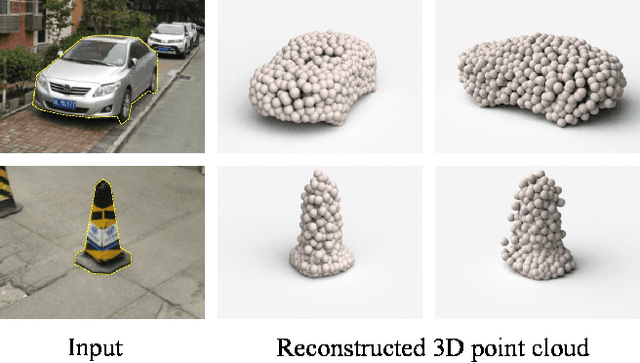
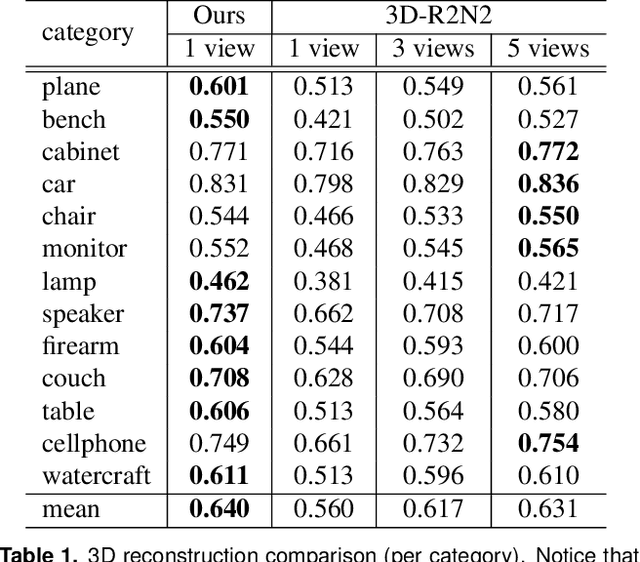
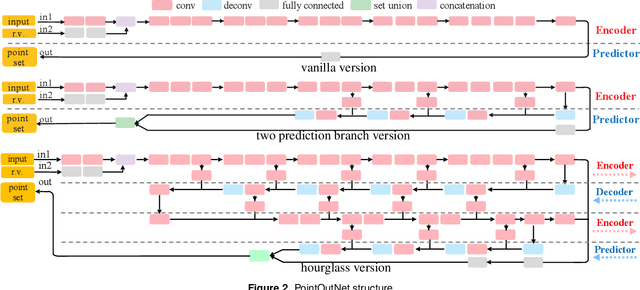
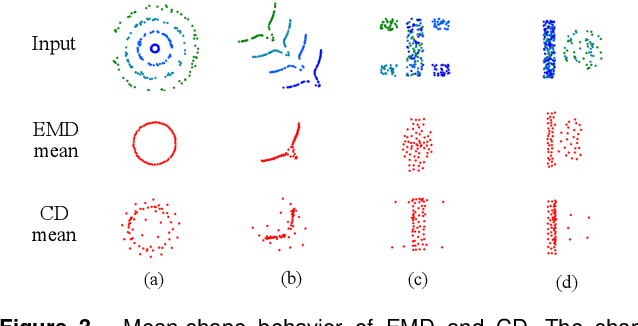
Generation of 3D data by deep neural network has been attracting increasing attention in the research community. The majority of extant works resort to regular representations such as volumetric grids or collection of images; however, these representations obscure the natural invariance of 3D shapes under geometric transformations and also suffer from a number of other issues. In this paper we address the problem of 3D reconstruction from a single image, generating a straight-forward form of output -- point cloud coordinates. Along with this problem arises a unique and interesting issue, that the groundtruth shape for an input image may be ambiguous. Driven by this unorthodox output form and the inherent ambiguity in groundtruth, we design architecture, loss function and learning paradigm that are novel and effective. Our final solution is a conditional shape sampler, capable of predicting multiple plausible 3D point clouds from an input image. In experiments not only can our system outperform state-of-the-art methods on single image based 3d reconstruction benchmarks; but it also shows a strong performance for 3d shape completion and promising ability in making multiple plausible predictions.
Spatial Action Maps for Mobile Manipulation
Apr 20, 2020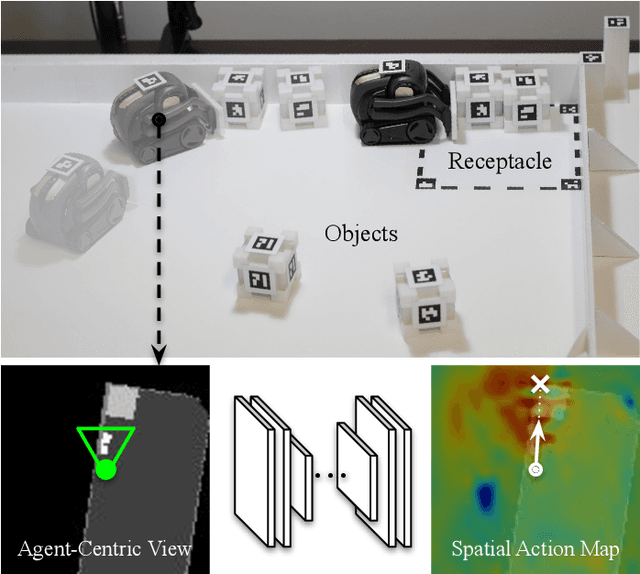
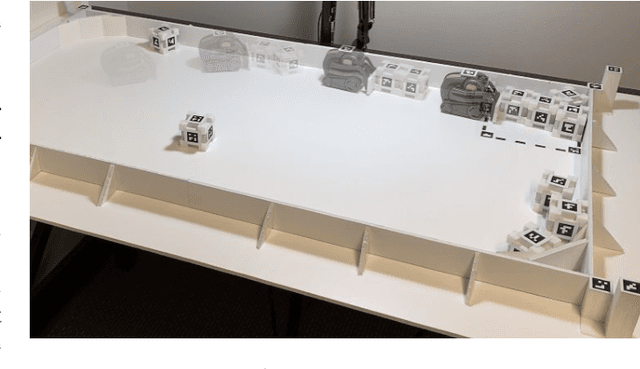


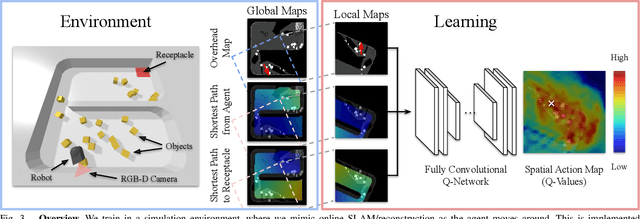
This paper proposes a new action representation for learning to perform complex mobile manipulation tasks. In a typical deep Q-learning setup, a convolutional neural network (ConvNet) is trained to map from an image representing the current state (e.g., a birds-eye view of a SLAM reconstruction of the scene) to predicted Q-values for a small set of steering command actions (step forward, turn right, turn left, etc.). Instead, we propose an action representation in the same domain as the state: "spatial action maps." In our proposal, the set of possible actions is represented by pixels of an image, where each pixel represents a trajectory to the corresponding scene location along a shortest path through obstacles of the partially reconstructed scene. A significant advantage of this approach is that the spatial position of each state-action value prediction represents a local milestone (local end-point) for the agent's policy, which may be easily recognizable in local visual patterns of the state image. A second advantage is that atomic actions can perform long-range plans (follow the shortest path to a point on the other side of the scene), and thus it is simpler to learn complex behaviors with a deep Q-network. A third advantage is that we can use a fully convolutional network (FCN) with skip connections to learn the mapping from state images to pixel-aligned action images efficiently. During experiments with a robot that learns to push objects to a goal location, we find that policies learned with this proposed action representation achieve significantly better performance than traditional alternatives.
Automatic phantom test pattern classification through transfer learning with deep neural networks
Jan 22, 2020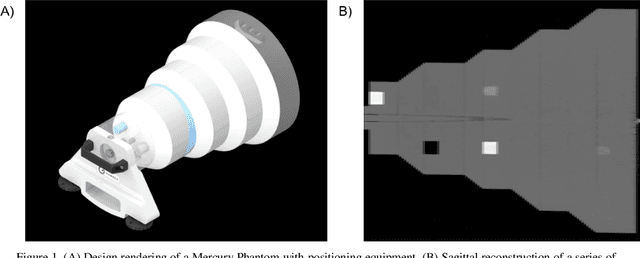
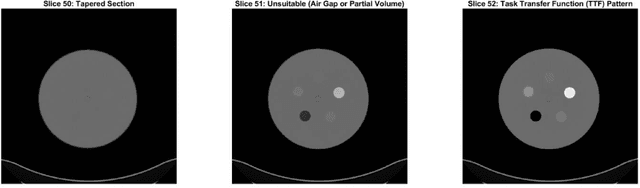
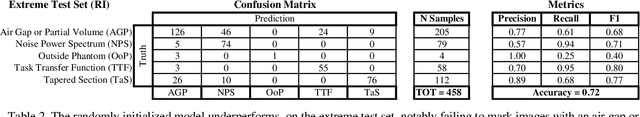
Imaging phantoms are test patterns used to measure image quality in computer tomography (CT) systems. A new phantom platform (Mercury Phantom, Gammex) provides test patterns for estimating the task transfer function (TTF) or noise power spectrum (NPF) and simulates different patient sizes. Determining which image slices are suitable for analysis currently requires manual annotation of these patterns by an expert, as subtle defects may make an image unsuitable for measurement. We propose a method of automatically classifying these test patterns in a series of phantom images using deep learning techniques. By adapting a convolutional neural network based on the VGG19 architecture with weights trained on ImageNet, we use transfer learning to produce a classifier for this domain. The classifier is trained and evaluated with over 3,500 phantom images acquired at a university medical center. Input channels for color images are successfully adapted to convey contextual information for phantom images. A series of ablation studies are employed to verify design aspects of the classifier and evaluate its performance under varying training conditions. Our solution makes extensive use of image augmentation to produce a classifier that accurately classifies typical phantom images with 98% accuracy, while maintaining as much as 86% accuracy when the phantom is improperly imaged.
ROSE: A Retinal OCT-Angiography Vessel Segmentation Dataset and New Model
Jul 10, 2020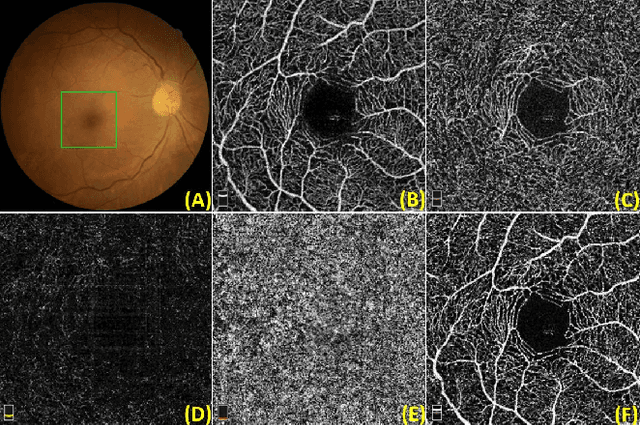
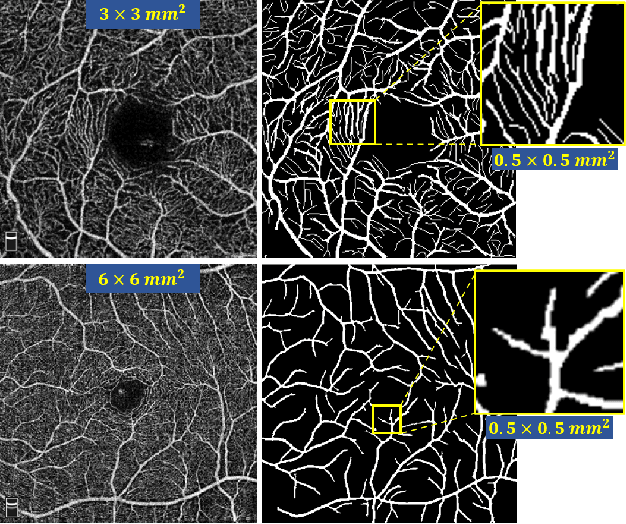
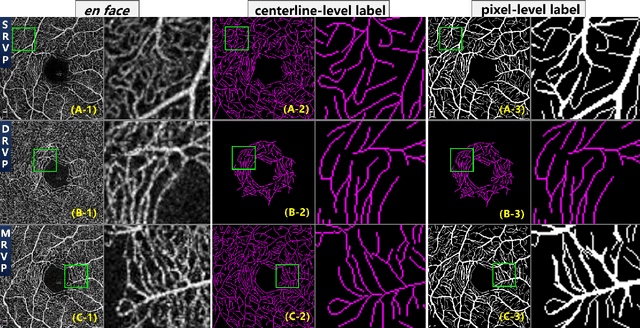
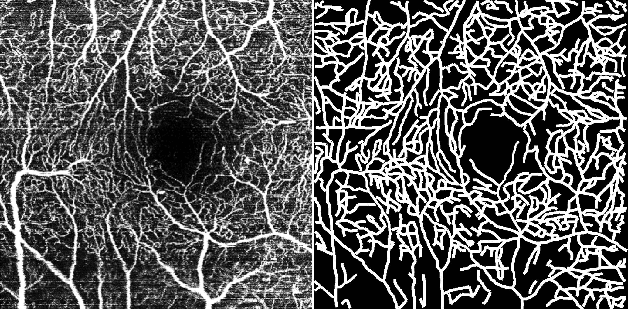
Optical Coherence Tomography Angiography (OCT-A) is a non-invasive imaging technique, and has been increasingly used to image the retinal vasculature at capillary level resolution. However, automated segmentation of retinal vessels in OCT-A has been under-studied due to various challenges such as low capillary visibility and high vessel complexity, despite its significance in understanding many eye-related diseases. In addition, there is no publicly available OCT-A dataset with manually graded vessels for training and validation. To address these issues, for the first time in the field of retinal image analysis we construct a dedicated Retinal OCT-A SEgmentation dataset (ROSE), which consists of 229 OCT-A images with vessel annotations at either centerline-level or pixel level. This dataset has been released for public access to assist researchers in the community in undertaking research in related topics. Secondly, we propose a novel Split-based Coarse-to-Fine vessel segmentation network (SCF-Net), with the ability to detect thick and thin vessels separately. In the SCF-Net, a split-based coarse segmentation (SCS) module is first introduced to produce a preliminary confidence map of vessels, and a split-based refinement (SRN) module is then used to optimize the shape/contour of the retinal microvasculature. Thirdly, we perform a thorough evaluation of the state-of-the-art vessel segmentation models and our SCF-Net on the proposed ROSE dataset. The experimental results demonstrate that our SCF-Net yields better vessel segmentation performance in OCT-A than both traditional methods and other deep learning methods.
Detection of prostate cancer in whole-slide images through end-to-end training with image-level labels
Jun 05, 2020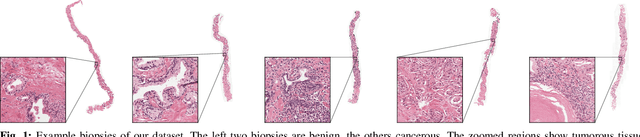
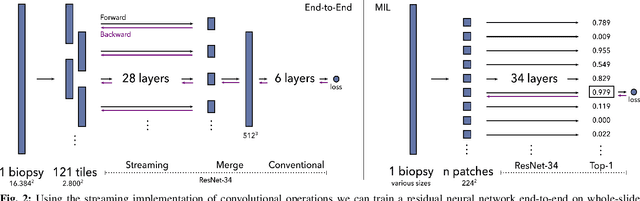
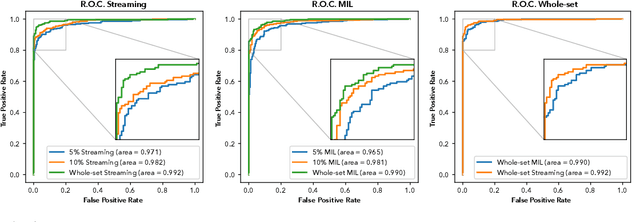

Prostate cancer is the most prevalent cancer among men in Western countries, with 1.1 million new diagnoses every year. The gold standard for the diagnosis of prostate cancer is a pathologists' evaluation of prostate tissue. To potentially assist pathologists deep-learning-based cancer detection systems have been developed. Many of the state-of-the-art models are patch-based convolutional neural networks, as the use of entire scanned slides is hampered by memory limitations on accelerator cards. Patch-based systems typically require detailed, pixel-level annotations for effective training. However, such annotations are seldom readily available, in contrast to the clinical reports of pathologists, which contain slide-level labels. As such, developing algorithms which do not require manual pixel-wise annotations, but can learn using only the clinical report would be a significant advancement for the field. In this paper, we propose to use a streaming implementation of convolutional layers, to train a modern CNN (ResNet-34) with 21 million parameters end-to-end on 4712 prostate biopsies. The method enables the use of entire biopsy images at high-resolution directly by reducing the GPU memory requirements by 2.4 TB. We show that modern CNNs, trained using our streaming approach, can extract meaningful features from high-resolution images without additional heuristics, reaching similar performance as state-of-the-art patch-based and multiple-instance learning methods. By circumventing the need for manual annotations, this approach can function as a blueprint for other tasks in histopathological diagnosis. The source code to reproduce the streaming models is available at https://github.com/DIAGNijmegen/pathology-streaming-pipeline .
Blind Super-Resolution Kernel Estimation using an Internal-GAN
Sep 18, 2019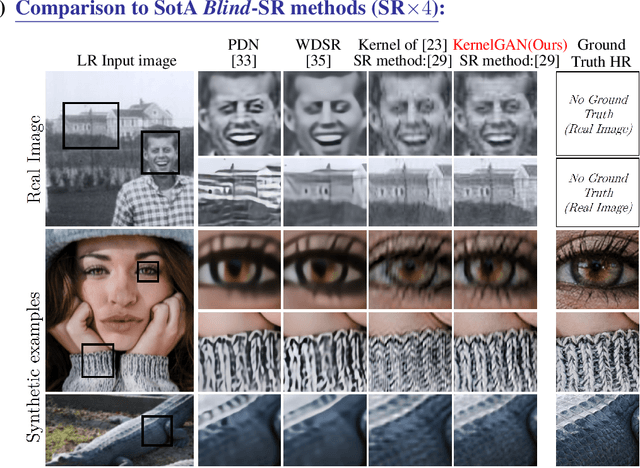
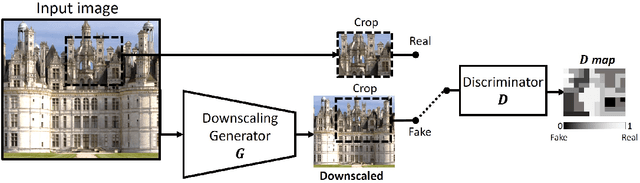
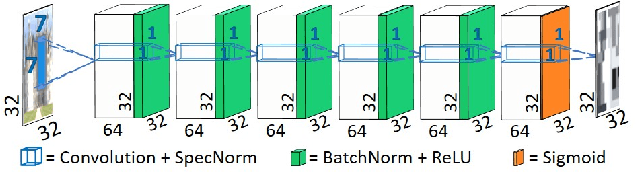
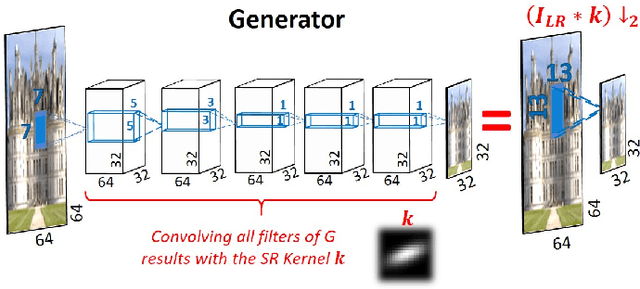
Super resolution (SR) methods typically assume that the low-resolution (LR) image was downscaled from the unknown high-resolution (HR) image by a fixed 'ideal' downscaling kernel (e.g. Bicubic downscaling). However, this is rarely the case in real LR images, in contrast to synthetically generated SR datasets. When the assumed downscaling kernel deviates from the true one, the performance of SR methods significantly deteriorates. This gave rise to Blind-SR - namely, SR when the downscaling kernel ("SR-kernel") is unknown. It was further shown that the true SR-kernel is the one that maximizes the recurrence of patches across scales of the LR image. In this paper we show how this powerful cross-scale recurrence property can be realized using Deep Internal Learning. We introduce "KernelGAN", an image-specific Internal-GAN, which trains solely on the LR test image at test time, and learns its internal distribution of patches. Its Generator is trained to produce a downscaled version of the LR test image, such that its Discriminator cannot distinguish between the patch distribution of the downscaled image, and the patch distribution of the original LR image. The Generator, once trained, constitutes the downscaling operation with the correct image-specific SR-kernel. KernelGAN is fully unsupervised, requires no training data other than the input image itself, and leads to state-of-the-art results in Blind-SR when plugged into existing SR algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge