"Image": models, code, and papers
DC-UNet: Rethinking the U-Net Architecture with Dual Channel Efficient CNN for Medical Images Segmentation
May 31, 2020
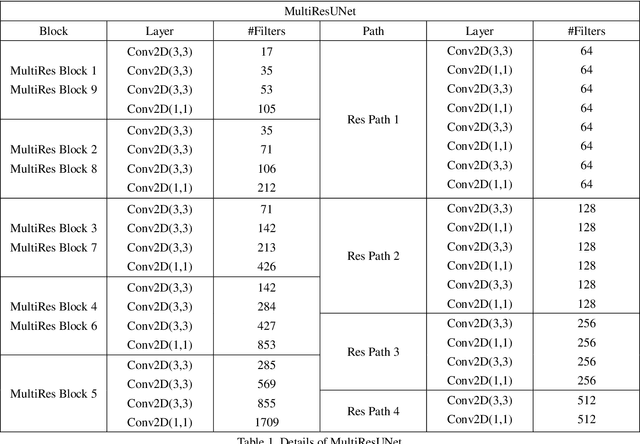
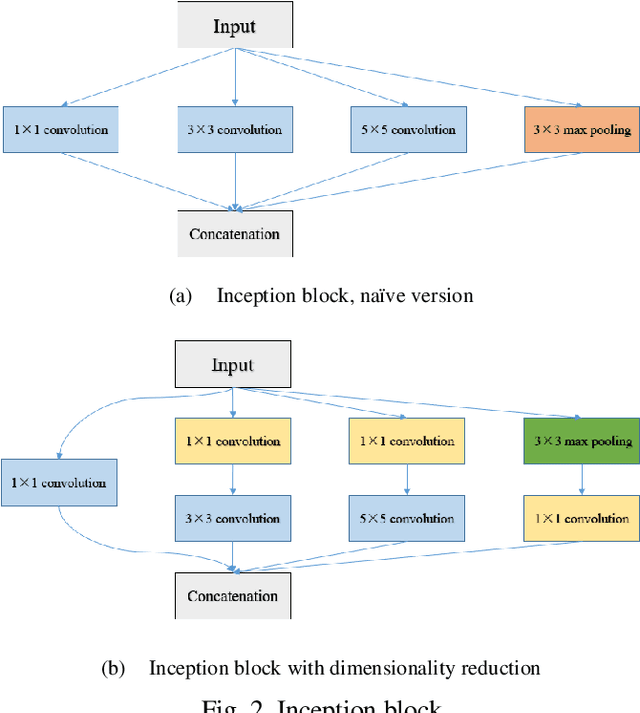
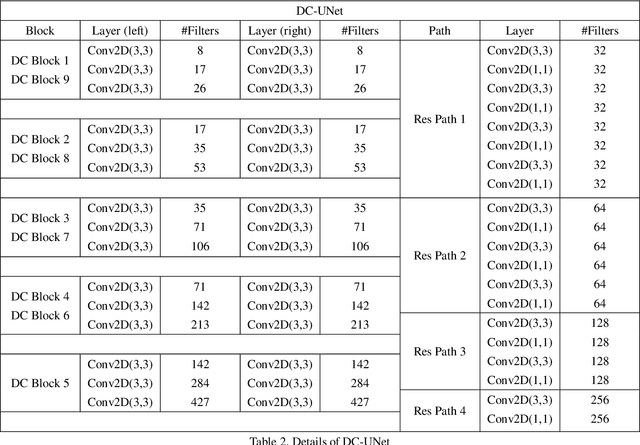
Recently, deep learning has become much more popular in computer vision area. The Convolution Neural Network (CNN) has brought a breakthrough in images segmentation areas, especially, for medical images. In this regard, U-Net is the predominant approach to medical image segmentation task. The U-Net not only performs well in segmenting multimodal medical images generally, but also in some tough cases of them. However, we found that the classical U-Net architecture has limitation in several aspects. Therefore, we applied modifications: 1) designed efficient CNN architecture to replace encoder and decoder, 2) applied residual module to replace skip connection between encoder and decoder to improve based on the-state-of-the-art U-Net model. Following these modifications, we designed a novel architecture--DC-UNet, as a potential successor to the U-Net architecture. We created a new effective CNN architecture and build the DC-UNet based on this CNN. We have evaluated our model on three datasets with tough cases and have obtained a relative improvement in performance of 2.90%, 1.49% and 11.42% respectively compared with classical U-Net. In addition, we used the Tanimoto similarity to replace the Jaccard similarity for gray-to-gray image comparisons.
Label-Free Segmentation of COVID-19 Lesions in Lung CT
Sep 15, 2020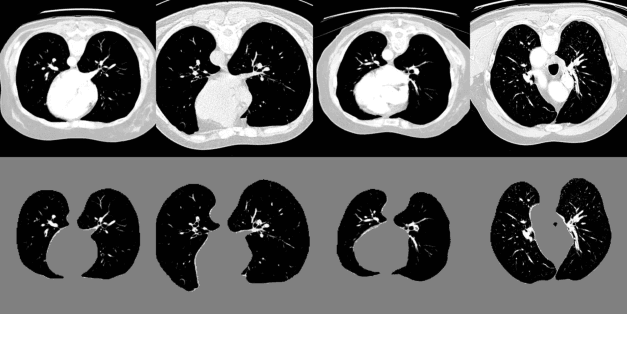
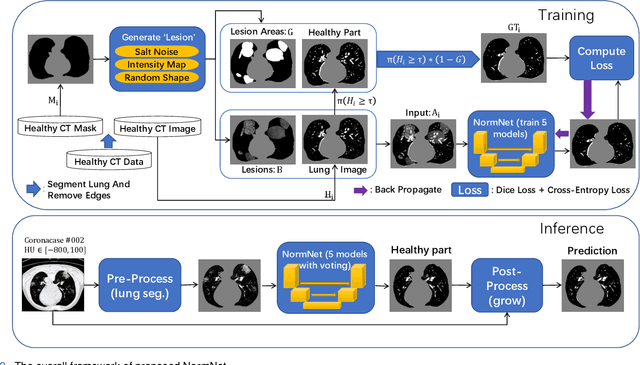
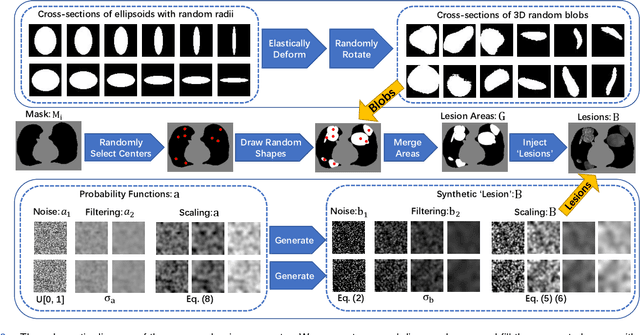
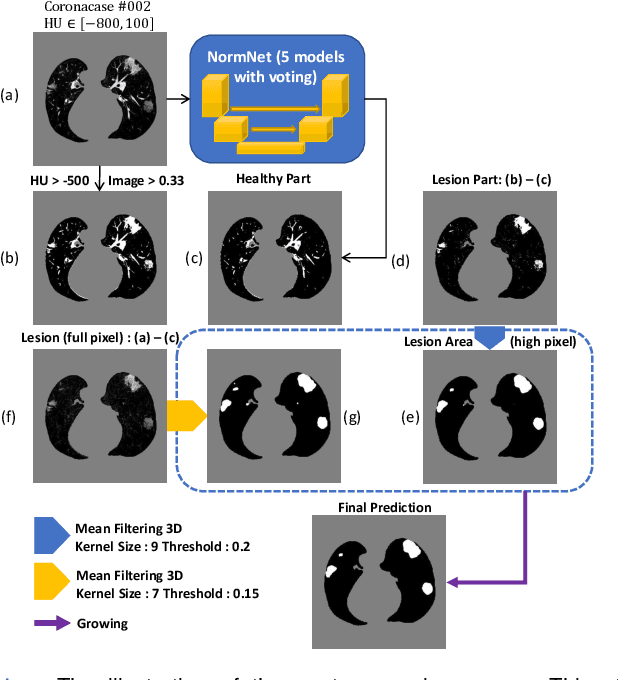
Scarcity of annotated images hampers the building of automated solution for reliable COVID-19 diagnosis and evaluation from CT. To alleviate the burden of data annotation, we herein present a label-free approach for segmenting COVID-19 lesions in CT via pixel-level anomaly modeling that mines out the relevant knowledge from normal CT lung scans. Our modeling is inspired by the observation that the parts of tracheae and vessels, which lay in the high-intensity range where lesions belong to, exhibit strong patterns. To facilitate the learning of such patterns at a pixel level, we synthesize `lesions' using a set of surprisingly simple operations and insert the synthesized `lesions' into normal CT lung scans to form training pairs, from which we learn a normalcy-converting network (NormNet) that turns an 'abnormal' image back to normal. Our experiments on three different datasets validate the effectiveness of NormNet, which conspicuously outperforms a variety of unsupervised anomaly detection (UAD) methods.
Langevin Cooling for Domain Translation
Aug 31, 2020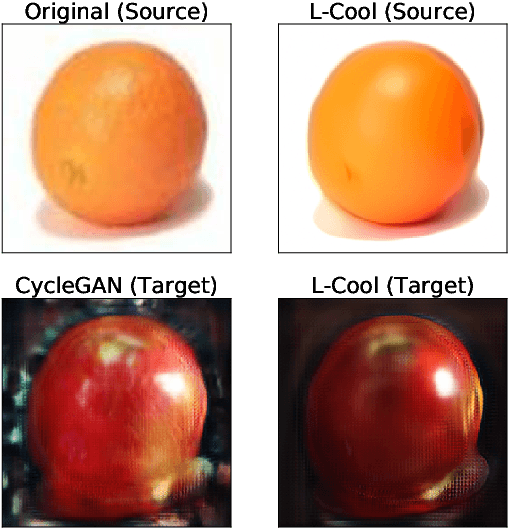
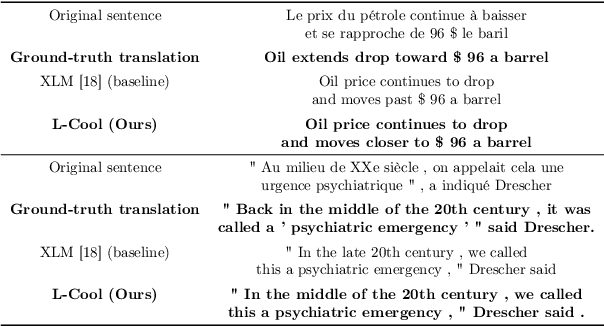
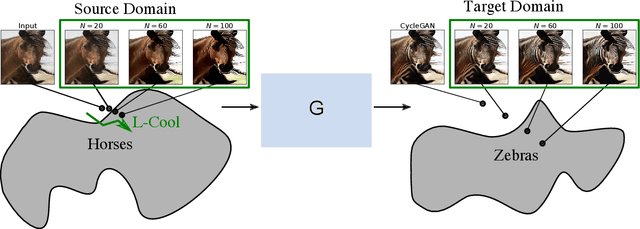
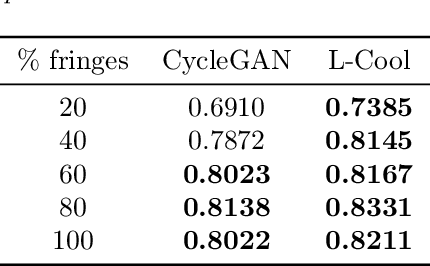
Domain translation is the task of finding correspondence between two domains. Several Deep Neural Network (DNN) models, e.g., CycleGAN and cross-lingual language models, have shown remarkable successes on this task under the unsupervised setting---the mappings between the domains are learned from two independent sets of training data in both domains (without paired samples). However, those methods typically do not perform well on a significant proportion of test samples. In this paper, we hypothesize that many of such unsuccessful samples lie at the fringe---relatively low-density areas---of data distribution, where the DNN was not trained very well, and propose to perform Langevin dynamics to bring such fringe samples towards high density areas. We demonstrate qualitatively and quantitatively that our strategy, called Langevin Cooling (L-Cool), enhances state-of-the-art methods in image translation and language translation tasks.
Cost-Effective Active Learning for Deep Image Classification
Jan 13, 2017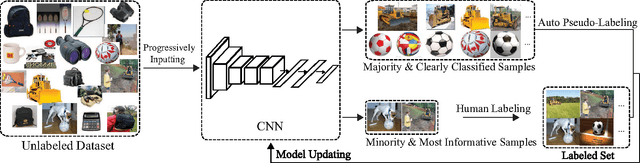

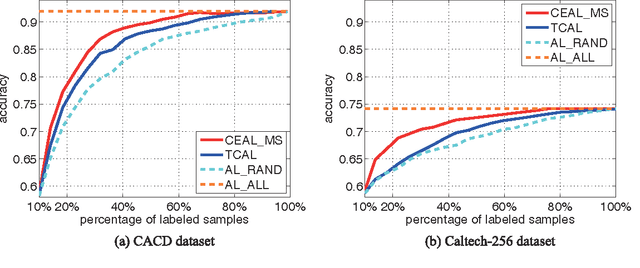
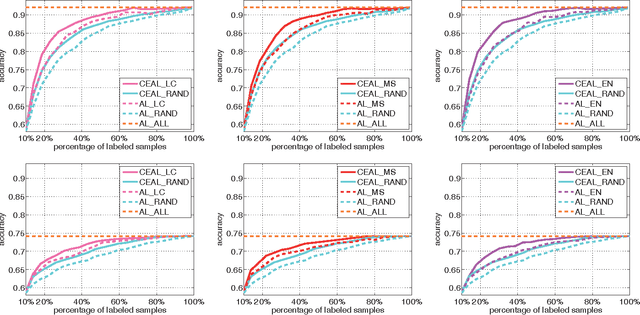
Recent successes in learning-based image classification, however, heavily rely on the large number of annotated training samples, which may require considerable human efforts. In this paper, we propose a novel active learning framework, which is capable of building a competitive classifier with optimal feature representation via a limited amount of labeled training instances in an incremental learning manner. Our approach advances the existing active learning methods in two aspects. First, we incorporate deep convolutional neural networks into active learning. Through the properly designed framework, the feature representation and the classifier can be simultaneously updated with progressively annotated informative samples. Second, we present a cost-effective sample selection strategy to improve the classification performance with less manual annotations. Unlike traditional methods focusing on only the uncertain samples of low prediction confidence, we especially discover the large amount of high confidence samples from the unlabeled set for feature learning. Specifically, these high confidence samples are automatically selected and iteratively assigned pseudo-labels. We thus call our framework "Cost-Effective Active Learning" (CEAL) standing for the two advantages.Extensive experiments demonstrate that the proposed CEAL framework can achieve promising results on two challenging image classification datasets, i.e., face recognition on CACD database [1] and object categorization on Caltech-256 [2].
Developing and Deploying Machine Learning Pipelines against Real-Time Image Streams from the PACS
Apr 20, 2020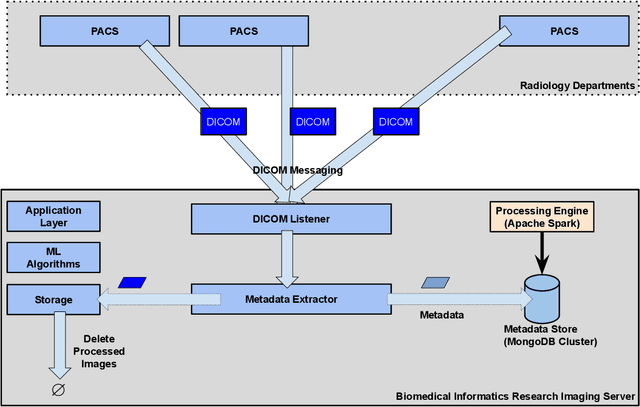
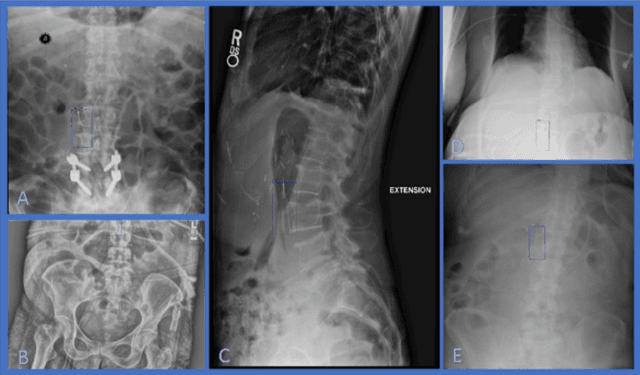
Executing machine learning (ML) pipelines on radiology images is hard due to limited computing resources in clinical environments, whereas running them in research clusters in real-time requires efficient data transfer capabilities. We propose Niffler, an integrated ML framework that runs in research clusters that receives radiology images in real-time from hospitals' Picture Archiving and Communication Systems (PACS). Niffler consists of an inter-domain data streaming approach that exploits the Digital Imaging and Communications in Medicine (DICOM) protocol to fetch data from the PACS to the data processing servers for executing the ML pipelines. It provides metadata extraction capabilities and Application programming interfaces (APIs) to apply filters on the DICOM images and run the ML pipelines. The outcomes of the ML pipelines can then be shared back with the end-users in a de-identified manner. Evaluations on the Niffler prototype highlight the feasibility and efficiency in running the ML pipelines in real-time from a research cluster on the images received in real-time from hospital PACS.
Deep Residual 3D U-Net for Joint Segmentation and Texture Classification of Nodules in Lung
Jun 26, 2020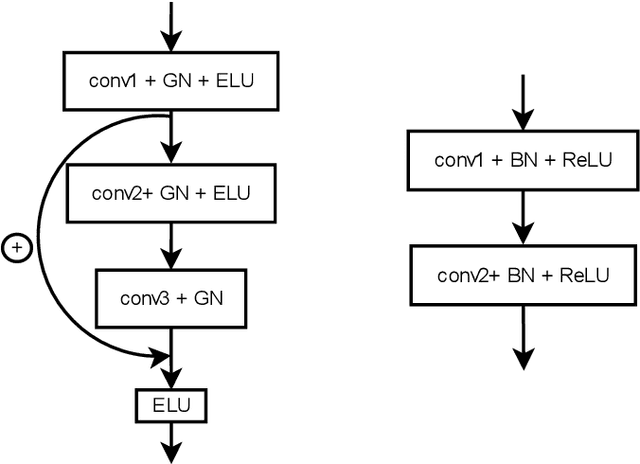
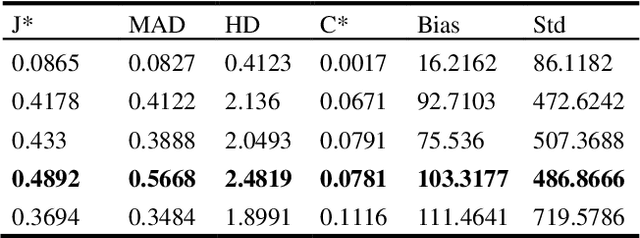
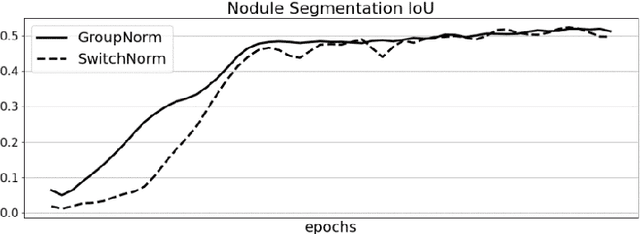
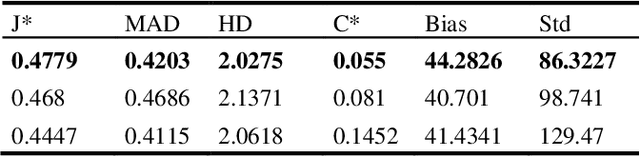
In this work we present a method for lung nodules segmentation, their texture classification and subsequent follow-up recommendation from the CT image of lung. Our method consists of neural network model based on popular U-Net architecture family but modified for the joint nodule segmentation and its texture classification tasks and an ensemble-based model for the follow-up recommendation. This solution was evaluated within the LNDb medical imaging challenge and produced the best nodule segmentation result on the final leaderboard.
* 10 pages, 5 figures, 2 tables, accepted for publication at ICIAR 2020 (LNDb Grand Challenge)
Switchable Deep Beamformer
Aug 31, 2020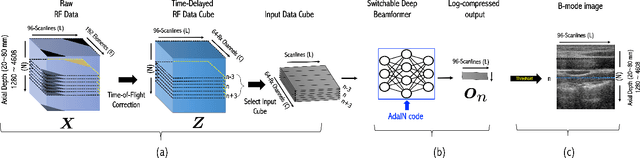
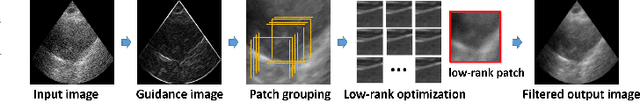
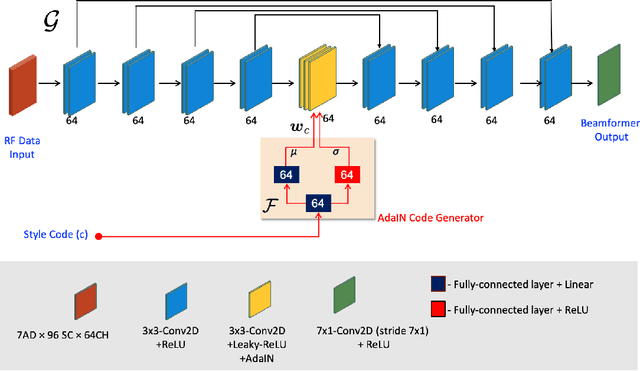
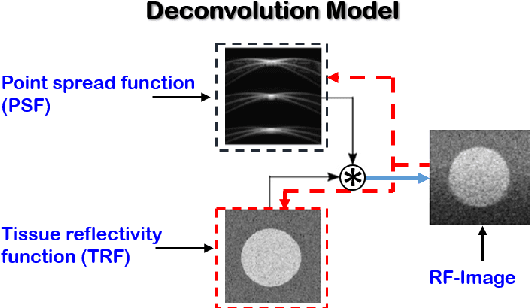

Recent proposals of deep beamformers using deep neural networks have attracted significant attention as computational efficient alternatives to adaptive and compressive beamformers. Moreover, deep beamformers are versatile in that image post-processing algorithms can be combined with the beamforming. Unfortunately, in the current technology, a separate beamformer should be trained and stored for each application, demanding significant scanner resources. To address this problem, here we propose a {\em switchable} deep beamformer that can produce various types of output such as DAS, speckle removal, deconvolution, etc., using a single network with a simple switch. In particular, the switch is implemented through Adaptive Instanace Normalization (AdaIN) layers, so that various output can be generated by merely changing the AdaIN code. Experimental results using B-mode focused ultrasound confirm the flexibility and efficacy of the proposed methods for various applications.
SegET: Deep Neural Network with Rich Contextual Features for Cellular Structures Segmentation in Electron Tomography Image
Nov 28, 2018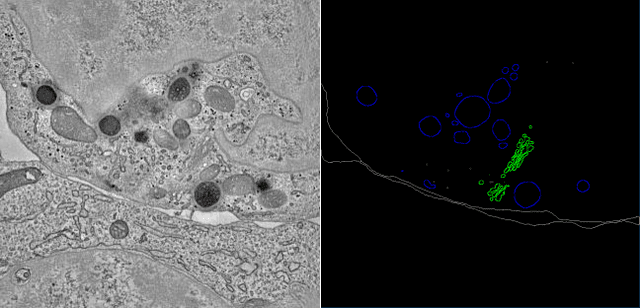
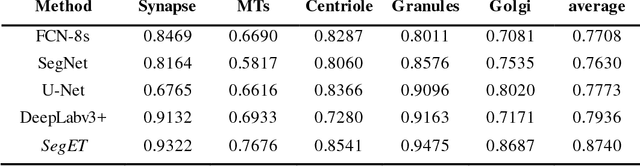
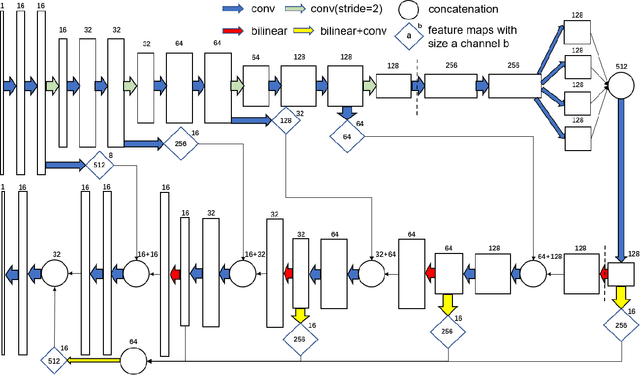
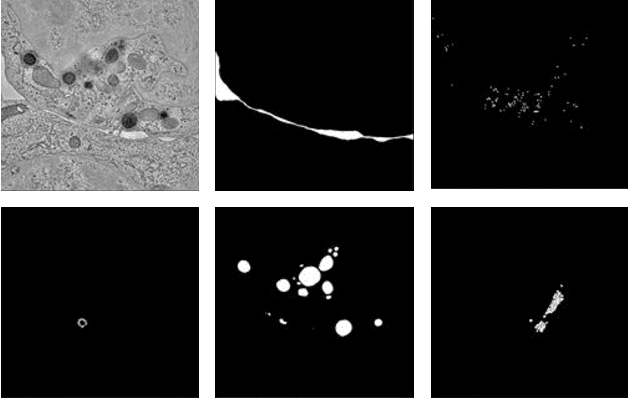
Electron tomography (ET) allows high-resolution reconstructions of macromolecular complexes at nearnative state. Cellular structures segmentation in the reconstruction data from electron tomographic images is often required for analyzing and visualizing biological structures, making it a powerful tool for quantitative descriptions of whole cell structures and understanding biological functions. However, these cellular structures are rather difficult to automatically separate or quantify from view owing to complex molecular environment and the limitations of reconstruction data of ET. In this paper, we propose a single end-to-end deep fully-convolutional semantic segmentation network dubbed SegET with rich contextual features which fully exploitsthe multi-scale and multi-level contextual information and reduces the loss of details of cellular structures in ET images. We trained and evaluated our network on the electron tomogram of the CTL Immunological Synapse from Cell Image library. Our results demonstrate that SegET can automatically segment accurately and outperform all other baseline methods on each individual structure in our ET dataset.
Estimating and abstracting the 3D structure of bones using neural networks on X-ray (2D) images
Jan 16, 2020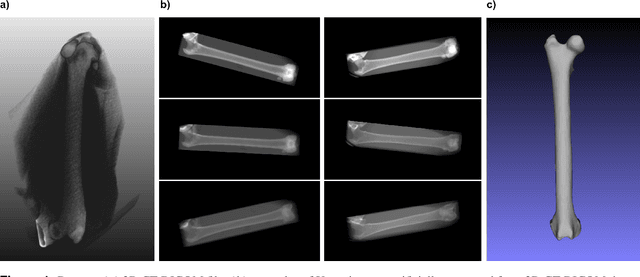
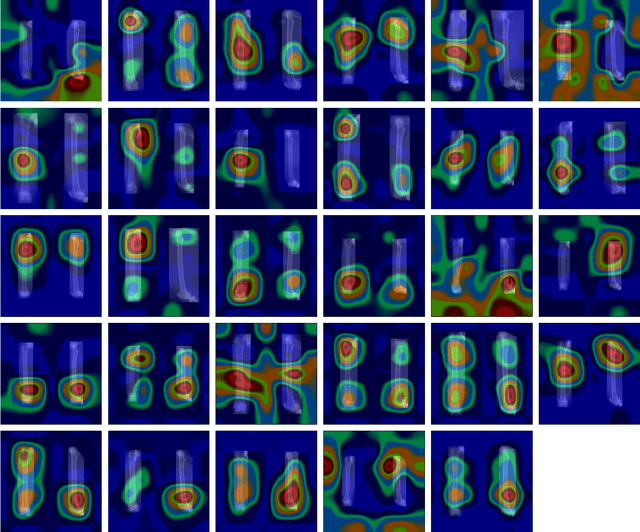


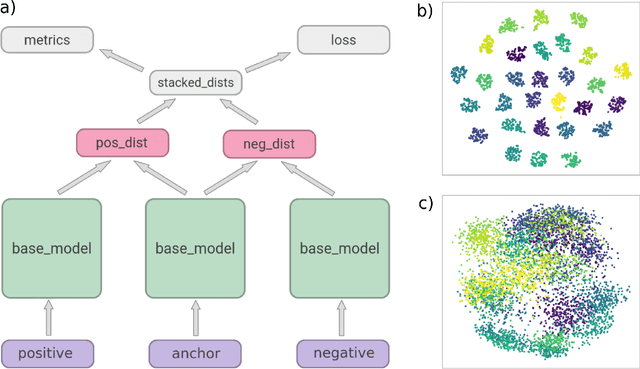
In this paper, we present a deep-learning based method for estimating the 3D structure of a bone from a pair of 2D X-ray images. Our triplet loss-trained neural network selects the most closely matching 3D bone shape from a predefined set of shapes. Our predictions have an average root mean square (RMS) distance of 1.08 mm between the predicted and true shapes, making it more accurate than the average error achieved by eight other examined 3D bone reconstruction approaches. The prediction process that we use is fully automated and unlike many competing approaches, it does not rely on any previous knowledge about bone geometry. Additionally, our neural network can determine the identity of a bone based only on its X-ray image. It computes a low-dimensional representation ("embedding") of each 2D X-ray image and henceforth compares different X-ray images based only on their embeddings. An embedding holds enough information to uniquely identify the bone CT belonging to the input X-ray image with a 100% accuracy and can therefore serve as a kind of fingerprint for that bone. Possible applications include faster, image content-based bone database searches for forensic purposes.
ConvSequential-SLAM: A Sequence-based, Training-less Visual Place Recognition Technique for Changing Environments
Sep 28, 2020
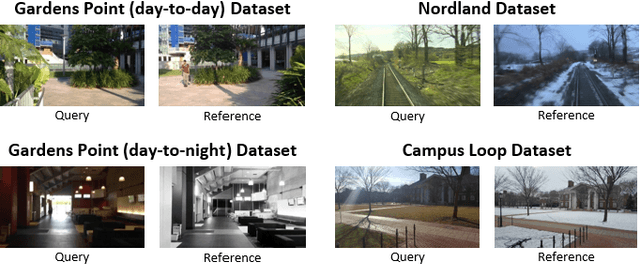


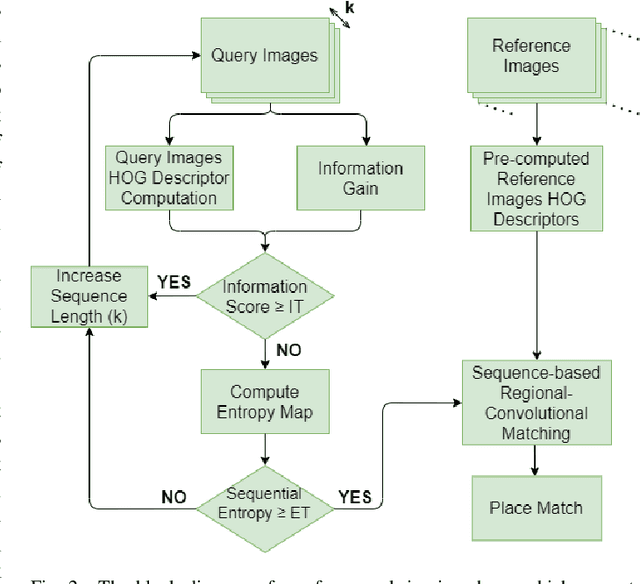
Visual Place Recognition (VPR) is the ability to correctly recall a previously visited place under changing viewpoints and appearances. A large number of handcrafted and deep-learning-based VPR techniques exist, where the former suffer from appearance changes and the latter have significant computational needs. In this paper, we present a new handcrafted VPR technique that achieves state-of-the-art place matching performance under challenging conditions. Our technique combines the best of 2 existing trainingless VPR techniques, SeqSLAM and CoHOG, which are each robust to conditional and viewpoint changes, respectively. This blend, namely ConvSequential-SLAM, utilises sequential information and block-normalisation to handle appearance changes, while using regional-convolutional matching to achieve viewpoint-invariance. We analyse content-overlap in-between query frames to find a minimum sequence length, while also re-using the image entropy information for environment-based sequence length tuning. State-of-the-art performance is reported in contrast to 8 contemporary VPR techniques on 4 public datasets. Qualitative insights and an ablation study on sequence length are also provided.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge