"Image": models, code, and papers
Does Normalization Methods Play a Role for Hyperspectral Image Classification?
Oct 09, 2017
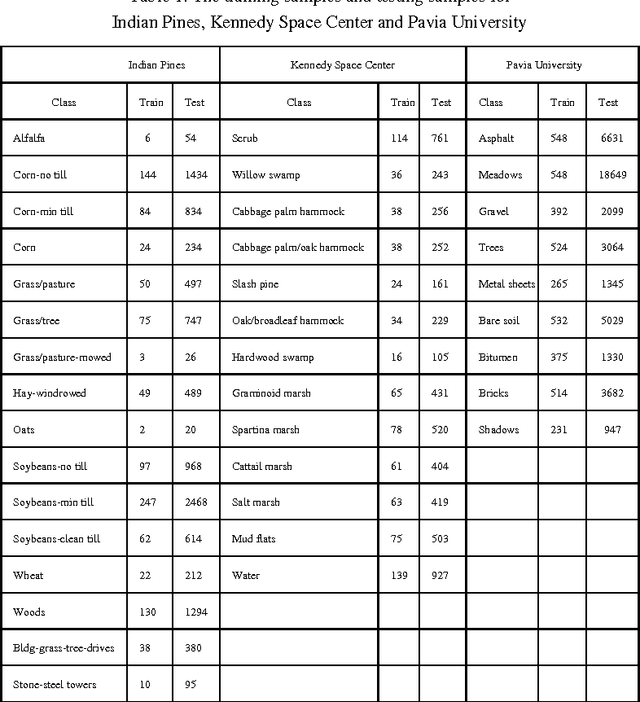
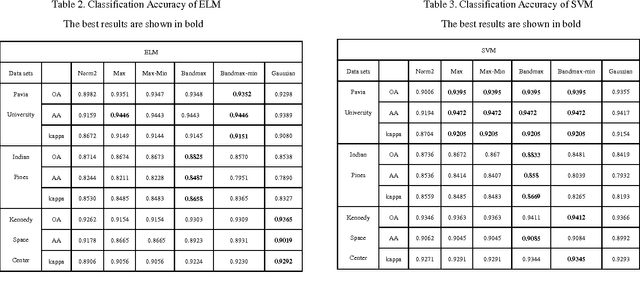
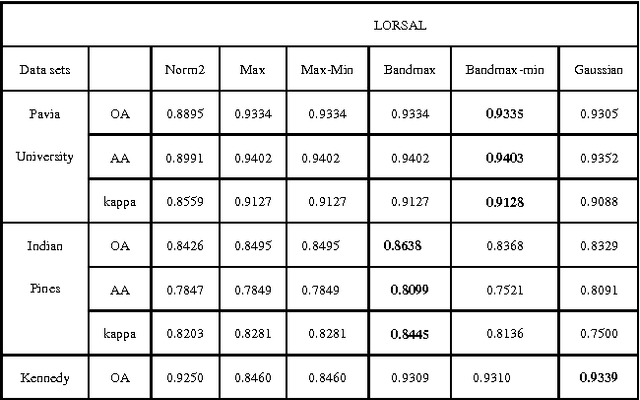
For Hyperspectral image (HSI) datasets, each class have their salient feature and classifiers classify HSI datasets according to the class's saliency features, however, there will be different salient features when use different normalization method. In this letter, we report the effect on classifiers by different normalization methods and recommend the best normalization methods for classifier after analyzing the impact of different normalization methods on classifiers. Pavia University datasets, Indian Pines datasets and Kennedy Space Center datasets will apply to several typical classifiers in order to evaluate and analysis the impact of different normalization methods on typical classifiers.
Classifying degraded images over various levels of degradation
Jun 15, 2020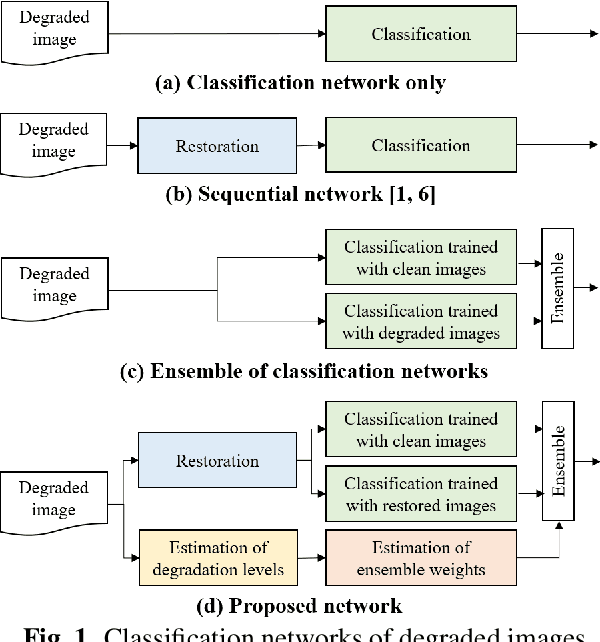
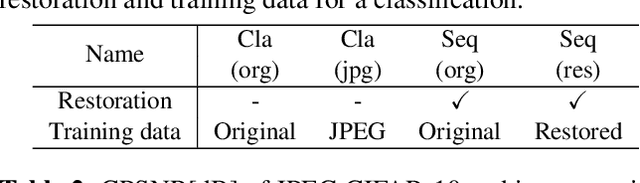
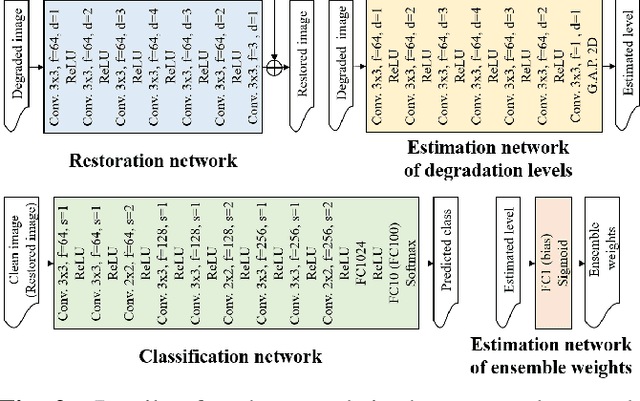
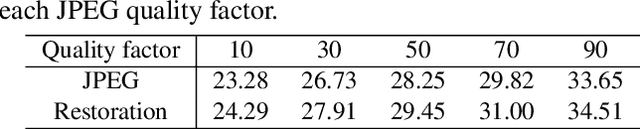
Classification for degraded images having various levels of degradation is very important in practical applications. This paper proposes a convolutional neural network to classify degraded images by using a restoration network and an ensemble learning. The results demonstrate that the proposed network can classify degraded images over various levels of degradation well. This paper also reveals how the image-quality of training data for a classification network affects the classification performance of degraded images.
FastPathology: An open-source platform for deep learning-based research and decision support in digital pathology
Nov 11, 2020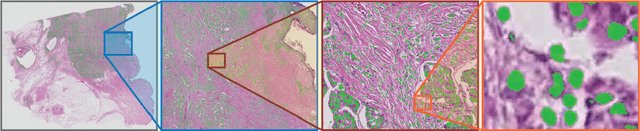
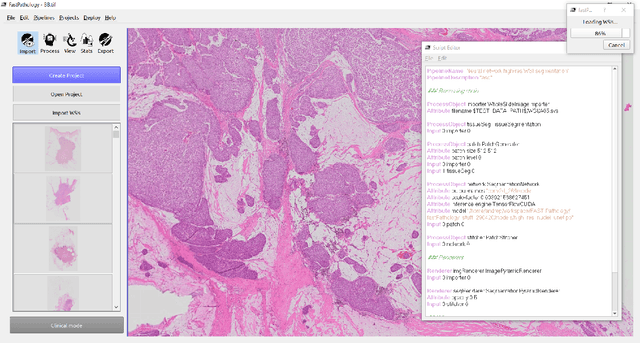
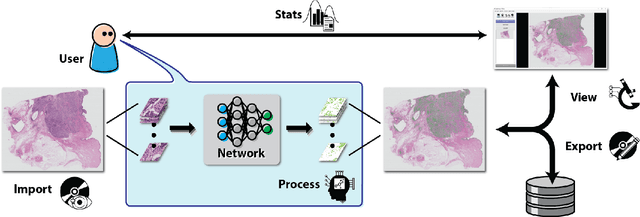
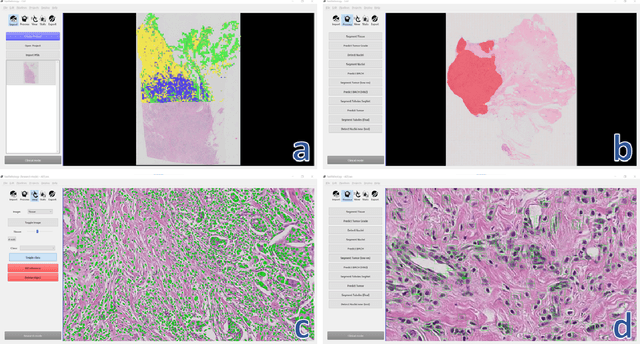
Deep convolutional neural networks (CNNs) are the current state-of-the-art for digital analysis of histopathological images. The large size of whole-slide microscopy images (WSIs) requires advanced memory handling to read, display and process these images. There are several open-source platforms for working with WSIs, but few support deployment of CNN models. These applications use third-party solutions for inference, making them less user-friendly and unsuitable for high-performance image analysis. To make deployment of CNNs user-friendly and feasible on low-end machines, we have developed a new platform, FastPathology, using the FAST framework and C++. It minimizes memory usage for reading and processing WSIs, deployment of CNN models, and real-time interactive visualization of results. Runtime experiments were conducted on four different use cases, using different architectures, inference engines, hardware configurations and operating systems. Memory usage for reading, visualizing, zooming and panning a WSI were measured, using FastPathology and three existing platforms. FastPathology performed similarly in terms of memory to the other C++ based application, while using considerably less than the two Java-based platforms. The choice of neural network model, inference engine, hardware and processors influenced runtime considerably. Thus, FastPathology includes all steps needed for efficient visualization and processing of WSIs in a single application, including inference of CNNs with real-time display of the results. Source code, binary releases and test data can be found online on GitHub at https://github.com/SINTEFMedtek/FAST-Pathology/.
A General-Purpose Dehazing Algorithm based on Local Contrast Enhancement Approaches
May 31, 2020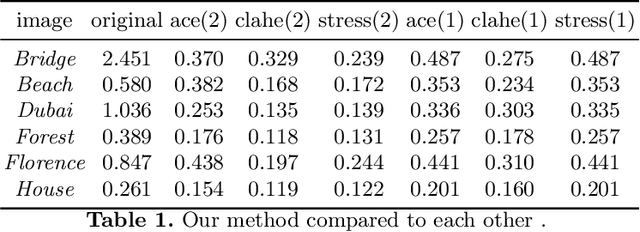

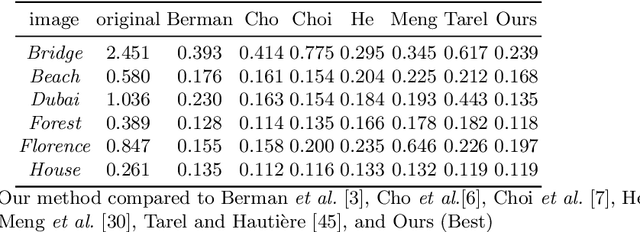

Dehazing is in the image processing and computer vision communities, the task of enhancing the image taken in foggy conditions. To better understand this type of algorithm, we present in this document a dehazing method which is suitable for several local contrast adjustment algorithms. We base it on two filters. The first filter is built with a step of normalization with some other statistical tricks while the last represents the local contrast improvement algorithm. Thus, it can work on both CPU and GPU for real-time applications. We hope that our approach will open the door to new ideas in the community. Other advantages of our method are first that it does not need to be trained, then it does not need additional optimization processing. Furthermore, it can be used as a pre-treatment or post-processing step in many vision tasks. In addition, it does not need to convert the problem into a physical interpretation, and finally that it is very fast. This family of defogging algorithms is fairly simple, but it shows promising results compared to state-of-the-art algorithms based not only on a visual assessment but also on objective criteria.
Cost-Effective Active Learning for Deep Image Classification
Jan 13, 2017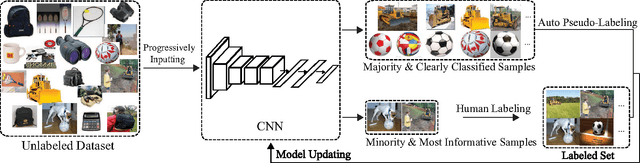

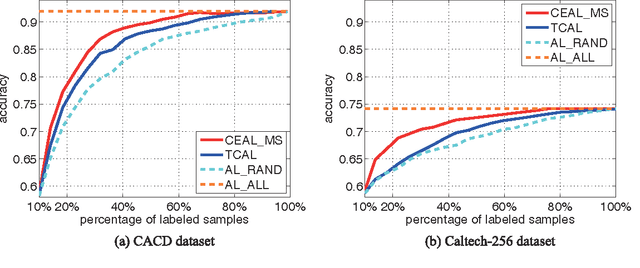
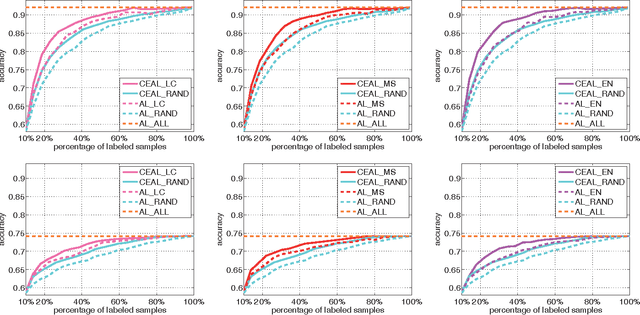
Recent successes in learning-based image classification, however, heavily rely on the large number of annotated training samples, which may require considerable human efforts. In this paper, we propose a novel active learning framework, which is capable of building a competitive classifier with optimal feature representation via a limited amount of labeled training instances in an incremental learning manner. Our approach advances the existing active learning methods in two aspects. First, we incorporate deep convolutional neural networks into active learning. Through the properly designed framework, the feature representation and the classifier can be simultaneously updated with progressively annotated informative samples. Second, we present a cost-effective sample selection strategy to improve the classification performance with less manual annotations. Unlike traditional methods focusing on only the uncertain samples of low prediction confidence, we especially discover the large amount of high confidence samples from the unlabeled set for feature learning. Specifically, these high confidence samples are automatically selected and iteratively assigned pseudo-labels. We thus call our framework "Cost-Effective Active Learning" (CEAL) standing for the two advantages.Extensive experiments demonstrate that the proposed CEAL framework can achieve promising results on two challenging image classification datasets, i.e., face recognition on CACD database [1] and object categorization on Caltech-256 [2].
Interpretable and synergistic deep learning for visual explanation and statistical estimations of segmentation of disease features from medical images
Nov 11, 2020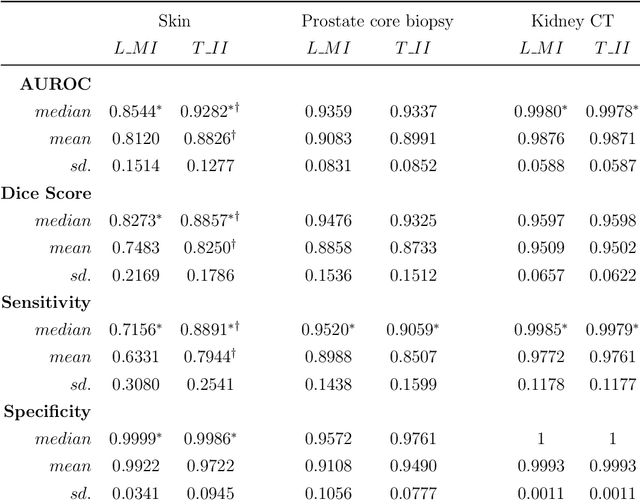
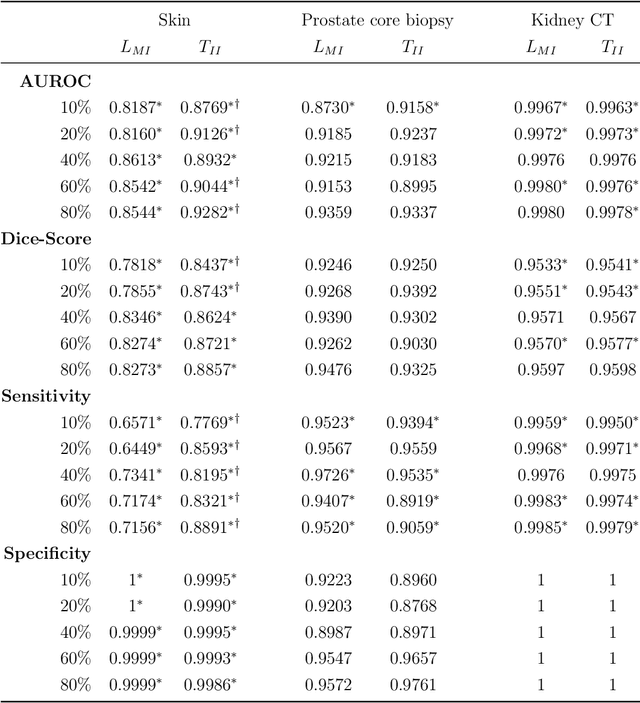
Deep learning (DL) models for disease classification or segmentation from medical images are increasingly trained using transfer learning (TL) from unrelated natural world images. However, shortcomings and utility of TL for specialized tasks in the medical imaging domain remain unknown and are based on assumptions that increasing training data will improve performance. We report detailed comparisons, rigorous statistical analysis and comparisons of widely used DL architecture for binary segmentation after TL with ImageNet initialization (TII-models) with supervised learning with only medical images(LMI-models) of macroscopic optical skin cancer, microscopic prostate core biopsy and Computed Tomography (CT) DICOM images. Through visual inspection of TII and LMI model outputs and their Grad-CAM counterparts, our results identify several counter intuitive scenarios where automated segmentation of one tumor by both models or the use of individual segmentation output masks in various combinations from individual models leads to 10% increase in performance. We also report sophisticated ensemble DL strategies for achieving clinical grade medical image segmentation and model explanations under low data regimes. For example; estimating performance, explanations and replicability of LMI and TII models described by us can be used for situations in which sparsity promotes better learning. A free GitHub repository of TII and LMI models, code and more than 10,000 medical images and their Grad-CAM output from this study can be used as starting points for advanced computational medicine and DL research for biomedical discovery and applications.
Gen-LaneNet: A Generalized and Scalable Approach for 3D Lane Detection
Mar 24, 2020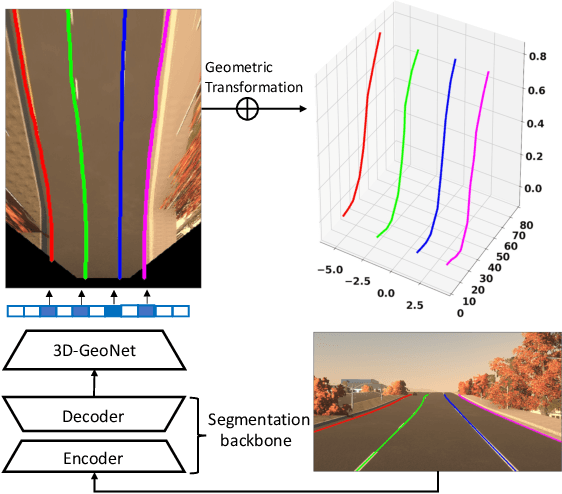
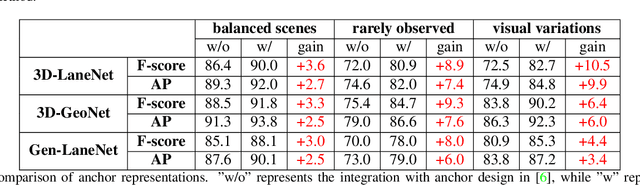
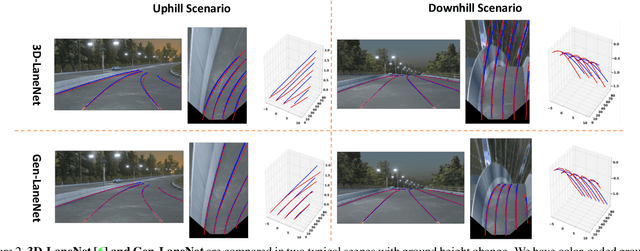
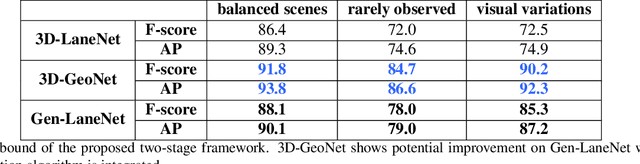
We present a generalized and scalable method, called Gen-LaneNet, to detect 3D lanes from a single image. The method, inspired by the latest state-of-the-art 3D-LaneNet, is a unified framework solving image encoding, spatial transform of features and 3D lane prediction in a single network. However, we propose unique designs for Gen-LaneNet in two folds. First, we introduce a new geometry-guided lane anchor representation in a new coordinate frame and apply a specific geometric transformation to directly calculate real 3D lane points from the network output. We demonstrate that aligning the lane points with the underlying top-view features in the new coordinate frame is critical towards a generalized method in handling unfamiliar scenes. Second, we present a scalable two-stage framework that decouples the learning of image segmentation subnetwork and geometry encoding subnetwork. Compared to 3D-LaneNet, the proposed Gen-LaneNet drastically reduces the amount of 3D lane labels required to achieve a robust solution in real-world application. Moreover, we release a new synthetic dataset and its construction strategy to encourage the development and evaluation of 3D lane detection methods. In experiments, we conduct extensive ablation study to substantiate the proposed Gen-LaneNet significantly outperforms 3D-LaneNet in average precision(AP) and F-score.
Image Restoration Using Very Deep Convolutional Encoder-Decoder Networks with Symmetric Skip Connections
Sep 01, 2016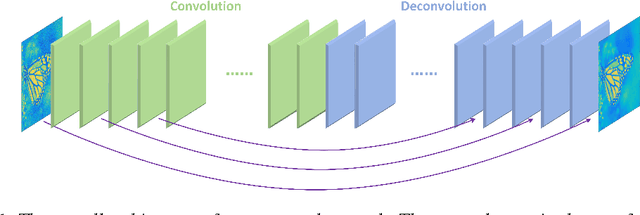
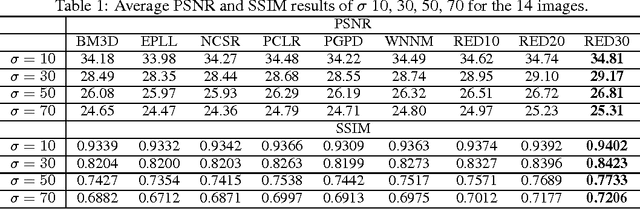
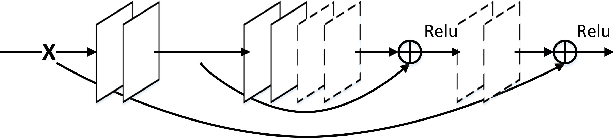
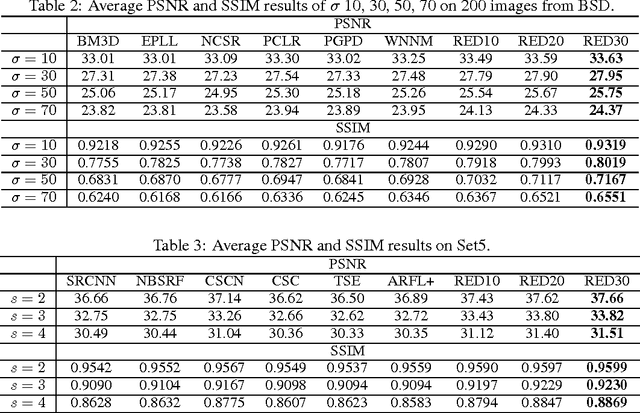
In this paper, we propose a very deep fully convolutional encoding-decoding framework for image restoration such as denoising and super-resolution. The network is composed of multiple layers of convolution and de-convolution operators, learning end-to-end mappings from corrupted images to the original ones. The convolutional layers act as the feature extractor, which capture the abstraction of image contents while eliminating noises/corruptions. De-convolutional layers are then used to recover the image details. We propose to symmetrically link convolutional and de-convolutional layers with skip-layer connections, with which the training converges much faster and attains a higher-quality local optimum. First, The skip connections allow the signal to be back-propagated to bottom layers directly, and thus tackles the problem of gradient vanishing, making training deep networks easier and achieving restoration performance gains consequently. Second, these skip connections pass image details from convolutional layers to de-convolutional layers, which is beneficial in recovering the original image. Significantly, with the large capacity, we can handle different levels of noises using a single model. Experimental results show that our network achieves better performance than all previously reported state-of-the-art methods.
Rotation-invariant Mixed Graphical Model Network for 2D Hand Pose Estimation
Feb 05, 2020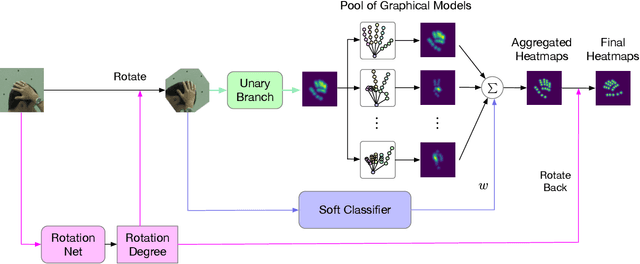
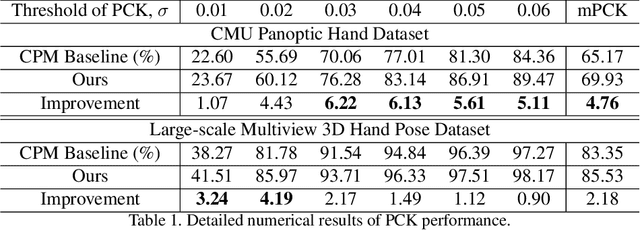
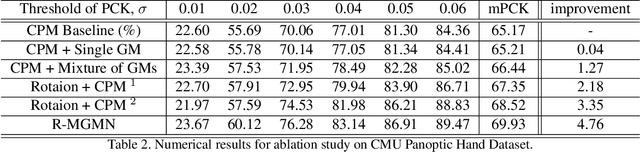

In this paper, we propose a new architecture named Rotation-invariant Mixed Graphical Model Network (R-MGMN) to solve the problem of 2D hand pose estimation from a monocular RGB image. By integrating a rotation net, the R-MGMN is invariant to rotations of the hand in the image. It also has a pool of graphical models, from which a combination of graphical models could be selected, conditioning on the input image. Belief propagation is performed on each graphical model separately, generating a set of marginal distributions, which are taken as the confidence maps of hand keypoint positions. Final confidence maps are obtained by aggregating these confidence maps together. We evaluate the R-MGMN on two public hand pose datasets. Experiment results show our model outperforms the state-of-the-art algorithm which is widely used in 2D hand pose estimation by a noticeable margin.
SegET: Deep Neural Network with Rich Contextual Features for Cellular Structures Segmentation in Electron Tomography Image
Nov 28, 2018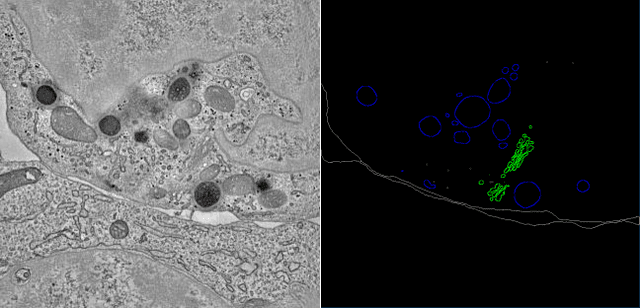
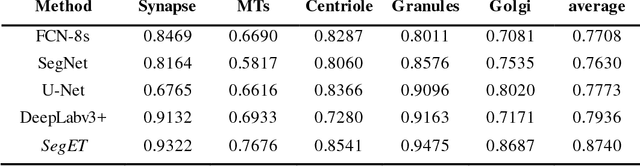
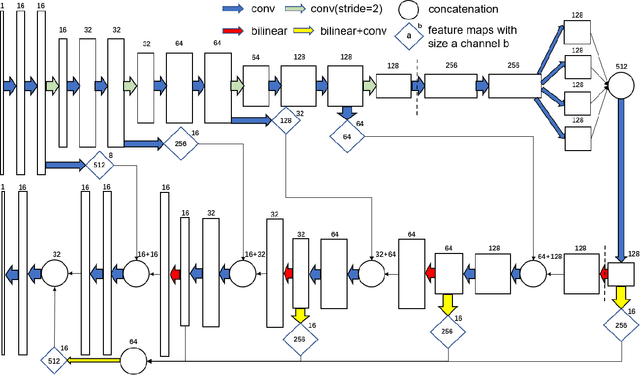
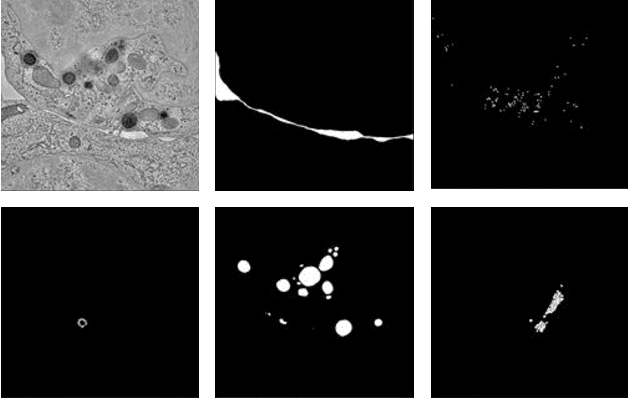
Electron tomography (ET) allows high-resolution reconstructions of macromolecular complexes at nearnative state. Cellular structures segmentation in the reconstruction data from electron tomographic images is often required for analyzing and visualizing biological structures, making it a powerful tool for quantitative descriptions of whole cell structures and understanding biological functions. However, these cellular structures are rather difficult to automatically separate or quantify from view owing to complex molecular environment and the limitations of reconstruction data of ET. In this paper, we propose a single end-to-end deep fully-convolutional semantic segmentation network dubbed SegET with rich contextual features which fully exploitsthe multi-scale and multi-level contextual information and reduces the loss of details of cellular structures in ET images. We trained and evaluated our network on the electron tomogram of the CTL Immunological Synapse from Cell Image library. Our results demonstrate that SegET can automatically segment accurately and outperform all other baseline methods on each individual structure in our ET dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge